1M8Y
 
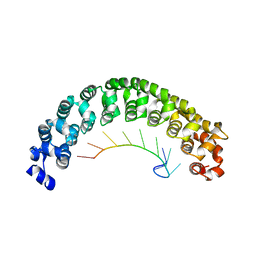 | | CRYSTAL STRUCTURE OF THE PUMILIO-HOMOLOGY DOMAIN FROM HUMAN PUMILIO1 IN COMPLEX WITH NRE2-10 RNA | | Descriptor: | 5'-R(P*AP*UP*UP*GP*UP*AP*CP*AP*UP*A)-3', Pumilio 1 | | Authors: | Wang, X, McLachlan, J, Zamore, P.D, Hall, T.M.T. | | Deposit date: | 2002-07-26 | | Release date: | 2002-09-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | MODULAR RECOGNITION OF RNA BY A HUMAN PUMILIO-HOMOLOGY DOMAIN
CELL(CAMBRIDGE,MASS.), 110, 2002
|
|
8IAJ
 
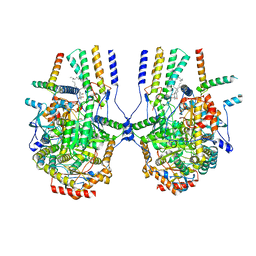 | | Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A) complex | | Descriptor: | N-[(2S,3R,4E)-1,3-dihydroxyoctadec-4-en-2-yl]tetracosanamide, PYRIDOXAL-5'-PHOSPHATE, Protein ORM2, ... | | Authors: | Xie, T, Gong, X. | | Deposit date: | 2023-02-08 | | Release date: | 2024-02-14 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Collaborative regulation of yeast SPT-Orm2 complex by phosphorylation and ceramide.
Cell Rep, 43, 2024
|
|
1M9K
 
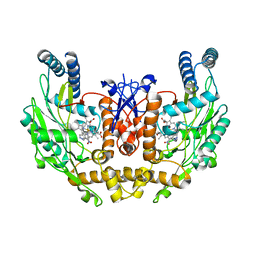 | | Human Endothelial Nitric Oxide Synthase with 7-Nitroindazole Bound | | Descriptor: | 7-NITROINDAZOLE, PROTOPORPHYRIN IX CONTAINING FE, ZINC ION, ... | | Authors: | Rosenfeld, R.J, Garcin, E.D, Panda, K, Andersson, G, Aberg, A, Wallace, A.V, Stuehr, D.J, Tainer, J.A, Getzoff, E.D. | | Deposit date: | 2002-07-29 | | Release date: | 2002-08-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Conformational Changes in Nitric Oxide Synthases Induced by Chlorzoxazone and Nitroindazoles: Crystallographic and Computational Analyses of Inhibitor Potency
Biochemistry, 41, 2002
|
|
5F5S
 
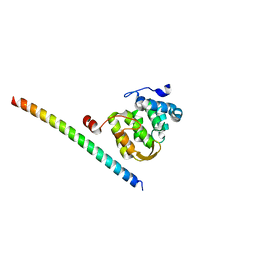 | |
5F5Z
 
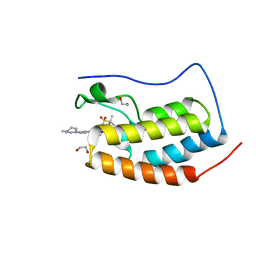 | | Crystal structure of the first bromodomain of human BRD4 in complex with MA2-014 | | Descriptor: | 1,2-ETHANEDIOL, 2-methyl-~{N}-[3-[[5-methyl-2-[[4-(4-methylpiperazin-1-yl)phenyl]amino]pyrimidin-4-yl]amino]phenyl]propane-2-sulfonamide, Bromodomain-containing protein 4 | | Authors: | Ember, S.W, Zhu, J.-Y, Schonbrunn, E. | | Deposit date: | 2015-12-04 | | Release date: | 2017-04-05 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Potent Dual BET Bromodomain-Kinase Inhibitors as Value-Added Multitargeted Chemical Probes and Cancer Therapeutics.
Mol. Cancer Ther., 16, 2017
|
|
8I0B
 
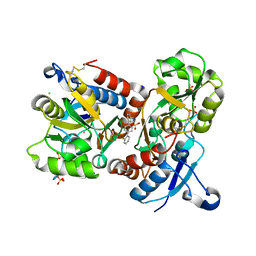 | | The crystal structure of human glutamate receptor 2 in complex with LT-102 | | Descriptor: | 8-[4-(2-fluorophenyl)phenyl]-3,4-dihydro-1,2$l^{6},3-benzoxathiazine 2,2-dioxide, CHLORIDE ION, Glutamate receptor 2,Isoform Flip of Glutamate receptor 2, ... | | Authors: | Qi, X.Y, Wu, C.Y. | | Deposit date: | 2023-01-10 | | Release date: | 2024-02-07 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Structure of the human GluA2 LBD in complex with LT-102
To Be Published
|
|
5F6M
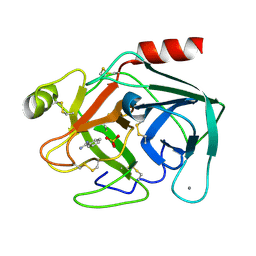 
 | | Isotropic Trypsin Model for Comparison of Diffuse Scattering | | Descriptor: | BENZAMIDINE, CALCIUM ION, Cationic trypsin, ... | | Authors: | Van Benschoten, A.H, Wall, M.E, Fraser, J.S. | | Deposit date: | 2015-12-06 | | Release date: | 2016-01-06 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Measuring and modeling diffuse scattering in protein X-ray crystallography.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
5EWI
 
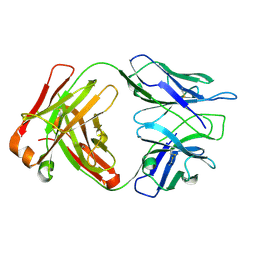 | |
1Z3V
 
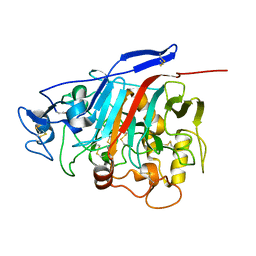 | | Structure of Phanerochaete chrysosporium cellobiohydrolase Cel7D (CBH58) in complex with lactose | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-galactopyranose-(1-4)-beta-D-glucopyranose, cellulase | | Authors: | Ubhayasekera, W, Munoz, I.G, Stahlberg, J, Mowbray, S.L. | | Deposit date: | 2005-03-14 | | Release date: | 2005-04-26 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | Structures of Phanerochaete chrysosporium Cel7D in complex with product and inhibitors
Febs J., 272, 2005
|
|
5F79
 
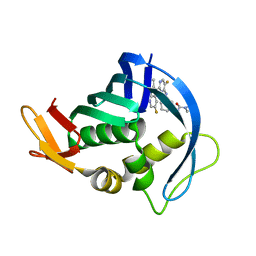 | | Influenza PB2 bound to an azaindole inhibitor | | Descriptor: | N-[(1R,3S)-3-({5-fluoro-2-[5-fluoro-2-(hydroxymethyl)-1H-pyrrolo[2,3-b]pyridin-3-yl]pyrimidin-4-yl}amino)cyclohexyl]pyrrolidine-1-carboxamide, Polymerase basic protein 2 | | Authors: | Jacobs, M.D. | | Deposit date: | 2015-12-07 | | Release date: | 2016-12-07 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Novel 2-Substituted 7-Azaindole and 7-Azaindazole Analogues as Potential Antiviral Agents for the Treatment of Influenza.
ACS Med Chem Lett, 8, 2017
|
|
8IA1
 
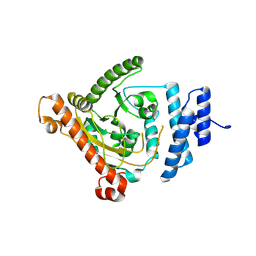 | |
8IWB
 
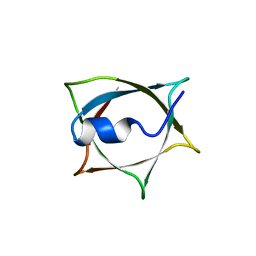 | | Crystal structure of Q9PR55 at pH 7.5 | | Descriptor: | Uncharacterized protein UU089.1 | | Authors: | Hsu, M.F, Ko, T.P, Huang, K.F, Chen, Y.R, Huang, J.S, Hsu, S.T.D. | | Deposit date: | 2023-03-29 | | Release date: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Structure, dynamics, and stability of the smallest and most complex 7 1 protein knot.
J.Biol.Chem., 300, 2023
|
|
8IAM
 
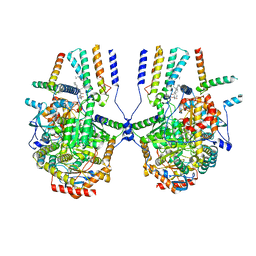 | | Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3D) complex | | Descriptor: | Chimera of Long chain base biosynthesis protein 1 and Serine palmitoyltransferase 1, N-[(2S,3R,4E)-1,3-dihydroxyoctadec-4-en-2-yl]tetracosanamide, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Xie, T, Gong, X. | | Deposit date: | 2023-02-08 | | Release date: | 2024-02-14 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Collaborative regulation of yeast SPT-Orm2 complex by phosphorylation and ceramide.
Cell Rep, 43, 2024
|
|
8I90
 
 | |
8I9M
 
 | | The RAGE and HMGB1 complex | | Descriptor: | Advanced glycosylation end product-specific receptor, High mobility group protein B1 | | Authors: | Han, C.W, Kim, H.J. | | Deposit date: | 2023-02-07 | | Release date: | 2024-02-14 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (5.19 Å) | | Cite: | The RAGE and HMGB1 complex
To Be Published
|
|
8I83
 
 | |
8IDX
 
 | | Structure of p205 HIN | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, Interferon-activable protein 205-B, ... | | Authors: | Li, Y.L, Jin, T.C. | | Deposit date: | 2023-02-14 | | Release date: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of p205 HIN
to Be published
|
|
5EYS
 
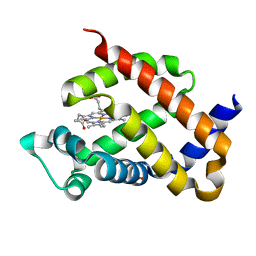 | | Crystal structure of murine neuroglobin mutant F106W at ambient pressure | | Descriptor: | Neuroglobin, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION | | Authors: | Colloc'h, N, Girard, E, Vallone, B, Prange, T. | | Deposit date: | 2015-11-25 | | Release date: | 2016-10-19 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Determinants of neuroglobin plasticity highlighted by joint coarse-grained simulations and high pressure crystallography.
Sci Rep, 7, 2017
|
|
1MLZ
 
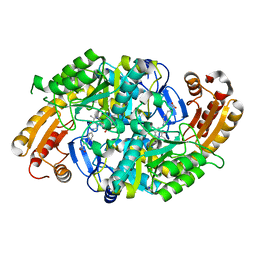 | | Crystal Structure of 7,8-Diaminopelargonic Acid Synthase in complex with the trans-isomer of amiclenomycin. | | Descriptor: | 7,8-diamino-pelargonic acid aminotransferase, PYRIDOXAL-5'-PHOSPHATE, SODIUM ION, ... | | Authors: | Sandmark, J, Mann, S, Marquet, A, Schneider, G. | | Deposit date: | 2002-09-02 | | Release date: | 2002-12-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural basis for the inhibition of the biosynthesis of biotin by the antibiotic amiclenomycin
J.Biol.Chem., 277, 2002
|
|
5EZ0
 
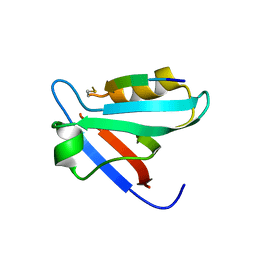 | | CRYSTAL STRUCTURE OF THE PTPN4 PDZ DOMAIN COMPLEXED WITH THE PDZ BINDING MOTIF OF THE MITOGEN ACTIVATED PROTEIN KINASE P38GAMMA. | | Descriptor: | Mitogen-activated protein kinase 12, SULFATE ION, Tyrosine-protein phosphatase non-receptor type 4 | | Authors: | Maisonneuve, P, Vaney, M.C, Caillet-Saguy, C, Lafon, M, Delepierre, M, Cordier, F, Wolff, N. | | Deposit date: | 2015-11-26 | | Release date: | 2016-06-08 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Molecular Basis of the Interaction of the Human Protein Tyrosine Phosphatase Non-receptor Type 4 (PTPN4) with the Mitogen-activated Protein Kinase p38 gamma.
J.Biol.Chem., 291, 2016
|
|
8IVI
 
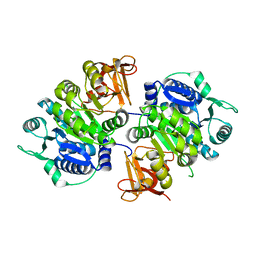 | | crystal structure of a medium-long chain fatty acyl-CoA ligase | | Descriptor: | Medium/long-chain-fatty-acid--CoA ligase FadD8 | | Authors: | Li, S. | | Deposit date: | 2023-03-27 | | Release date: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Structural study of medium-long chain fatty acyl-CoA ligase FadD8 from Mycobacterium tuberculosis.
Biochem.Biophys.Res.Commun., 672, 2023
|
|
8HZT
 
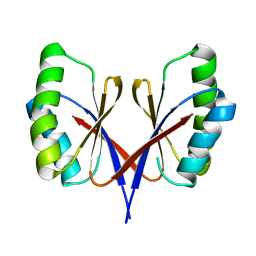 | |
5EX5
 
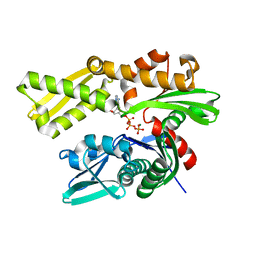 | | Crystal structure of human GRP78 (70kDa heat shock protein 5 / BIP) ATPase domain in complex with 7-deaza-ADP and inorganic phosphate | | Descriptor: | 7-deazaadenosine-5'-diphosphate, 78 kDa glucose-regulated protein, MAGNESIUM ION, ... | | Authors: | Hughes, S.J, Antoshchenko, T, Song, J.H, Pizarro, J, Park, H.W. | | Deposit date: | 2015-11-23 | | Release date: | 2016-06-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Probing the ATP Site of GRP78 with Nucleotide Triphosphate Analogs.
Plos One, 11, 2016
|
|
8I94
 
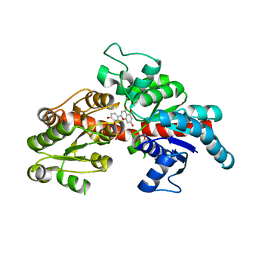 | | Structure of flavone 4'-O-glucoside 7-O-glucosyltransferase from Nemophila menziesii, complex with luteolin | | Descriptor: | 2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-4H-chromen-4-one, Glycosyltransferase, SULFATE ION | | Authors: | Murayama, K, Kato-Murayama, M, Shirouzu, M. | | Deposit date: | 2023-02-06 | | Release date: | 2024-02-14 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Molecular basis of ligand recognition specificity of flavone glucosyltransferases in Nemophila menziesii.
Arch.Biochem.Biophys., 753, 2024
|
|
5EXM
 
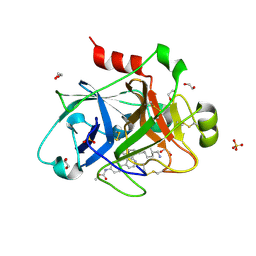 | |
