1KDF
 
 | | NORTH-ATLANTIC OCEAN POUT ANTIFREEZE PROTEIN TYPE III ISOFORM HPLC12 MUTANT, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | ANTIFREEZE PROTEIN | | Authors: | Sonnichsen, F.D, Deluca, C.I, Davies, P.L, Sykes, B.D. | | Deposit date: | 1996-07-08 | | Release date: | 1997-04-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Refined solution structure of type III antifreeze protein: hydrophobic groups may be involved in the energetics of the protein-ice interaction.
Structure, 4, 1996
|
|
1KUL
 
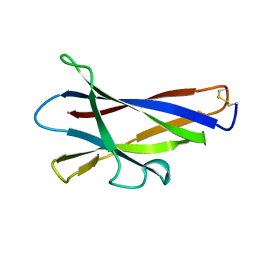 | | GLUCOAMYLASE, GRANULAR STARCH-BINDING DOMAIN, NMR, 5 STRUCTURES | | Descriptor: | GLUCOAMYLASE | | Authors: | Sorimachi, K, Jacks, A.J, Le Gal-Coeffet, M.-F, Williamson, G, Archer, D.B, Williamson, M.P. | | Deposit date: | 1996-01-12 | | Release date: | 1996-07-11 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the granular starch binding domain of glucoamylase from Aspergillus niger by nuclear magnetic resonance spectroscopy.
J.Mol.Biol., 259, 1996
|
|
1KUM
 
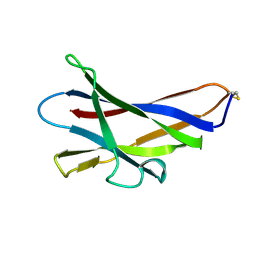 | | GLUCOAMYLASE, GRANULAR STARCH-BINDING DOMAIN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | GLUCOAMYLASE | | Authors: | Sorimachi, K, Jacks, A.J, Le Gal-Coeffet, M.-F, Williamson, G, Archer, D.B, Williamson, M.P. | | Deposit date: | 1996-01-12 | | Release date: | 1996-07-11 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the granular starch binding domain of glucoamylase from Aspergillus niger by nuclear magnetic resonance spectroscopy.
J.Mol.Biol., 259, 1996
|
|
1L3Y
 
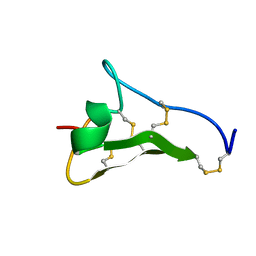 | |
2PPH
 
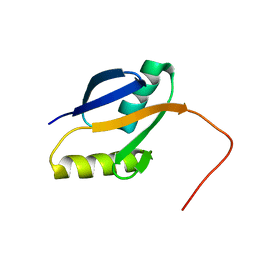 | | solution structure of human MEKK3 PB1 domain | | Descriptor: | Mitogen-activated protein kinase kinase kinase 3 | | Authors: | Hu, Q, Zhang, J, Wu, J, Shi, Y. | | Deposit date: | 2007-04-30 | | Release date: | 2007-05-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Insight into the Binding Properties of MEKK3 PB1 to MEK5 PB1 from Its Solution Structure.
Biochemistry, 46, 2007
|
|
1JI9
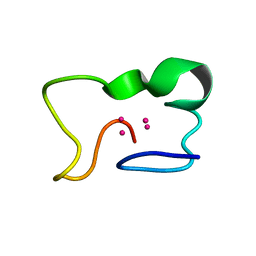 
 | |
1ZU2
 
 | |
1IGL
 
 | | SOLUTION STRUCTURE OF HUMAN INSULIN-LIKE GROWTH FACTOR II RELATIONSHIP TO RECEPTOR AND BINDING PROTEIN INTERACTIONS | | Descriptor: | INSULIN-LIKE GROWTH FACTOR II | | Authors: | Torres, A.M, Forbes, B.E, Aplin, S.E, Wallace, J.C, Francis, G.L, Norton, R.S. | | Deposit date: | 1994-12-29 | | Release date: | 1995-02-14 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of human insulin-like growth factor II. Relationship to receptor and binding protein interactions.
J.Mol.Biol., 248, 1995
|
|
2AFJ
 
 | | SPRY domain-containing SOCS box protein 2 (SSB-2) | | Descriptor: | gene rich cluster, C9 gene | | Authors: | Masters, S.L, Yao, S, Willson, T.A, Zhang, J.G, Palmer, K.R, Smith, B.J, Babon, J.J, Nicola, N.A, Norton, R.S, Nicholson, S.E. | | Deposit date: | 2005-07-26 | | Release date: | 2006-01-03 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The SPRY domain of SSB-2 adopts a novel fold that presents conserved Par-4-binding residues
Nat.Struct.Mol.Biol., 13, 2006
|
|
1ISK
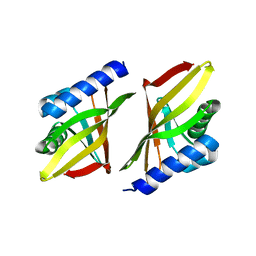 
 | | 3-OXO-DELTA5-STEROID ISOMERASE, NMR, 20 STRUCTURES | | Descriptor: | 3-OXO-DELTA5-STEROID ISOMERASE | | Authors: | Wu, Z.R, Ebrahimian, S, Zawrotny, M.E, Thornburg, L.D, Perez-Alvarado, G.C, Brothers, P, Pollack, R.M, Summers, M.F. | | Deposit date: | 1997-03-12 | | Release date: | 1997-11-12 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of 3-oxo-delta5-steroid isomerase.
Science, 276, 1997
|
|
2P0X
 
 | | solution structure of a non-biological ATP-binding protein | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, ZINC ION, abiotic ATP-binding, ... | | Authors: | Mansy, S.S, Szostak, J.W, Chaput, J.C. | | Deposit date: | 2007-03-01 | | Release date: | 2007-08-07 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure and Evolutionary Analysis of a Non-biological ATP-binding Protein
J.Mol.Biol., 371, 2007
|
|
1IDZ
 
 | |
1JY6
 | |
1JSA
 
 | | MYRISTOYLATED RECOVERIN WITH TWO CALCIUMS BOUND, NMR, 24 STRUCTURES | | Descriptor: | CALCIUM ION, MYRISTIC ACID, RECOVERIN | | Authors: | Ames, J.B, Ishima, R, Tanaka, T, Gordon, J.I, Stryer, L, Ikura, M. | | Deposit date: | 1997-06-04 | | Release date: | 1997-10-15 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Molecular mechanics of calcium-myristoyl switches.
Nature, 389, 1997
|
|
1JTB
 
 | | LIPID TRANSFER PROTEIN COMPLEXED WITH PALMITOYL COENZYME A, NMR, 16 STRUCTURES | | Descriptor: | COENZYME A, LIPID TRANSFER PROTEIN, PALMITIC ACID | | Authors: | Lerche, M.H, Kragelund, B.B, Bech, L.M, Poulsen, F.M. | | Deposit date: | 1996-12-03 | | Release date: | 1997-07-07 | | Last modified: | 2024-06-05 | | Method: | SOLUTION NMR | | Cite: | Barley lipid-transfer protein complexed with palmitoyl CoA: the structure reveals a hydrophobic binding site that can expand to fit both large and small lipid-like ligands.
Structure, 5, 1997
|
|
2NAZ
 
 | | The solution NMR structure of the C-terminal effector domain of BfmR from Acinetobacter baumannii | | Descriptor: | Transcriptional regulatory protein RstA | | Authors: | Olson, A.L, Thompson, R.J, Cavanagh, J, Feldmann, E.A, Bobay, B.G. | | Deposit date: | 2016-01-15 | | Release date: | 2017-01-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Structure of the Biofilm-controlling Response Regulator BfmR from Acinetobacter baumannii Reveals Details of Its DNA-binding Mechanism.
J.Mol.Biol., 430, 2018
|
|
2N6L
 
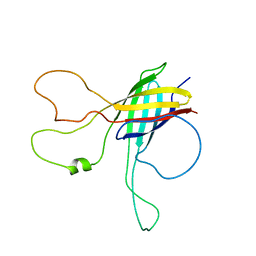 | | Solution NMR structure of Outer Membrane Protein G from Pseudomonas aeruginosa | | Descriptor: | Outer membrane protein OprG | | Authors: | Kucharska, I, Seelheim, P, Edrington, T.C, Liang, B, Tamm, L.K. | | Deposit date: | 2015-08-25 | | Release date: | 2015-12-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | OprG Harnesses the Dynamics of its Extracellular Loops to Transport Small Amino Acids across the Outer Membrane of Pseudomonas aeruginosa.
Structure, 23, 2015
|
|
1M5Z
 
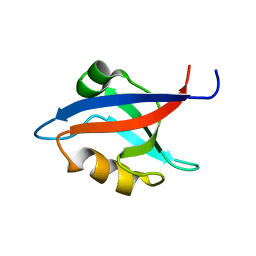 | | The PDZ7 of Glutamate Receptor Interacting Protein Binds to its Target via a Novel Hydrophobic Surface Area | | Descriptor: | AMPA receptor interacting protein | | Authors: | Feng, W, Fan, J, Jiang, M, Shi, Y, Zhang, M. | | Deposit date: | 2002-07-11 | | Release date: | 2002-11-06 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The PDZ7 of Glutamate Receptor Interacting Protein Binds to its Target via a Novel Hydrophobic Surface Area
J.Biol.Chem., 277, 2002
|
|
1J7R
 
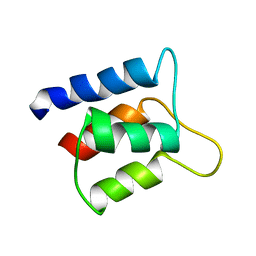 | | Solution structure and backbone dynamics of the defunct EF-hand domain of Calcium Vector Protein | | Descriptor: | Calcium Vector Protein | | Authors: | Theret, I, Baladi, S, Cox, J.A, Gallay, J, Sakamoto, H, Craescu, C.T. | | Deposit date: | 2001-05-18 | | Release date: | 2001-06-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure and backbone dynamics of the defunct domain of calcium vector protein.
Biochemistry, 40, 2001
|
|
1M8B
 
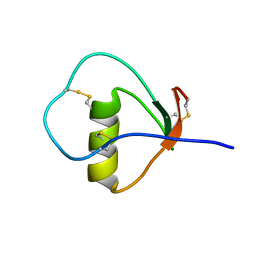 | | Solution structure of the C State of turkey ovomucoid at pH 2.5 | | Descriptor: | Ovomucoid | | Authors: | Song, J, Laskowski Jr, M, Qasim, M.A, Markley, J.L. | | Deposit date: | 2002-07-24 | | Release date: | 2002-09-04 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Two conformational states of Turkey ovomucoid third domain at low pH: three-dimensional structures, internal dynamics, and interconversion kinetics and thermodynamics.
Biochemistry, 42, 2003
|
|
2N5U
 | |
2LED
 
 | |
2LEE
 
 | |
2LHA
 | | Solution structure of C2B with IP6 | | Descriptor: | INOSITOL HEXAKISPHOSPHATE, Synaptotagmin-1 | | Authors: | Joung, M, Mohan, S.K, Yu, C. | | Deposit date: | 2011-08-08 | | Release date: | 2012-05-02 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Molecular Level Interaction of Inositol Hexaphosphate with the C2B Domain of Human Synaptotagmin I
Biochemistry, 2012
|
|
2M2S
 | | Solution structure of the antimicrobial peptide [Aba5,7,12,14]BTD-2 | | Descriptor: | [Aba5,7,12,14]BTD-2 | | Authors: | Conibear, A.C, Rosengren, K, Daly, N.L, Troiera Henriques, S, Craik, D.J. | | Deposit date: | 2013-01-02 | | Release date: | 2013-02-27 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | The cyclic cystine ladder in theta-defensins is important for structure and stability, but not antibacterial activity.
J.Biol.Chem., 288, 2013
|
|
