7MWF
 
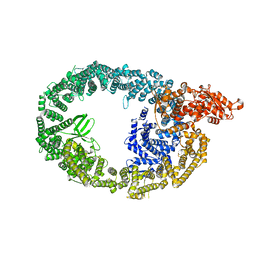 | | HUWE1 in map with focus on interface | | Descriptor: | E3 ubiquitin-protein ligase HUWE1 | | Authors: | Hunkeler, M, Fischer, E.S. | | Deposit date: | 2021-05-16 | | Release date: | 2021-07-28 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Solenoid architecture of HUWE1 contributes to ligase activity and substrate recognition.
Mol.Cell, 81, 2021
|
|
7O3D
 
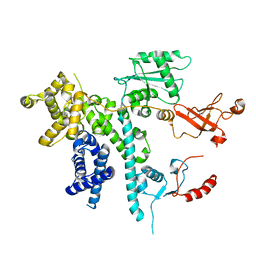 | | Cooperation between the intrinsically disordered and ordered regions of Spt6 regulates nucleosome and Pol II CTD binding, and nucleosome assembly | | Descriptor: | Transcription elongation factor SPT6 | | Authors: | Kasiliauskaite, A, Kubicek, K, Klumpler, T, Zanova, M, Zapletal, D, Novacek, J, Stefl, R. | | Deposit date: | 2021-04-01 | | Release date: | 2022-04-13 | | Last modified: | 2022-06-15 | | Method: | ELECTRON MICROSCOPY (3.71 Å) | | Cite: | Cooperation between intrinsically disordered and ordered regions of Spt6 regulates nucleosome and Pol II CTD binding, and nucleosome assembly.
Nucleic Acids Res., 50, 2022
|
|
7O6B
 
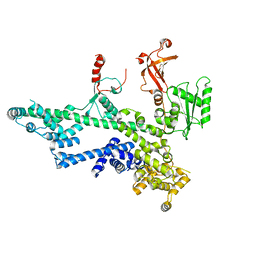 | | Cooperation between the intrinsically disordered and ordered regions of Spt6 regulates nucleosome and Pol II CTD binding, and nucleosome assembly | | Descriptor: | Transcription elongation factor SPT6 | | Authors: | Kasiliauskaite, A, Kubicek, K, Klumpler, T, Zanova, M, Zapletal, D, Novacek, J, Stefl, R. | | Deposit date: | 2021-04-09 | | Release date: | 2022-04-20 | | Last modified: | 2022-06-22 | | Method: | ELECTRON MICROSCOPY (3.88 Å) | | Cite: | Cooperation between intrinsically disordered and ordered regions of Spt6 regulates nucleosome and Pol II CTD binding, and nucleosome assembly.
Nucleic Acids Res., 50, 2022
|
|
7ODE
 
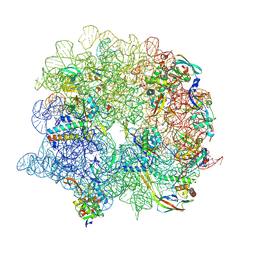 | | E. coli 50S ribosome LiCl core particle | | Descriptor: | 23S rRNA, 50S ribosomal protein L13, 50S ribosomal protein L14, ... | | Authors: | Larsson, D.S.D, Selmer, M. | | Deposit date: | 2021-04-29 | | Release date: | 2022-06-01 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (2.84 Å) | | Cite: | Structural Consequences of Deproteinating the 50S Ribosome.
Biomolecules, 12, 2022
|
|
8IOG
 
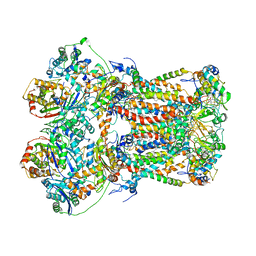 | |
6O9J
 
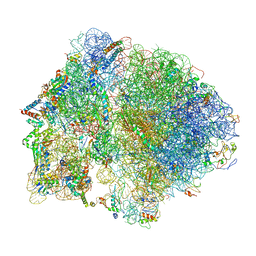 | | 70S Elongation Competent Ribosome | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Frank, J, Gonzalez Jr, R.L, Kaledhonkar, S, Fu, Z, Caban, K, Li, W, Chen, B, Sun, M. | | Deposit date: | 2019-03-14 | | Release date: | 2019-05-29 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Late steps in bacterial translation initiation visualized using time-resolved cryo-EM.
Nature, 570, 2019
|
|
9F18
 
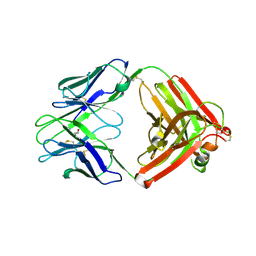 | |
9FJN
 | | Solution NMR structure of a peptide encompassing residues 2-19 of the human formin INF2 | | Descriptor: | Inverted formin-2 | | Authors: | Jimenez, M.A, Comas, L, Labat-de-Hoz, L, Correas, I, Alonso, M.A. | | Deposit date: | 2024-05-31 | | Release date: | 2024-09-11 | | Method: | SOLUTION NMR | | Cite: | Structure and function of the N-terminal extension of the formin INF2.
Cell Mol Life Sci, 79, 2022
|
|
9CL9
 
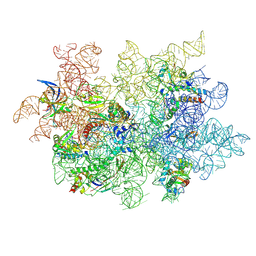 | | WT 12C IM fraction, B-b3 with RluB bound | | Descriptor: | 23S rRNA, Large ribosomal subunit protein bL20, Large ribosomal subunit protein bL21, ... | | Authors: | Lee, J, Sheng, K, Williamson, J.R. | | Deposit date: | 2024-07-10 | | Release date: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (5.04 Å) | | Cite: | 50S ribosome assembly intermediates at low temperature reveal bound RluB
To Be Published
|
|
9DLL
 | | NMR structures of small molecules bound to a model of an RNA CAG repeat expansion. | | Descriptor: | 4-carbamimidamidophenyl 4-carbamimidamidobenzoate, CAG 13-mer RNA (5'-R(*GP*AP*CP*AP*GP*CP*AP*GP*CP*UP*GP*UP*C)-3') | | Authors: | Chen, J.L, Taghavi, A, Disney, M.D, Fountain, M.A, Childs-Disney, J.L. | | Deposit date: | 2024-09-11 | | Release date: | 2024-09-25 | | Method: | SOLUTION NMR | | Cite: | NMR structures of small molecules bound to a model of an RNA CAG repeat expansion.
Nucleic Acids Res., 2024
|
|
9FJW
 
 | | Solution NMR structure of a peptide encompassing residues 2-36 of the human formin INF2 | | Descriptor: | Inverted formin-2 | | Authors: | Jimenez, M.A, Comas, L, Labat-de-Hoz, L, Correas, I, Alonso, M.A. | | Deposit date: | 2024-05-31 | | Release date: | 2024-09-11 | | Method: | SOLUTION NMR | | Cite: | Structure and function of the N-terminal extension of the formin INF2.
Cell Mol Life Sci, 79, 2022
|
|
9EP9
 
 | | NMR solution structure of lipid transfer protei Sola l7 from tomato seeds | | Descriptor: | Non-specific lipid-transfer protein | | Authors: | Parron-Ballesteros, J, Mantin-Pedraz, L, G.Gordo, R, Mayorga, C, Villaba, M, Batanero, E, Pantoja-Uceda, D, Turnay, J. | | Deposit date: | 2024-03-18 | | Release date: | 2024-09-04 | | Last modified: | 2024-10-09 | | Method: | SOLUTION NMR | | Cite: | Long-chain fatty acids block allergic reaction against lipid transfer protein Sola l 7 from tomato seeds.
Protein Sci., 33, 2024
|
|
9F1I
 
 | | Crystal structure of a first-in-class antibody for alpha-1,6-fucosylated prostate-specific antigen, target bound | | Descriptor: | 1,2-ETHANEDIOL, 2-[2-(2-azanylethoxy)ethoxy]ethanoic acid, Heavy chain rabbit fab, ... | | Authors: | Halldorsson, S. | | Deposit date: | 2024-04-19 | | Release date: | 2024-07-31 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Development of a first-in-class antibody and a specific assay for alpha-1,6-fucosylated prostate-specific antigen.
Sci Rep, 14, 2024
|
|
9BH5
 | | High-resolution C. elegans 80S ribosome structure - class 1 | | Descriptor: | 18S rRNA, 28S rRNA, 4-{(2R)-2-[(1S,3S,5S)-3,5-dimethyl-2-oxocyclohexyl]-2-hydroxyethyl}piperidine-2,6-dione, ... | | Authors: | Sehgal, E, Serrao, V.H.B, Arribere, J. | | Deposit date: | 2024-04-19 | | Release date: | 2024-09-04 | | Last modified: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (2.63 Å) | | Cite: | High-Resolution Reconstruction of a C. elegans Ribosome Sheds Light on Evolutionary Dynamics and Tissue Specificity.
Rna, 2024
|
|
9F3D
 
 | |
9F3E
 
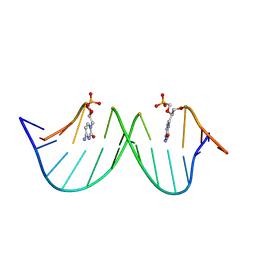 | |
9EVX
 | | cryoEM structure of Photosystem II averaged across S2-S3 states at 1.71 Angstrom resolution | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | Authors: | Hussein, R, Graca, A, Zouni, A, Messinger, J, Schroder, W.P. | | Deposit date: | 2024-04-02 | | Release date: | 2024-06-12 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (1.71 Å) | | Cite: | Cryo-electron microscopy reveals hydrogen positions and water networks in photosystem II.
Science, 384, 2024
|
|
1A11
 
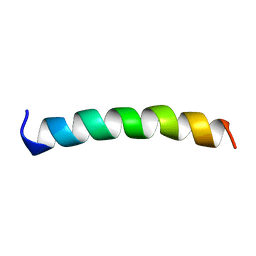 | | NMR STRUCTURE OF MEMBRANE SPANNING SEGMENT 2 OF THE ACETYLCHOLINE RECEPTOR IN DPC MICELLES, 10 STRUCTURES | | Descriptor: | ACETYLCHOLINE RECEPTOR M2 | | Authors: | Gesell, J.J, Sun, W, Montal, M, Opella, S.J. | | Deposit date: | 1997-12-19 | | Release date: | 1998-04-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structures of the M2 channel-lining segments from nicotinic acetylcholine and NMDA receptors by NMR spectroscopy.
Nat.Struct.Biol., 6, 1999
|
|
7UX3
 
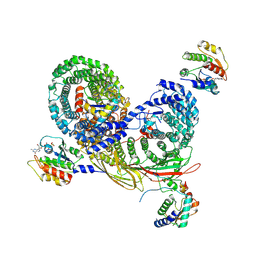 | | Asymmetric unit of AP-1, Arf1, Nef lattice on MHC-I lipopeptide incorporated narrow membrane tubes | | Descriptor: | ADP-ribosylation factor 1, AP-1 complex subunit beta-1, AP-1 complex subunit gamma-1, ... | | Authors: | Hooy, R.M, Hurley, J.H. | | Deposit date: | 2022-05-04 | | Release date: | 2023-05-10 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (9.6 Å) | | Cite: | Self-assembly and structure of a clathrin-independent AP-1:Arf1 tubular membrane coat.
Sci Adv, 8, 2022
|
|
1AXP
 
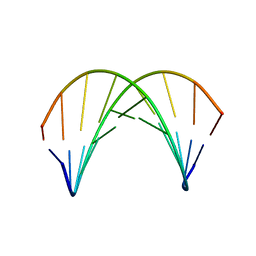 | | DNA DUPLEX CONTAINING A PURINE-RICH STRAND, NMR, 6 STRUCTURES | | Descriptor: | DNA (D(GAAGAGAAGC)(DOT)D(GCTTCTCTTC)) | | Authors: | Gyi, J.I, Lane, A.N, Conn, G.L, Brown, T. | | Deposit date: | 1997-10-17 | | Release date: | 1998-04-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structures of DNA.RNA hybrids with purine-rich and pyrimidine-rich strands: comparison with the homologous DNA and RNA duplexes.
Biochemistry, 37, 1998
|
|
1BUQ
 | | SOLUTION STRUCTURE OF DELTA-5-3-KETOSTEROID ISOMERASE COMPLEXED WITH THE STEROID 19-NORTESTOSTERONE-HEMISUCCINATE | | Descriptor: | PROTEIN (3-KETOSTEROID ISOMERASE-19-NORTESTOSTERONE-HEMISUCCINATE), SUCCINIC ACID MONO-(13-METHYL-3-OXO-2,3,6,7,8,9,10,11,12,13,14,15,16,17-TETRADECAHYDRO-1H-CYCLOPENTA[A]PHENANTHREN-17-YL) ESTER | | Authors: | Massiah, M.A, Abeygunawardana, C, Gittis, A.G, Mildvan, A.S. | | Deposit date: | 1998-09-04 | | Release date: | 1999-01-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Delta 5-3-ketosteroid isomerase complexed with the steroid 19-nortestosterone hemisuccinate.
Biochemistry, 37, 1998
|
|
8HGX
 
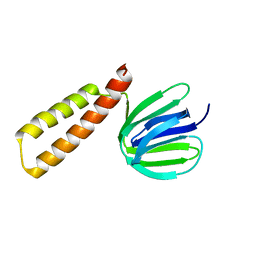 | |
8T1O
 
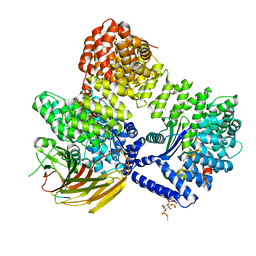 | | AP2 bound to MSP2N2 nanodisc with Tgn38 cargo peptide; composite map | | Descriptor: | AP-2 complex subunit alpha-2, AP-2 complex subunit beta, AP-2 complex subunit mu, ... | | Authors: | Baker, R.W, Cannon, K.S, Reta, S. | | Deposit date: | 2023-06-02 | | Release date: | 2023-07-12 | | Last modified: | 2024-06-19 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Lipid nanodiscs as a template for high-resolution cryo-EM structures of peripheral membrane proteins.
J.Struct.Biol., 215, 2023
|
|
8SW7
 
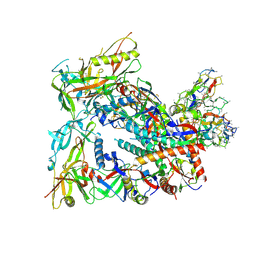 | | BG505 Boost2 SOSIP.664 in complex with NHP polyclonal antibody FP1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, BG505 Boost 2 gp120, BG505 Boost 2 gp41, ... | | Authors: | Pratap, P.P, Antansijevic, A, Ozorowski, G, Ward, A.B. | | Deposit date: | 2023-05-17 | | Release date: | 2023-07-26 | | Method: | ELECTRON MICROSCOPY (3.37 Å) | | Cite: | Focusing antibody responses to the fusion peptide in rhesus macaques.
Biorxiv, 2023
|
|
7TJQ
 
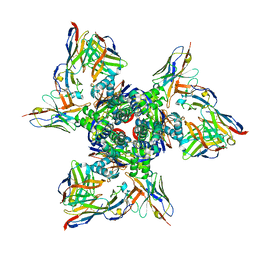 | | SAN27-14 bound to a antigenic site V on prefusion-stabilized hMPV F | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Fusion glycoprotein F0, MPE8 Fab heavy chain, ... | | Authors: | Hsieh, C.-L, McLellan, J.S, Rush, S.A. | | Deposit date: | 2022-01-16 | | Release date: | 2022-09-14 | | Last modified: | 2022-11-23 | | Method: | ELECTRON MICROSCOPY (3.13 Å) | | Cite: | Characterization of prefusion-F-specific antibodies elicited by natural infection with human metapneumovirus.
Cell Rep, 40, 2022
|
|
