4NRV
 
 | | Crystal Structure of non-edited human NEIL1 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Endonuclease 8-like 1 | | Authors: | Prakash, A, Doublie, S. | | Deposit date: | 2013-11-27 | | Release date: | 2014-01-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Genome and cancer single nucleotide polymorphisms of the human NEIL1 DNA glycosylase: Activity, structure, and the effect of editing.
Dna Repair, 14, 2014
|
|
5FBW
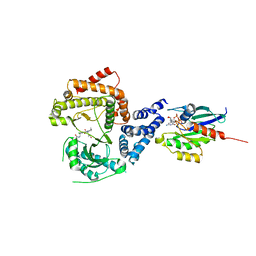 
 | | PI4KB in complex with Rab11 and the MI369 Inhibitor | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, Phosphatidylinositol 4-kinase beta,Phosphatidylinositol 4-kinase beta, Ras-related protein Rab-11A, ... | | Authors: | Chalupska, D, Mejdrova, I, Nencka, R, Boura, E. | | Deposit date: | 2015-12-14 | | Release date: | 2016-12-28 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.487 Å) | | Cite: | PI4KB in complex with Rab11 and the MI369 Inhibitor
To Be Published
|
|
1YWR
 
 | | Crystal Structure Analysis of inactive P38 kinase domain in complex with a Monocyclic Pyrazolone Inhibitor | | Descriptor: | 4-(4-FLUOROPHENYL)-1-METHYL-5-(2-{[(1S)-1-PHENYLETHYL]AMINO}PYRIMIDIN-4-YL)-2-PIPERIDIN-4-YL-1,2-DIHYDRO-3H-PYRAZOL-3-ONE, Mitogen-activated protein kinase 14 | | Authors: | Golebiowski, A, Townes, J.A, Laufersweiler, M.J, Brugel, T.A, Clark, M.P, Clark, C.M, Djung, J.F, Laughlin, S.K, Sabat, M.P, Bookland, R.G, Vanrens, J.C, De, B, Hsieh, L.C, Janusz, M.J, Walter, R.L, Webster, M.E, Mekel, M.J. | | Deposit date: | 2005-02-18 | | Release date: | 2005-05-10 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The development of monocyclic pyrazolone based cytokine synthesis inhibitors.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
4NDA
 
 | | Crystal structure of 3-nitro-tyrosine tRNA synthetase (5B) bound to 3-nitro-tyrosine | | Descriptor: | GLYCEROL, META-NITRO-TYROSINE, SODIUM ION, ... | | Authors: | Cooley, R.B, Driggers, C.M, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2013-10-25 | | Release date: | 2014-03-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis of Improved Second-Generation 3-Nitro-tyrosine tRNA Synthetases.
Biochemistry, 53, 2014
|
|
1Z36
 
 | | Crystal structure of Trichomonas vaginalis purine nucleoside phosphorylase complexed with formycin A | | Descriptor: | (1S)-1-(7-amino-1H-pyrazolo[4,3-d]pyrimidin-3-yl)-1,4-anhydro-D-ribitol, purine nucleoside phosphorylase | | Authors: | Zhang, Y, Wang, W.H, Wu, S.W, Wang, C.C, Ealick, S.E. | | Deposit date: | 2005-03-10 | | Release date: | 2005-03-29 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Identification of a subversive substrate of Trichomonas vaginalis purine nucleoside phosphorylase and the crystal structure of the enzyme-substrate complex.
J.Biol.Chem., 280, 2005
|
|
7SC6
 
 | |
3VKM
 
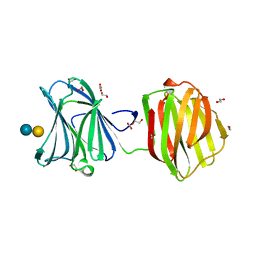 | | Protease-resistant mutant form of Human Galectin-8 in complex with sialyllactose and lactose | | Descriptor: | 1,2-ETHANEDIOL, Galectin-8, N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose, ... | | Authors: | Yoshida, H, Yamashita, S, Teraoka, M, Nakakita, S, Nishi, N, Kamitori, S. | | Deposit date: | 2011-11-18 | | Release date: | 2012-09-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | X-ray structure of a protease-resistant mutant form of human galectin-8 with two carbohydrate recognition domains
Febs J., 279, 2012
|
|
5FNH
 
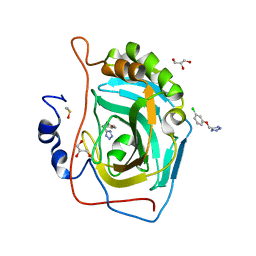 | | Native state mass spectrometry, surface plasmon resonance and X-ray crystallography correlate strongly as a fragment screening combination | | Descriptor: | 5-[(3-chloranylphenoxy)methyl]-1,2,4-triaza-3-azanidacyclopenta-1,4-diene, CARBONIC ANHYDRASE 2, DIMETHYL SULFOXIDE, ... | | Authors: | Woods, L.A, Dolezal, O, Ren, B, Ryan, J.H, Peat, T.S, Poulsen, S.A. | | Deposit date: | 2015-11-15 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Native State Mass Spectrometry, Surface Plasmon Resonance and X-Ray Crystallography Correlate Strongly as a Fragment Screening Combination.
J.Med.Chem., 59, 2016
|
|
9GFD
 
 | | Crystal structure of ASO binding Fab fragment with ASO139 | | Descriptor: | 1,2-ETHANEDIOL, 1-[(1~{S},3~{R},4~{R},7~{S})-7-[[(1~{R},3~{R},4~{R},7~{S})-7-[[(1~{R},3~{R},4~{R},7~{S})-3-(6-aminopurin-9-yl)-7-[[(2~{R},3~{S},5~{R})-5-(4-azanyl-2-oxidanylidene-pyrimidin-1-yl)-3-oxidanyl-oxolan-2-yl]methoxy-sulfanyl-phosphoryl]oxy-2,5-dioxabicyclo[2.2.1]heptan-1-yl]methoxy-sulfanyl-phosphoryl]oxy-3-(4-azanyl-5-methyl-2-oxidanylidene-pyrimidin-1-yl)-2,5-dioxabicyclo[2.2.1]heptan-1-yl]methoxy-sulfanyl-phosphoryl]oxy-1-(hydroxymethyl)-2,5-dioxabicyclo[2.2.1]heptan-3-yl]-5-methyl-pyrimidine-2,4-dione, Fab Fragment light chain, ... | | Authors: | Hsia, H.-E, Zanini, C, Simonneau, C, Fraidling, J, Kraft, T, Mayer, K, Sommer, A, Indlekofer, A, Wirth, T, Benz, J, Geroges, G, Langer, M.L, Gassner, C, Larraillet, V, Manso, M, Ravn, J, Hofer, K, Emrich, T, Niewoehner, J, Schumacher, F, Brinkmann, U. | | Deposit date: | 2024-08-09 | | Release date: | 2025-08-20 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Improved Targeted Delivery of Antisense Oligonucleotide Conjugates
with the Antibody Mask
To Be Published
|
|
7SCF
 
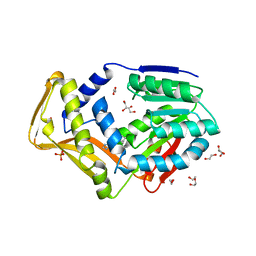 | | M. tb EgtD in complex with HD2 | | Descriptor: | (2S)-3-(1H-imidazol-5-yl)-2-(1H-pyrrol-1-yl)propanoic acid, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Sudasinghe, T.D, Ronning, D.R. | | Deposit date: | 2021-09-28 | | Release date: | 2021-12-01 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Inhibitors of Mycobacterium tuberculosis EgtD target both substrate binding sites to limit hercynine production.
Sci Rep, 11, 2021
|
|
3E37
 
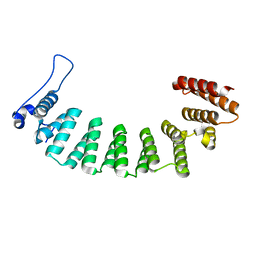 | |
7SF6
 
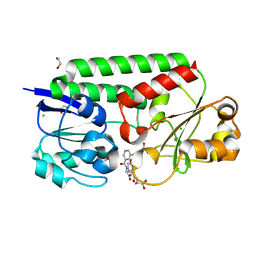 | | Crystal Structure of Siderophore Binding Protein FatB from Desulfitobacterium hafniense | | Descriptor: | 1,2-ETHANEDIOL, 2-(2,3-DIHYDROXY-BENZOYLAMINO)-3-HYDROXY-PROPIONIC ACID, CHLORIDE ION, ... | | Authors: | Kim, Y, Patel, H.P, Nordquist, K.A, Schaab, K.M, Sha, J, Babnigg, G, Bond, A.H, Joachimiak, A, Midwest Center for Structural Genomics, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2021-10-03 | | Release date: | 2021-12-15 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Crystal Structure of Siderophore Binding Protein FatB from Desulfitobacterium hafniense
To Be Published
|
|
7SSE
 
 | | Crystal structure of the WDR domain of human DCAF1 in complex with CYCA-117-70 | | Descriptor: | DDB1- and CUL4-associated factor 1, N-[(3R)-1-(3-fluorophenyl)piperidin-3-yl]-6-(morpholin-4-yl)pyrimidin-4-amine | | Authors: | Kimani, S, Owen, J, Li, A, Dong, A, Li, Y, Hutchinson, A, Seitova, A, Shahani, V.M, Schapira, M, Arrowsmith, C.H, Edwards, A.M, Halabelian, L, Structural Genomics Consortium (SGC) | | Deposit date: | 2021-11-10 | | Release date: | 2021-12-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Discovery of a Novel DCAF1 Ligand Using a Drug-Target Interaction Prediction Model: Generalizing Machine Learning to New Drug Targets.
J.Chem.Inf.Model., 63, 2023
|
|
6DIQ
 
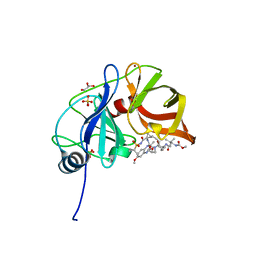 | |
1Z2Q
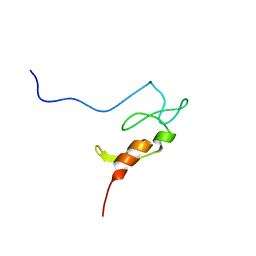 
 | |
2PBW
 
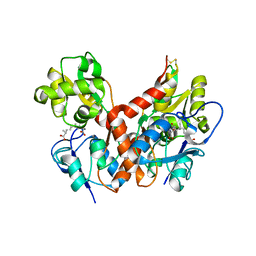 | | Crystal Structure of the Ligand-Binding Core of iGluR5 in Complex with the Partial agonist Domoic Acid at 2.5 A Resolution | | Descriptor: | (2S,3S,4S)-2-CARBOXY-4-[(1Z,3E,5R)-5-CARBOXY-1-METHYL-1,3-HEXADIENYL]-3-PYRROLIDINEACETIC ACID, Glutamate receptor, ionotropic kainate 1 | | Authors: | Hald, H, Naur, P, Gajhede, M, Kastrup, J.S. | | Deposit date: | 2007-03-29 | | Release date: | 2007-07-03 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Partial agonism and antagonism of the ionotropic glutamate receptor iGLuR5: structures of the ligand-binding core in complex with domoic acid and 2-amino-3-[5-tert-butyl-3-(phosphonomethoxy)-4-isoxazolyl]propionic acid.
J.Biol.Chem., 282, 2007
|
|
3N18
 
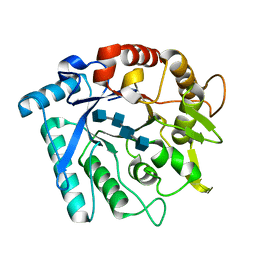 | | Crystal stricture of E145G/Y227F chitinase in complex with NAG from Bacillus cereus NCTU2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Chitinase A | | Authors: | Hsieh, Y.-C, Wu, Y.-J, Wu, W.-G, Li, Y.-K, Chen, C.-J. | | Deposit date: | 2010-05-15 | | Release date: | 2010-08-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structures of bacillus cereus NCTU2 chitinase complexes with chitooligomers reveal novel substrate binding for catalysis: a chitinase without chitin-binding and insertion domains
J.Biol.Chem., 285, 2010
|
|
8PQW
 
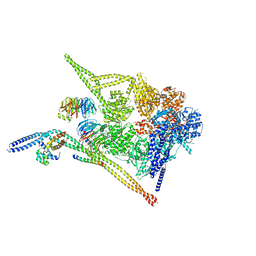 | | Cytoplasmic dynein-1 motor domain bound to dynactin-p150glued and LIS1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Cytoplasmic dynein 1 heavy chain 1, ... | | Authors: | Singh, K, Lau, C.K, Manigrasso, G, Gassmann, R, Carter, A.P. | | Deposit date: | 2023-07-12 | | Release date: | 2024-03-27 | | Last modified: | 2024-04-10 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Molecular mechanism of dynein-dynactin complex assembly by LIS1.
Science, 383, 2024
|
|
5FDL
 
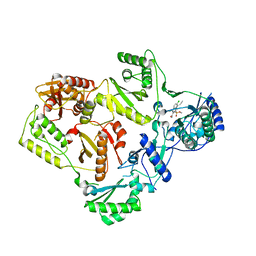 | | Crystal Structure of K103N/Y181C Mutant HIV-1 Reverse Transcriptase (RT) in Complex with IDX899 | | Descriptor: | P51 Reverse transcriptase, P66 Reverse transcriptase, methyl (R)-(2-carbamoyl-5-chloro-1H-indol-3-yl)[3-(2-cyanoethyl)-5-methylphenyl]phosphinate | | Authors: | Dousson, C.B, Alexandre, F.-R, Convard, T, Fisher, M, Lamers, M.B.A.C, Leonard, P.M. | | Deposit date: | 2015-12-16 | | Release date: | 2016-02-17 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Discovery of the Aryl-phospho-indole IDX899, a Highly Potent Anti-HIV Non-nucleoside Reverse Transcriptase Inhibitor.
J.Med.Chem., 59, 2016
|
|
4ND6
 
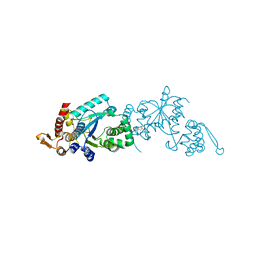 | | Crystal structure of apo 3-nitro-tyrosine tRNA synthetase (5B) in the open form | | Descriptor: | GLYCEROL, Tyrosine-tRNA ligase | | Authors: | Cooley, R.B, Driggers, C.M, Karplus, P.A, Mehl, R.A. | | Deposit date: | 2013-10-25 | | Release date: | 2014-03-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis of Improved Second-Generation 3-Nitro-tyrosine tRNA Synthetases.
Biochemistry, 53, 2014
|
|
7SPP
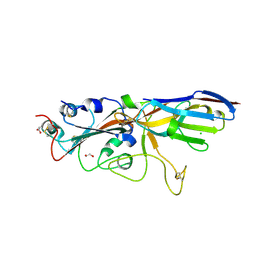 
 | |
1MFU
 
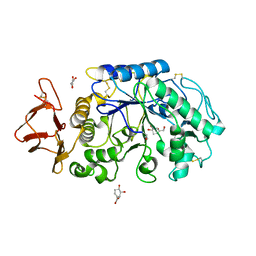 | | Probing the role of a mobile loop in human salivary amylase: Structural studies on the loop-deleted mutant | | Descriptor: | 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 4-amino-4,6-dideoxy-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, 5-HYDROXYMETHYL-CHONDURITOL, ... | | Authors: | Ramasubbu, N, Ragunath, C, Mishra, P.J. | | Deposit date: | 2002-08-13 | | Release date: | 2002-11-20 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Probing the role of a mobile loop in substrate binding and enzyme activity of
human salivary amylase.
J.Mol.Biol., 325, 2003
|
|
9GTC
 
 | | Crystal structure of human lysosomal acid-alpha-glucosidase, GAA, in complex with iminosugar compound 4g | | Descriptor: | (2~{R},3~{R},4~{R},5~{S})-2-(hydroxymethyl)-2-pentyl-piperidine-3,4,5-triol, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Sulzenbacher, G, Roig-Zamboni, V, Moracci, M, Parenti, G, Py, S. | | Deposit date: | 2024-09-17 | | Release date: | 2025-10-01 | | Last modified: | 2025-10-08 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | C -Branched Iminosugars as Selective Pharmacological Chaperones of Lysosomal alpha-Glucosidase for the Treatment of Pompe Disease.
J.Med.Chem., 68, 2025
|
|
9GTD
 
 | | Crystal structure of human lysosomal acid-alpha-glucosidase, GAA, in complex with iminosugar compound 4i | | Descriptor: | (2~{R},3~{R},4~{R},5~{S})-2-(2-cyclohexylethyl)-2-(hydroxymethyl)piperidine-3,4,5-triol, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Sulzenbacher, G, Roig-Zamboni, V, Moracci, M, Parenti, G, Py, S. | | Deposit date: | 2024-09-17 | | Release date: | 2025-10-01 | | Last modified: | 2025-10-08 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | C -Branched Iminosugars as Selective Pharmacological Chaperones of Lysosomal alpha-Glucosidase for the Treatment of Pompe Disease.
J.Med.Chem., 68, 2025
|
|
9GTN
 
 | | Crystal structure of human lysosomal acid-alpha-glucosidase, GAA, in complex with iminosugar compound 4k | | Descriptor: | (2~{R},3~{R},4~{R},5~{S})-2-(2-hydroxyethyl)-2-(hydroxymethyl)piperidine-3,4,5-triol, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Sulzenbacher, G, Roig-Zamboni, V, Moracci, M, Parenti, G, Py, S. | | Deposit date: | 2024-09-18 | | Release date: | 2025-10-01 | | Last modified: | 2025-10-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | C -Branched Iminosugars as Selective Pharmacological Chaperones of Lysosomal alpha-Glucosidase for the Treatment of Pompe Disease.
J.Med.Chem., 68, 2025
|
|
