8FW0
 
 | | MtrR from Neisseria gonorrhoeae bound to beta-Estradiol | | Descriptor: | ESTRADIOL, GLYCEROL, HTH-type transcriptional regulator MtrR, ... | | Authors: | Hooks, G.H, Brennan, R.G. | | Deposit date: | 2023-01-20 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Hormonal steroids induce multidrug resistance and stress response genes in Neisseria gonorrhoeae by binding to MtrR.
Nat Commun, 15, 2024
|
|
8FW3
 
 | | MtrR from Neisseria gonorrhoeae bound to Testosterone | | Descriptor: | HTH-type transcriptional regulator MtrR, PHOSPHATE ION, TESTOSTERONE | | Authors: | Hooks, G.H, Brennan, R.G. | | Deposit date: | 2023-01-20 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Hormonal steroids induce multidrug resistance and stress response genes in Neisseria gonorrhoeae by binding to MtrR.
Nat Commun, 15, 2024
|
|
8FW8
 
 | | MtrR from Neisseria gonorrhoeae bound to Progesterone | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, HTH-type transcriptional regulator MtrR, PHOSPHATE ION, ... | | Authors: | Hooks, G.H, Brennan, R.G. | | Deposit date: | 2023-01-20 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Hormonal steroids induce multidrug resistance and stress response genes in Neisseria gonorrhoeae by binding to MtrR.
Nat Commun, 15, 2024
|
|
3BUK
 | | Crystal Structure of the Neurotrophin-3 and p75NTR Symmetrical Complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Neurotrophin-3, Tumor necrosis factor receptor superfamily member 16 | | Authors: | Jiang, T, Gong, Y, Cao, P, Yu, H.J. | | Deposit date: | 2008-01-02 | | Release date: | 2008-07-15 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the neurotrophin-3 and p75NTR symmetrical complex.
Nature, 454, 2008
|
|
2F4M
 
 | | The Mouse PNGase-HR23 Complex Reveals a Complete Remodulation of the Protein-Protein Interface Compared to its Yeast Orthologs | | Descriptor: | CHLORIDE ION, UV excision repair protein RAD23 homolog B, ZINC ION, ... | | Authors: | Zhao, G, Zhou, X, Wang, L, Kisker, C, Lennarz, W.J, Schindelin, H. | | Deposit date: | 2005-11-23 | | Release date: | 2006-03-07 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure of the mouse peptide N-glycanase-HR23 complex suggests co-evolution of the endoplasmic reticulum-associated degradation and DNA repair pathways.
J.Biol.Chem., 281, 2006
|
|
1TRH
 
 | |
2F4O
 
 | | The Mouse PNGase-HR23 Complex Reveals a Complete Remodulation of the Protein-Protein Interface Compared to its Yeast Orthologs | | Descriptor: | CHLORIDE ION, PHQ-VAL-ALA-ASP-CF0, XP-C repair complementing complex 58 kDa protein, ... | | Authors: | Zhao, G, Zhou, X, Wang, L, Kisker, C, Lennarz, W.J, Schindelin, H. | | Deposit date: | 2005-11-23 | | Release date: | 2006-03-07 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structure of the mouse peptide N-glycanase-HR23 complex suggests co-evolution of the endoplasmic reticulum-associated degradation and DNA repair pathways.
J.Biol.Chem., 281, 2006
|
|
2VOM
 
 | | Structural basis of human triosephosphate isomerase deficiency. Mutation E104D and correlation to solvent perturbation. | | Descriptor: | TRIOSEPHOSPHATE ISOMERASE | | Authors: | Rodriguez-Almazan, C, Arreola-Alemon, R, Rodriguez-Larrea, D, Aguirre-Lopez, B, de Gomez-Puyou, M.T, Perez-Montfort, R, Costas, M, Gomez-Puyou, A, Torres-Larios, A. | | Deposit date: | 2008-02-19 | | Release date: | 2008-06-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural Basis of Human Triosephosphate Isomerase Deficiency: Mutation E104D is Related to Alterations of a Conserved Water Network at the Dimer Interface.
J.Biol.Chem., 283, 2008
|
|
2N97
 
 | | DD homodimer | | Descriptor: | Tumor necrosis factor receptor superfamily member 16 | | Authors: | Lin, Z, Ibanez, C.F. | | Deposit date: | 2015-11-07 | | Release date: | 2015-12-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis of death domain signaling in the p75 neurotrophin receptor
Elife, 4, 2015
|
|
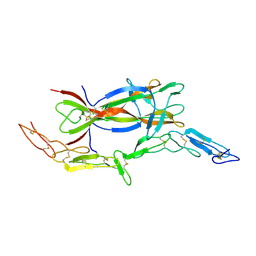
1SG1
 
 | |
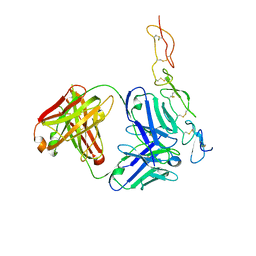
2H9G
 
 | |
3FWQ
 
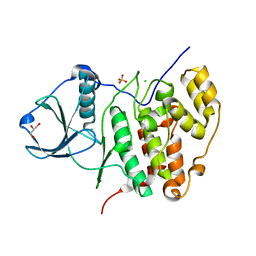 | | Inactive conformation of human protein kinase CK2 catalytic subunit | | Descriptor: | CHLORIDE ION, Casein kinase II subunit alpha, GLYCEROL, ... | | Authors: | Niefind, K, Raaf, J, Issinger, O.G. | | Deposit date: | 2009-01-19 | | Release date: | 2009-02-17 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | First inactive conformation of CK2 alpha, the catalytic subunit of protein kinase CK2
J.Mol.Biol., 386, 2009
|
|
3NJP
 
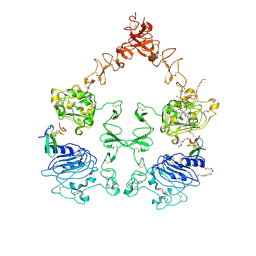 | | The Extracellular and Transmembrane Domain Interfaces in Epidermal Growth Factor Receptor Signaling | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Epidermal growth factor, Epidermal growth factor receptor, ... | | Authors: | Lu, C, Mi, L.-Z, Grey, M.J, Zhu, J, Graef, E, Yokoyama, S, Springer, T.A. | | Deposit date: | 2010-06-17 | | Release date: | 2010-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.304 Å) | | Cite: | Structural evidence for loose linkage between ligand binding and kinase activation in the epidermal growth factor receptor.
Mol.Cell.Biol., 30, 2010
|
|
2XT2
 
 | | Structure of the pentapeptide repeat protein AlbG, a resistance factor for the topoisomerase poison albicidin. | | Descriptor: | MCBG-LIKE PROTEIN, SULFATE ION | | Authors: | Vetting, M.W, Hegde, S.S, Blanchard, J.S. | | Deposit date: | 2010-10-05 | | Release date: | 2010-10-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.999 Å) | | Cite: | Pentapeptide-Repeat Proteins that Act as Topoisomerase Poison Resistance Factors Have a Common Dimer Interface.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
2XGV
 
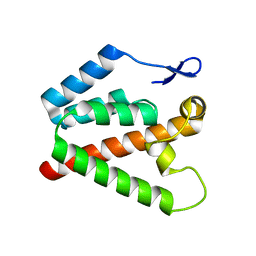 | | Structure of the N-terminal domain of capsid protein from Rabbit Endogenous Lentivirus (RELIK) | | Descriptor: | PSIV CAPSID N-TERMINAL DOMAIN | | Authors: | Goldstone, D.C, Robertson, L.E, Haire, L.F, Stoye, J.P, Taylor, I.A. | | Deposit date: | 2010-06-07 | | Release date: | 2010-09-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Analysis of Prehistoric Lentiviruses Uncovers an Ancient Molecular Interface.
Cell Host Microbe, 8, 2010
|
|
2XGU
 
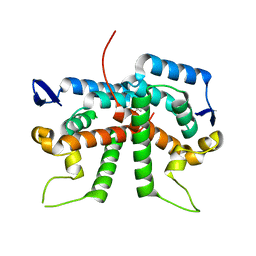 | | Structure of the N-terminal domain of capsid protein from Rabbit Endogenous Lentivirus (RELIK) | | Descriptor: | ACETATE ION, RELIK CAPSID N-TERMINAL DOMAIN | | Authors: | Goldstone, D.C, Taylor, I.A, Robertson, L.E, Haire, L.F, Stoye, J.P. | | Deposit date: | 2010-06-07 | | Release date: | 2010-09-22 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.502 Å) | | Cite: | Structural and Functional Analysis of Prehistoric Lentiviruses Uncovers an Ancient Molecular Interface.
Cell Host Microbe, 8, 2010
|
|
2N80
 
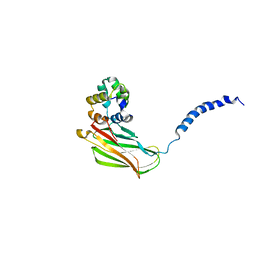 | | p75NTR DD:RhoGDI | | Descriptor: | Rho GDP-dissociation inhibitor 1, Tumor necrosis factor receptor superfamily member 16 | | Authors: | Lin, Z, Ibanez, C.F. | | Deposit date: | 2015-09-30 | | Release date: | 2015-12-23 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis of death domain signaling in the p75 neurotrophin receptor
Elife, 4, 2015
|
|
2N83
 
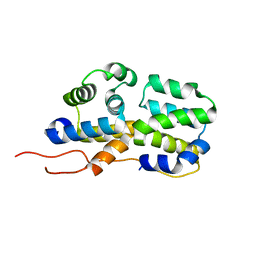 | | p75NTR DD:RIP2 CARD | | Descriptor: | Receptor-interacting serine/threonine-protein kinase 2, Tumor necrosis factor receptor superfamily member 16 | | Authors: | Lin, Z, Ibanez, C.F. | | Deposit date: | 2015-10-02 | | Release date: | 2015-12-23 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis of death domain signaling in the p75 neurotrophin receptor
Elife, 4, 2015
|
|
7OYJ
 
 | |
7OQV
 
 | |
7PDZ
 
 | | Structure of capping protein bound to the barbed end of a cytoplasmic actin filament | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin, cytoplasmic 1, ... | | Authors: | Funk, J, Merino, F, Schacks, M, Rottner, K, Raunser, S, Bieling, P. | | Deposit date: | 2021-08-09 | | Release date: | 2021-09-01 | | Last modified: | 2021-10-06 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | A barbed end interference mechanism reveals how capping protein promotes nucleation in branched actin networks.
Nat Commun, 12, 2021
|
|
7PT1
 
 | |
7PT2
 
 | |
7PT3
 
 | |
7PT4
 
 | | Actinobacterial 2-hydroxyacyl-CoA lyase (AcHACL) structure in complex with a covalently bound reaction intermediate as well as products formyl-CoA and acetone | | Descriptor: | 2-hydroxyacyl-CoA lyase, 3-[(4-AMINO-2-METHYLPYRIMIDIN-5-YL)METHYL]-2-{(1R,11R,15S,17R)-19-[(2R,3S,4R,5R)-5-(6-AMINO-9H-PURIN-9-YL)-4-HYDROXY-3-(PHOSPHONOOXY)TETRAHYDROFURAN-2-YL]-1,11,15,17-TETRAHYDROXY-12,12-DIMETHYL-15,17-DIOXIDO-6,10-DIOXO-14,16,18-TRIOXA-2-THIA-5,9-DIAZA-15,17-DIPHOSPHANONADEC-1-YL}-5-(2-{[(R)-HYDROXY(PHOSPHONOOXY)PHOSPHORYL]OXY}ETHYL)-4-METHYL-1,3-THIAZOL-3-IUM, ACETONE, ... | | Authors: | Zahn, M, Rohwerder, T. | | Deposit date: | 2021-09-25 | | Release date: | 2022-02-02 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Mechanistic details of the actinobacterial lyase-catalyzed degradation reaction of 2-hydroxyisobutyryl-CoA.
J.Biol.Chem., 298, 2022
|
|
