2EIF
 
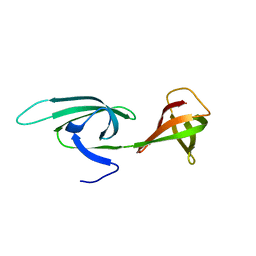 | | Eukaryotic translation initiation factor 5A from Methanococcus jannaschii | | Descriptor: | PROTEIN (EUKARYOTIC TRANSLATION INITIATION FACTOR 5A) | | Authors: | Kim, K.K, Hung, L.W, Kim, R, Kim, S.H, Berkeley Structural Genomics Center (BSGC) | | Deposit date: | 1998-10-12 | | Release date: | 1999-10-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structures of eukaryotic translation initiation factor 5A from Methanococcus jannaschii at 1.8 A resolution.
Proc.Natl.Acad.Sci.USA, 95, 1998
|
|
1UR0
 
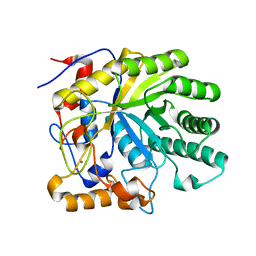 | | The structure of endo-beta-1,4-galactanase from Bacillus licheniformis in complex with two oligosaccharide products. | | Descriptor: | CALCIUM ION, GALACTANASE, TRIETHYLENE GLYCOL, ... | | Authors: | Ryttersgaard, C, Le Nours, J, Lo Leggio, L, Jorgensen, C.T, Christensen, L.L.H, Bjornvad, M, Larsen, S. | | Deposit date: | 2003-10-24 | | Release date: | 2004-10-20 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Structure of Endo-Beta-1,4-Galactanase from Bacillus Licheniformis in Complex with Two Oligosaccharide Products
J.Mol.Biol., 341, 2004
|
|
1C7R
 
 | | THE CRYSTAL STRUCTURE OF PHOSPHOGLUCOSE ISOMERASE/AUTOCRINE MOTILITY FACTOR/NEUROLEUKIN COMPLEXED WITH ITS CARBOHYDRATE PHOSPHATE INHIBITORS AND ITS SUBSTRATE RECOGNITION MECHANISM | | Descriptor: | 5-PHOSPHOARABINONIC ACID, PHOSPHOGLUCOSE ISOMERASE | | Authors: | Chou, C.-C, Meng, M, Sun, Y.-J, Hsiao, C.-D. | | Deposit date: | 2000-03-02 | | Release date: | 2000-09-13 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of phosphoglucose isomerase/autocrine motility factor/neuroleukin complexed with its carbohydrate phosphate inhibitors suggests its substrate/receptor recognition.
J.Biol.Chem., 275, 2000
|
|
1URG
 
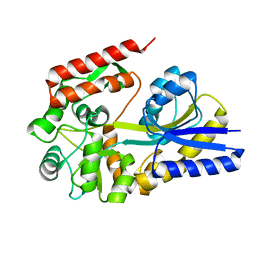 | | X-ray structures from the maltose-maltodextrin binding protein of the thermoacidophilic bacterium Alicyclobacillus acidocaldarius | | Descriptor: | MALTOSE-BINDING PROTEIN, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Schafer, K, Magnusson, U, Scheffel, F, Schiefner, A, Sandgren, M.O.J, Diederichs, K, Welte, W, Hulsmann, A, Schneider, E, Mowbray, S.L. | | Deposit date: | 2003-10-29 | | Release date: | 2003-12-11 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-Ray Structures of the Maltose-Maltodextrin-Binding Protein of the Thermoacidophilic Bacterium Alicyclobacillus Acidocaldarius Provide Insight Into Acid Stability of Proteins.
J.Mol.Biol., 335, 2004
|
|
2EMN
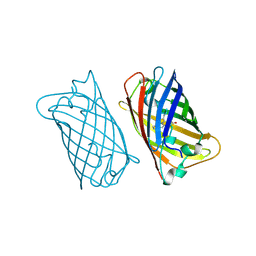 
 | |
1YQM
 
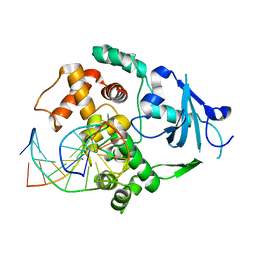 | | Catalytically inactive human 8-oxoguanine glycosylase crosslinked to 7-deazaguanine containing DNA | | Descriptor: | 5'-D(P*GP*GP*TP*AP*GP*AP*CP*CP*TP*GP*GP*AP*C)-3', 5'-D(P*GP*TP*CP*CP*AP*(7GU)P*GP*TP*CP*TP*AP*C)-3', CALCIUM ION, ... | | Authors: | Banerjee, A, Yang, W, Karplus, M, Verdine, G.L. | | Deposit date: | 2005-02-02 | | Release date: | 2005-04-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of a repair enzyme interrogating undamaged DNA elucidates recognition of damaged DNA.
Nature, 434, 2005
|
|
1YPO
 
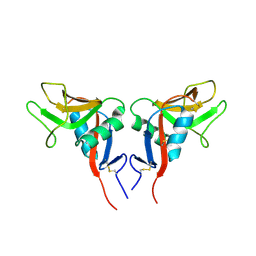 | |
1YQJ
 
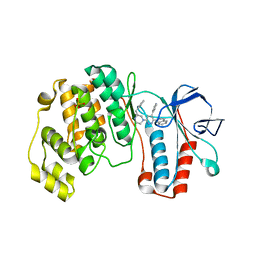 | | Crystal Structure of p38 Alpha in Complex with a Selective Pyridazine Inhibitor | | Descriptor: | 6((S)-3-BENZYLPIPERAZIN-1-YL)-3-(NAPHTHALEN-2-YL)-4-(PYRIDIN-4-YL)PYRAZINE, Mitogen-activated protein kinase 14, SULFATE ION | | Authors: | Tamayo, N, Liao, H, Goldberg, M, Syed, R, Li, V, Powers, D, Tudor, Y, Yu, V, Wong, M.L, Henkle, B, Middelton, S, Harvey, T, Jang, G, Hungate, R, Dominguez, C. | | Deposit date: | 2005-02-01 | | Release date: | 2005-04-26 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design and synthesis of potent pyridazine inhibitors of p38 MAP kinase.
Bioorg.Med.Chem.Lett., 15, 2005
|
|
1YSB
 
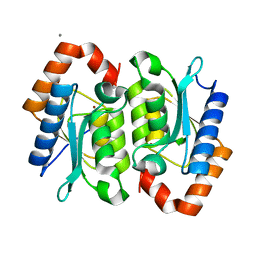 | | Yeast Cytosine Deaminase Triple Mutant | | Descriptor: | CALCIUM ION, Cytosine deaminase, ZINC ION | | Authors: | Korkegian, A, Black, M.E, Baker, D, Stoddard, B.L. | | Deposit date: | 2005-02-08 | | Release date: | 2005-05-17 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Computational thermostabilization of an enzyme.
Science, 308, 2005
|
|
1YSO
 
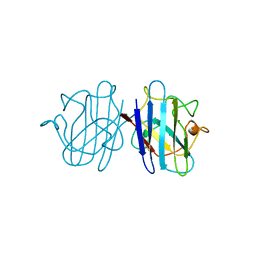 | | YEAST CU, ZN SUPEROXIDE DISMUTASE WITH THE REDUCED BRIDGE BROKEN | | Descriptor: | COPPER (I) ION, YEAST CU, ZN SUPEROXIDE DISMUTASE, ... | | Authors: | Parge, H.E, Crane, B.R, Tsang, J, Tainer, J.A. | | Deposit date: | 1995-12-21 | | Release date: | 1996-06-10 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Unusual trigonal-planar copper configuration revealed in the atomic structure of yeast copper-zinc superoxide dismutase.
Biochemistry, 35, 1996
|
|
1UUR
 
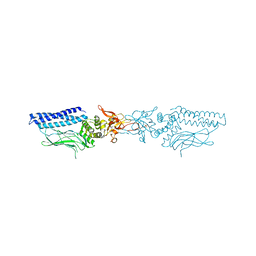 | | Structure of an activated Dictyostelium STAT in its DNA-unbound form | | Descriptor: | STATA PROTEIN | | Authors: | Soler-Lopez, M, Petosa, C, Fukuzawa, M, Ravelli, R, Williams, J.G, Muller, C.W. | | Deposit date: | 2004-01-09 | | Release date: | 2004-03-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of an Activated Dictyostelium Stat in its DNA-Unbound Form
Mol.Cell, 13, 2004
|
|
2EQ7
 
 | |
1YTF
 
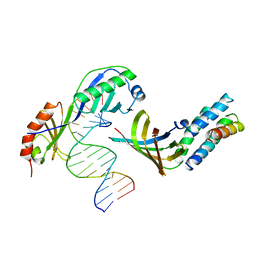 | | YEAST TFIIA/TBP/DNA COMPLEX | | Descriptor: | DNA (5'-D(*GP*TP*TP*TP*TP*AP*TP*AP*TP*AP*CP*AP*TP*AP*CP*A)-3'), DNA (5'-D(*TP*GP*TP*AP*TP*GP*TP*AP*TP*AP*TP*AP*AP*AP*AP*C)-3'), PROTEIN (TATA BINDING PROTEIN (TBP)), ... | | Authors: | Tan, S, Hunziker, Y, Sargent, D.F, Richmond, T.J. | | Deposit date: | 1996-04-05 | | Release date: | 1996-06-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of a yeast TFIIA/TBP/DNA complex.
Nature, 381, 1996
|
|
2ERE
 
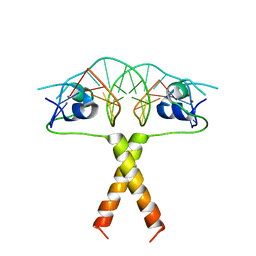 | |
1C7Q
 
 | | THE CRYSTAL STRUCTURE OF PHOSPHOGLUCOSE ISOMERASE/AUTOCRINE MOTILITY FACTOR/NEUROLEUKIN COMPLEXED WITH ITS CARBOHYDRATE PHOSPHATE INHIBITORS AND ITS SUBSTRATE RECOGNITION MECHANISM | | Descriptor: | N-BROMOACETYL-AMINOETHYL PHOSPHATE, PHOSPHOGLUCOSE ISOMERASE | | Authors: | Chou, C.-C, Meng, M, Sun, Y.-J, Hsiao, C.-D. | | Deposit date: | 2000-03-02 | | Release date: | 2000-09-13 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The crystal structure of phosphoglucose isomerase/autocrine motility factor/neuroleukin complexed with its carbohydrate phosphate inhibitors suggests its substrate/receptor recognition.
J.Biol.Chem., 275, 2000
|
|
1SMP
 
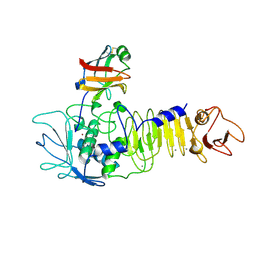 | | CRYSTAL STRUCTURE OF A COMPLEX BETWEEN SERRATIA MARCESCENS METALLO-PROTEASE AND AN INHIBITOR FROM ERWINIA CHRYSANTHEMI | | Descriptor: | CALCIUM ION, ERWINIA CHRYSANTHEMI INHIBITOR, SERRATIA METALLO PROTEINASE, ... | | Authors: | Baumann, U, Bauer, M, Letoffe, S, Delepelaire, P, Wandersman, C. | | Deposit date: | 1995-01-13 | | Release date: | 1996-04-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a complex between Serratia marcescens metallo-protease and an inhibitor from Erwinia chrysanthemi.
J.Mol.Biol., 248, 1995
|
|
2EO4
 
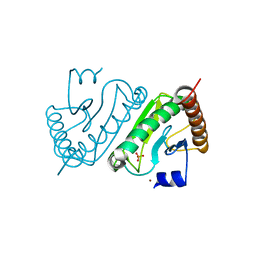 | |
1SPR
 
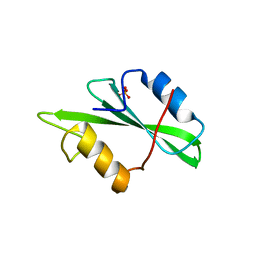 | |
1SNB
 
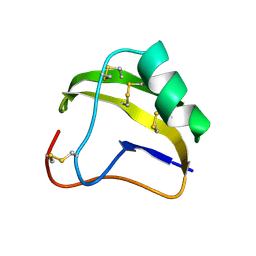 | | STRUCTURE OF SCORPION NEUROTOXIN BMK M8 | | Descriptor: | NEUROTOXIN BMK M8 | | Authors: | Wang, D.C, Zeng, Z.H, Li, H.M. | | Deposit date: | 1997-03-12 | | Release date: | 1997-05-15 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of an acidic neurotoxin from scorpion Buthus martensii Karsch at 1.85 A resolution.
J.Mol.Biol., 261, 1996
|
|
1YUK
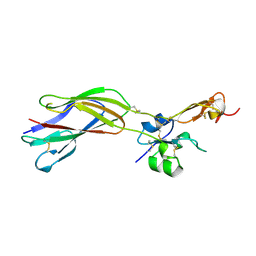 
 | | The crystal structure of the PSI/Hybrid domain/ I-EGF1 segment from the human integrin beta2 at 1.8 resolution | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, Integrin beta-2 A chain, Integrin beta-2 B chain | | Authors: | Shi, M, Sundramurthy, K, Liu, B, Tan, S.M, Law, S.K, Lescar, J. | | Deposit date: | 2005-02-14 | | Release date: | 2005-07-19 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Crystal Structure of the Plexin-Semaphorin-Integrin Domain/Hybrid Domain/I-EGF1 Segment from the Human Integrin {beta}2 Subunit at 1.8-A Resolution
J.Biol.Chem., 280, 2005
|
|
1YMK
 
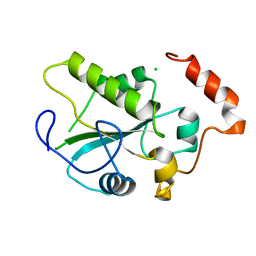 | | Crystal Structure of the CDC25B phosphatase catalytic domain in the apo form | | Descriptor: | CHLORIDE ION, M-phase inducer phosphatase 2 | | Authors: | Buhrman, G.K, Parker, B, Sohn, J, Rudolph, J, Mattos, C. | | Deposit date: | 2005-01-21 | | Release date: | 2005-04-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Mechanism of Oxidative Regulation of the Phosphatase Cdc25B via an Intramolecular Disulfide Bond
Biochemistry, 44, 2005
|
|
1SNK
 
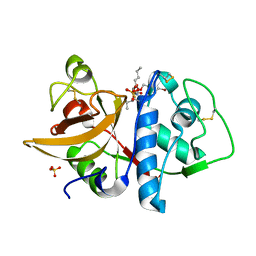 | | Cathepsin K complexed with carbamate derivatized norleucine aldehyde | | Descriptor: | Cathepsin K, N2-({[(4-BROMOPHENYL)METHYL]OXY}CARBONYL)-N1-[(1S)-1-FORMYLPENTYL]-L-LEUCINAMIDE, SULFATE ION | | Authors: | Boros, E.E, Deaton, D.N, Hassell, A.M, McFadyen, R.B, Miller, A.B, Miller, L.R, Shewchuk, L.M, Thompson, J.B, Willard Jr, D.H, Wright, L.L. | | Deposit date: | 2004-03-11 | | Release date: | 2004-06-22 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Exploration of the P(2)-P(3) SAR of aldehyde cathepsin K inhibitors
Bioorg.Med.Chem.Lett., 14, 2004
|
|
1YN5
 
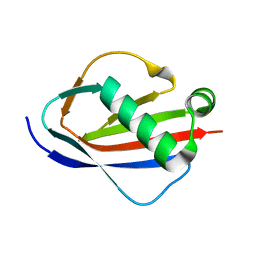 | | Crystal Structures of EAP Domains from Staphylococcus aureus Reveal an Unexpected Homology to Bacterial Superantigens | | Descriptor: | EapH2 | | Authors: | Geisbrecht, B.V, Hamaoka, B.Y, Perman, B, Zemla, A, Leahy, D.J. | | Deposit date: | 2005-01-23 | | Release date: | 2005-03-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Crystal Structures of EAP Domains from Staphylococcus aureus Reveal an Unexpected Homology to Bacterial Superantigens.
J.Biol.Chem., 280, 2005
|
|
2EO7
 
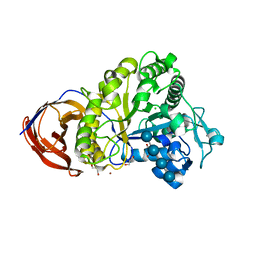 | | Crystal structure of Cel44A, GH family 44 endoglucanase from Clostridium thermocellum | | Descriptor: | CALCIUM ION, CHLORIDE ION, Endoglucanase, ... | | Authors: | Kitao, Y, Karita, S, Watanabe, N, Sakka, K, Tanaka, I. | | Deposit date: | 2007-03-29 | | Release date: | 2007-09-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of Cel44A, a glycoside hydrolase family 44 endoglucanase from Clostridium thermocellum.
J.Biol.Chem., 282, 2007
|
|
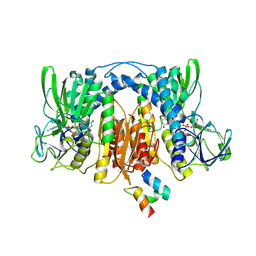
2EQ9
 
 | |
