3L22
 
 | |
3L2J
 
 | |
3KF0
 
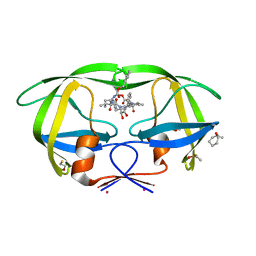 | | HIV Protease with fragment 4D9 bound | | Descriptor: | (1S,2S)-2-methylcyclohexanol, BETA-MERCAPTOETHANOL, DIMETHYL SULFOXIDE, ... | | Authors: | Stout, C.D, Perryman, A.L. | | Deposit date: | 2009-10-27 | | Release date: | 2010-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Fragment-based screen against HIV protease.
Chem.Biol.Drug Des., 75, 2010
|
|
3KFA
 
 | |
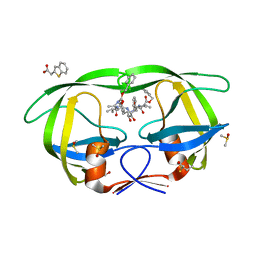
3KFS
 | |
3KI4
 
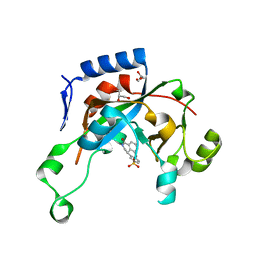 | | Catalytic fragment of Cholix toxin from Vibrio Cholerae in complex with inhibitor GP-P | | Descriptor: | (11bR)-3-oxo-1,2,3,11b-tetrahydrochromeno[4,3,2-de]isoquinoline-10-sulfonic acid, Cholix toxin, GLYCEROL | | Authors: | Jorgensen, R, Edwards, P.R, Merrill, A.R. | | Deposit date: | 2009-10-31 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure function analysis of soluble inhibitors of cholix toxin from Vibrio cholerae
To be Published
|
|
3KIZ
 
 | |
8ANY
 
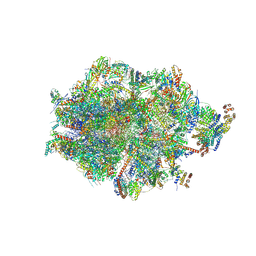 | | Human mitochondrial ribosome in complex with LRPPRC, SLIRP, A-site, P-site, E-site tRNAs and mRNA | | Descriptor: | 1,4-DIAMINOBUTANE, 12S mitochondrial rRNA, 16S mitochondrial rRNA, ... | | Authors: | Singh, V, Itoh, Y, Amunts, A. | | Deposit date: | 2022-08-06 | | Release date: | 2023-08-16 | | Last modified: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (2.85 Å) | | Cite: | Structural basis of LRPPRC-SLIRP-dependent translation by the mitoribosome.
Nat.Struct.Mol.Biol., 2024
|
|
3KDU
 
 | |
5J8A
 
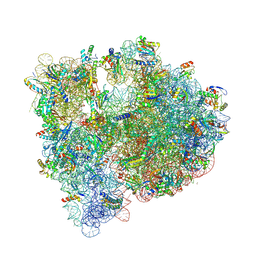 | | Structure of the E coli 70S ribosome with the U1052G mutation in 16S rRNA bound to tigecycline | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,2-ETHANEDIOL, 1,4-DIAMINOBUTANE, ... | | Authors: | Cocozaki, A, Ferguson, A. | | Deposit date: | 2016-04-07 | | Release date: | 2016-07-06 | | Last modified: | 2016-08-03 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Resistance mutations generate divergent antibiotic susceptibility profiles against translation inhibitors.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
3KEI
 
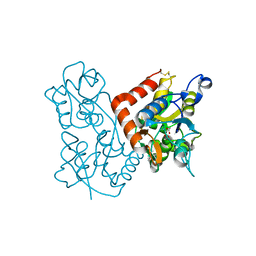 | |
5JFN
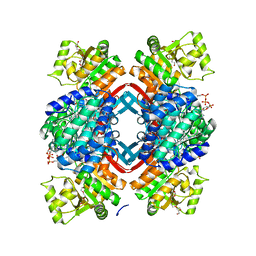 
 | | Crystal structure of Rhodopseudomonas palustris propionaldehyde dehydrogenase with bound CoA and acylated Cys330 | | Descriptor: | Aldehyde dehydrogenase, COENZYME A, PENTAETHYLENE GLYCOL, ... | | Authors: | Zarzycki, J, Sutter, M, Kerfeld, C.A. | | Deposit date: | 2016-04-19 | | Release date: | 2017-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | In Vitro Characterization and Concerted Function of Three Core Enzymes of a Glycyl Radical Enzyme - Associated Bacterial Microcompartment.
Sci Rep, 7, 2017
|
|
3KLB
 
 | |
3KF5
 
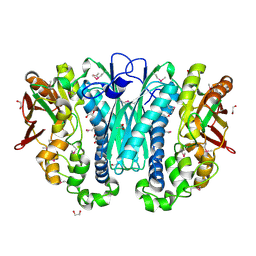
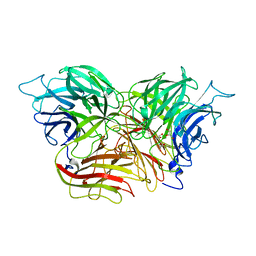 | | Structure of invertase from Schwanniomyces occidentalis | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Invertase | | Authors: | Sanz-Aparicio, J, Polo, A. | | Deposit date: | 2009-10-27 | | Release date: | 2010-02-23 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural and kinetic analysis of Schwanniomyces occidentalis invertase reveals a new oligomerization pattern and the role of its supplementary domain in substrate binding
J.Biol.Chem., 285, 2010
|
|
3KBZ
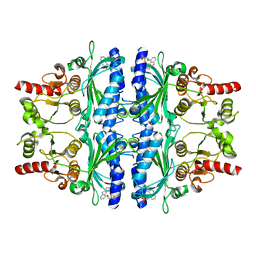 
 | | Crystal structure of human liver FBPase in complex with tricyclic inhibitor 6 | | Descriptor: | Fructose-1,6-bisphosphatase 1, {[(2-amino-8H-indeno[1,2-d][1,3]thiazol-4-yl)oxy]methyl}phosphonic acid | | Authors: | Takahashi, M, Sone, J, Hanzawa, H. | | Deposit date: | 2009-10-20 | | Release date: | 2010-02-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structure-based drug design of tricyclic 8H-indeno[1,2-d][1,3]thiazoles as potent FBPase inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
3KM5
 
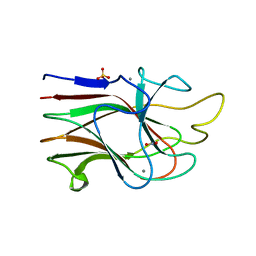 | | Crystal Structure Analysis of the K2 Cleaved Adhesin Domain of Lys-gingipain (Kgp) | | Descriptor: | CALCIUM ION, GLYCEROL, Lysine specific cysteine protease, ... | | Authors: | Li, N, Collyer, C.A, Hunter, N. | | Deposit date: | 2009-11-09 | | Release date: | 2010-03-31 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure determination and analysis of a haemolytic gingipain adhesin domain from Porphyromonas gingivalis
Mol.Microbiol., 76, 2010
|
|
3KMG
 
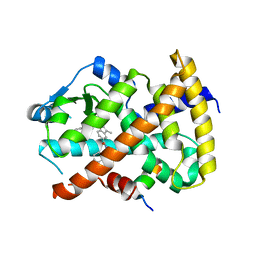 | |
8AEL
 
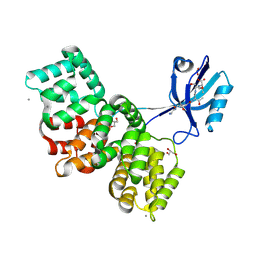 | | SYNJ2BP complex with a synthetic Vangl2 peptide (3mer). | | Descriptor: | CALCIUM ION, GLY-GLY-GLY-THR-SER-VAL, GLYCEROL, ... | | Authors: | Carrasco, K, Cousido Siah, A, Gogl, G, Betzi, S, McEwen, A, Kostmann, C, Trave, G. | | Deposit date: | 2022-07-13 | | Release date: | 2023-08-16 | | Last modified: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | SYNJ2BP PDZ domain in complex with a synthetic Vangl2 peptide.
To Be Published
|
|
3KN7
 
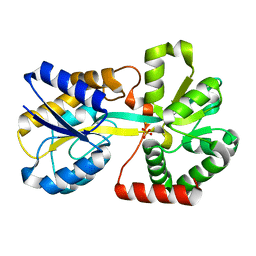 | |
3KDW
 
 | |
3KE5
 
 | |
3KNC
 
 | |
3KNP
 
 | |
3KNZ
 
 | |
3KEZ
 
 | |
