7D8C
 
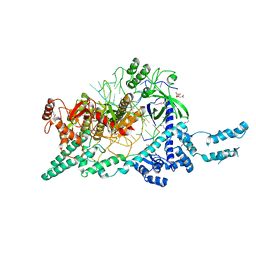 | | Crystal structure of the Cas12i1-crRNA binary complex | | Descriptor: | 12i1, CITRIC ACID, RNA (3-MER), ... | | Authors: | Zhang, B, Luo, D.Y, Li, Y, OuYang, S.Y. | | Deposit date: | 2020-10-07 | | Release date: | 2021-05-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Mechanistic insights into the R-loop formation and cleavage in CRISPR-Cas12i1.
Nat Commun, 12, 2021
|
|
6E82
 
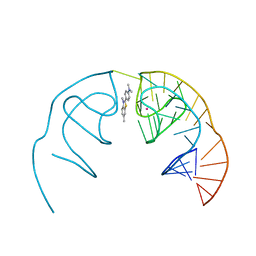 | |
2DUL
 
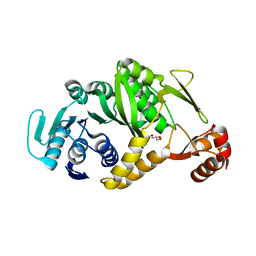 | | Crystal structure of tRNA G26 methyltransferase Trm1 in apo form from Pyrococcus horikoshii | | Descriptor: | GLYCEROL, N(2),N(2)-dimethylguanosine tRNA methyltransferase | | Authors: | Ihsanawati, Shirouzu, M, Bessho, Y, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-07-24 | | Release date: | 2007-01-24 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of tRNA N(2),N(2)-Guanosine Dimethyltransferase Trm1 from Pyrococcus horikoshii
J.Mol.Biol., 383, 2008
|
|
4MCF
 
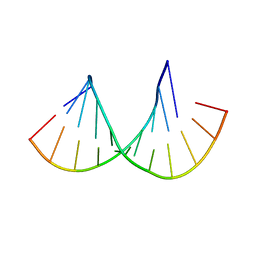 | | Crystal structure of the Gas5 GRE Mimic | | Descriptor: | Gas5 GREM Fwd, Gas5 GREM Rev, SULFATE ION | | Authors: | Hudson, W.H, Ortlund, E.A. | | Deposit date: | 2013-08-21 | | Release date: | 2014-11-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.899 Å) | | Cite: | Conserved sequence-specific lincRNA-steroid receptor interactions drive transcriptional repression and direct cell fate.
Nat Commun, 5, 2014
|
|
6E81
 
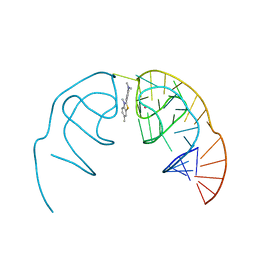 | | Crystal structure of the Corn aptamer in complex with ThT | | Descriptor: | 2-[4-(dimethylamino)phenyl]-3,6-dimethyl-1,3-benzothiazol-3-ium, POTASSIUM ION, RNA (36-MER) | | Authors: | Sjekloca, L, Ferre-D'Amare, A.R. | | Deposit date: | 2018-07-27 | | Release date: | 2019-07-31 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.721 Å) | | Cite: | Binding between G Quadruplexes at the Homodimer Interface of the Corn RNA Aptamer Strongly Activates Thioflavin T Fluorescence.
Cell Chem Biol, 26, 2019
|
|
1WAP
 
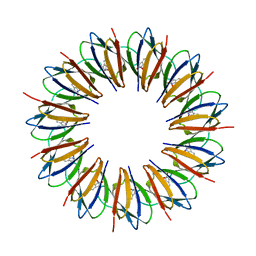 | |
4MCE
 
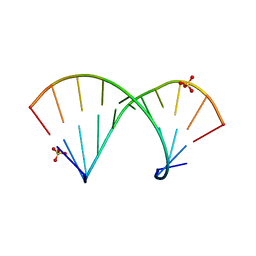 | | Crystal structure of the Gas5 GRE Mimic | | Descriptor: | Gas5 GREM Fwd, Gas5 GREM Rev, SULFATE ION | | Authors: | Hudson, W.H, Ortlund, E.A. | | Deposit date: | 2013-08-21 | | Release date: | 2014-09-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.205 Å) | | Cite: | Conserved sequence-specific lincRNA-steroid receptor interactions drive transcriptional repression and direct cell fate.
Nat Commun, 5, 2014
|
|
1YV2
 
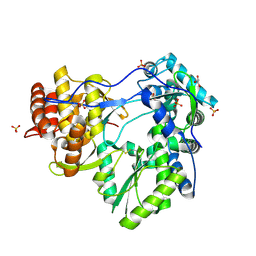 | | Hepatitis C virus NS5B RNA-dependent RNA Polymerase genotype 2a | | Descriptor: | GLYCEROL, RNA dependent RNA polymerase, SULFATE ION | | Authors: | Biswal, B.K, Cherney, M.M, Wang, M, Chan, L, Yannopoulos, C.G, Bilimoria, D, Nicolas, O, Bedard, J, James, M.N.G. | | Deposit date: | 2005-02-14 | | Release date: | 2005-03-22 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structures of the RNA-dependent RNA Polymerase Genotype 2a of Hepatitis C Virus Reveal Two Conformations and Suggest Mechanisms of Inhibition by Non-nucleoside Inhibitors
J.Biol.Chem., 280, 2005
|
|
4BJJ
 
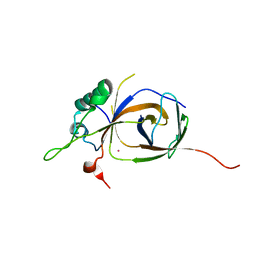 | | Sfc1-Sfc7 dimerization module | | Descriptor: | MERCURY (II) ION, TRANSCRIPTION FACTOR TAU SUBUNIT SFC1, TRANSCRIPTION FACTOR TAU SUBUNIT SFC7 | | Authors: | Taylor, N.M.I, Baudin, F, von Scheven, G, Muller, C.W. | | Deposit date: | 2013-04-18 | | Release date: | 2013-07-31 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | RNA Polymerase III-Specific General Transcription Factor Iiic Contains a Heterodimer Resembling Tfiif RAP30/RAP74.
Nucleic Acids Res., 41, 2013
|
|
4BJI
 
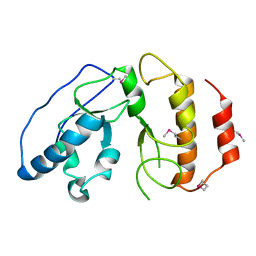 | | Sfc1-DBD | | Descriptor: | TRANSCRIPTION FACTOR TAU SUBUNIT SFC1 | | Authors: | Taylor, N.M.I, Baudin, F, von Scheven, G, Muller, C.W. | | Deposit date: | 2013-04-18 | | Release date: | 2013-07-31 | | Last modified: | 2013-11-06 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | RNA Polymerase III-Specific General Transcription Factor Iiic Contains a Heterodimer Resembling Tfiif RAP30/RAP74.
Nucleic Acids Res., 41, 2013
|
|
2CKZ
 
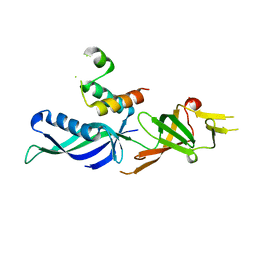 | | X-ray structure of RNA polymerase III subcomplex C17-C25. | | Descriptor: | DNA-DIRECTED RNA POLYMERASE III 18 KD POLYPEPTIDE, DNA-DIRECTED RNA POLYMERASE III 25 KD POLYPEPTIDE | | Authors: | Jasiak, A.J, Armache, K.-J, Martens, B, Jansen, R.-P, Cramer, P. | | Deposit date: | 2006-04-24 | | Release date: | 2006-07-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural Biology of RNA Polymerase III: Subcomplex C17/-C25 X-Ray Structure and 11-Subunit Enzyme Model
Mol.Cell, 23, 2006
|
|
2KXN
 
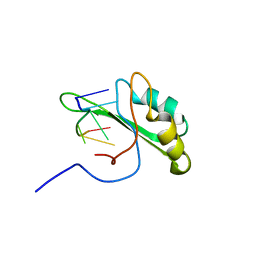 | | NMR structure of human Tra2beta1 RRM in complex with AAGAAC RNA | | Descriptor: | 5'-R(*AP*AP*GP*AP*AP*C)-3', Transformer-2 protein homolog beta | | Authors: | Clery, A, Jayne, S, Benderska, N, Dominguez, C, Stamm, S, Allain, F.H.-T. | | Deposit date: | 2010-05-10 | | Release date: | 2011-03-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Molecular basis of purine-rich RNA recognition by the human SR-like protein Tra2-beta1
Nat.Struct.Mol.Biol., 18, 2011
|
|
4LS9
 
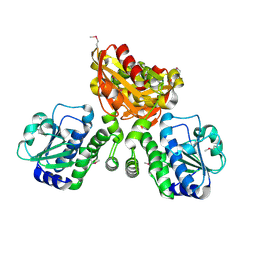 | | Structure of mycobacterial nrnA homolog reveals multifunctional nuclease activities | | Descriptor: | DHH family protein, GLYCEROL, MANGANESE (II) ION | | Authors: | Kumar, D, Srivastav, R, Grover, A, Manjasetty, B.A, Sharma, R, Taneja, B. | | Deposit date: | 2013-07-22 | | Release date: | 2014-07-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Unique subunit packing in mycobacterial nanoRNase leads to alternate substrate recognitions in DHH phosphodiesterases
Nucleic Acids Res., 42, 2014
|
|
2M22
 
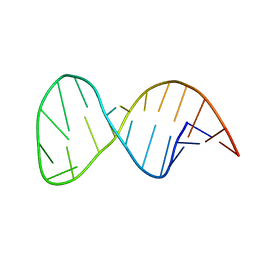 | | Solution structure of the helix II template boundary element from Tetrahymena telomerase RNA | | Descriptor: | 5'-R(*GP*GP*CP*AP*GP*AP*UP*CP*UP*GP*UP*AP*AP*UP*AP*GP*AP*AP*CP*UP*GP*CP*C)-3' | | Authors: | Cash, D.D, Richards, R.J, Theimer, C.A, Finger, D.L, Feigon, J. | | Deposit date: | 2012-12-11 | | Release date: | 2013-03-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the helix II template boundary element from Tetrahymena telomerase RNA
Nucleic Acids Res., 34, 2006
|
|
2IKF
 
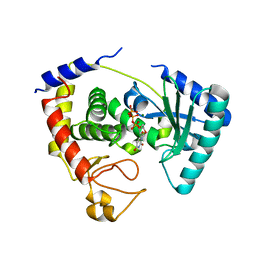 | |
1BRI
 
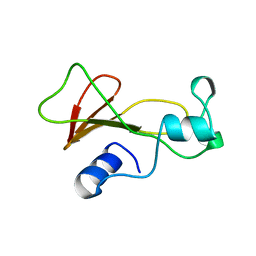 | | BARNASE MUTANT WITH ILE 76 REPLACED BY ALA | | Descriptor: | BARNASE | | Authors: | Cramer, P.C, Buckle, A, Fersht, A. | | Deposit date: | 1995-03-09 | | Release date: | 1995-07-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural and energetic responses to cavity-creating mutations in hydrophobic cores: observation of a buried water molecule and the hydrophilic nature of such hydrophobic cavities.
Biochemistry, 35, 1996
|
|
1BRK
 
 | | BARNASE MUTANT WITH ILE 96 REPLACED BY ALA | | Descriptor: | BARNASE, ZINC ION | | Authors: | Cramer, P.C, Buckle, A, Fersht, A. | | Deposit date: | 1995-03-09 | | Release date: | 1995-07-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and energetic responses to cavity-creating mutations in hydrophobic cores: observation of a buried water molecule and the hydrophilic nature of such hydrophobic cavities.
Biochemistry, 35, 1996
|
|
1RYM
 
 | |
1RYB
 
 | |
1RYN
 
 | |
3AXS
 
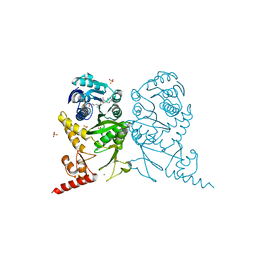 | | Complex structure of tRNA methyltransferase Trm1 from Aquifex aeolicus with sinefungin | | Descriptor: | Probable N(2),N(2)-dimethylguanosine tRNA methyltransferase Trm1, SINEFUNGIN, SULFATE ION, ... | | Authors: | Ihsanawati, Sengoku, T, Yokoyama, S, Bessho, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-04-13 | | Release date: | 2011-08-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.162 Å) | | Cite: | Substrate tRNA recognition mechanism of a multisite-specific tRNA methyltransferase, Aquifex aeolicus Trm1, based on the X-ray crystal structure
J.Biol.Chem., 286, 2011
|
|
4E48
 
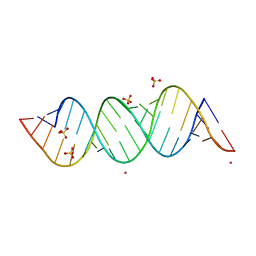 | |
3AXT
 
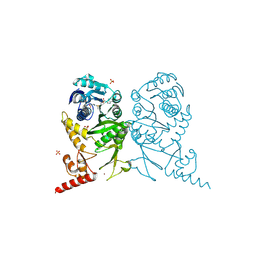 | | Complex structure of tRNA methyltransferase Trm1 from Aquifex aeolicus with S-adenosyl-L-Methionine | | Descriptor: | Probable N(2),N(2)-dimethylguanosine tRNA methyltransferase Trm1, S-ADENOSYLMETHIONINE, SULFATE ION, ... | | Authors: | Ihsanawati, Sengoku, T, Yokoyama, S, Bessho, Y, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2011-04-14 | | Release date: | 2011-08-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.491 Å) | | Cite: | Substrate tRNA recognition mechanism of a multisite-specific tRNA methyltransferase, Aquifex aeolicus Trm1, based on the X-ray crystal structure
J.Biol.Chem., 286, 2011
|
|
4YE2
 
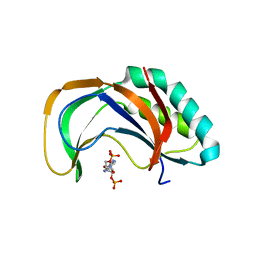 | |
1BRJ
 
 | | BARNASE MUTANT WITH ILE 88 REPLACED BY ALA | | Descriptor: | BARNASE, ZINC ION | | Authors: | Cramer, P.C, Buckle, A, Fersht, A. | | Deposit date: | 1995-03-09 | | Release date: | 1995-07-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and energetic responses to cavity-creating mutations in hydrophobic cores: observation of a buried water molecule and the hydrophilic nature of such hydrophobic cavities.
Biochemistry, 35, 1996
|
|
