8QC4
 
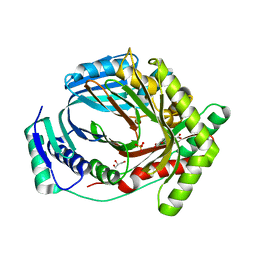 | | M. tuberculosis salicylate synthase MbtI in complex with 5-(3-carboxyphenyl)furan-2-carboxylic acid | | Descriptor: | 5-(3-carboxyphenyl)furan-2-carboxylic acid, GLYCEROL, SULFATE ION, ... | | Authors: | Mori, M, Villa, S, Meneghetti, F, Bellinzoni, M. | | Deposit date: | 2023-08-25 | | Release date: | 2023-11-15 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.578 Å) | | Cite: | Structural Study of a New MbtI-Inhibitor Complex: Towards an Optimized Model for Structure-Based Drug Discovery.
Pharmaceuticals, 16, 2023
|
|
7FR6
 
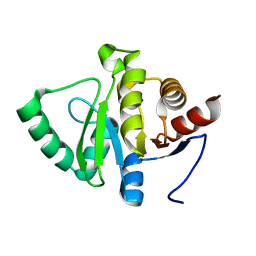 | |
9FSL
 
 | | Crystal structure of CyuA from Methanococcus maripaludis with [2Fe-2S] clusters solved by Fe-SAD | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CHLORIDE ION, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Pecqueur, L, He, N, Golinelli-Pimpaneau, B. | | Deposit date: | 2024-06-21 | | Release date: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (2.417 Å) | | Cite: | Insights into the phylogenetic distribution, structure and function of [4Fe-4S]-dependent L-cysteine desulfidase, an enzyme that supplies sulfide to the archaeon Methanococcus maripaludis
To Be Published
|
|
7FR8
 
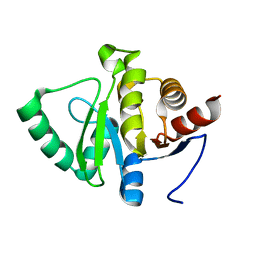 | |
7FR1
 
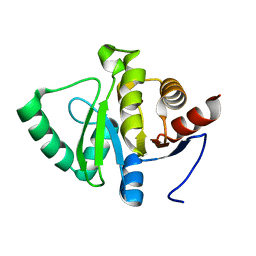 | |
7FRB
 
 | |
7FR5
 
 | |
7FRA
 
 | |
7FR7
 
 | |
7FR2
 
 | |
7FR0
 
 | |
7FR4
 
 | |
7FR3
 
 | |
7FRC
 
 | |
7FR9
 
 | |
7FRD
 
 | |
8QN5
 
 | | M. tuberculosis salicylate synthase MbtI in complex with methyl-AMT (new crystal form) | | Descriptor: | 3-{[(1Z)-1-carboxyprop-1-en-1-yl]oxy}-2-hydroxybenzoic acid, AMMONIUM ION, CITRATE ANION, ... | | Authors: | Mori, M, Villa, S, Meneghetti, M, Bellinzoni, M. | | Deposit date: | 2023-09-25 | | Release date: | 2023-11-15 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.544 Å) | | Cite: | Structural Study of a New MbtI-Inhibitor Complex: Towards an Optimized Model for Structure-Based Drug Discovery.
Pharmaceuticals, 16, 2023
|
|
9FQP
 
 | | EGFR Exon20 insertion mutant NPG bound with Compound 23 | | Descriptor: | (7~{S})-3-[(3-chloranyl-2-methoxy-phenyl)amino]-2-(3-fluoranylpyridin-4-yl)-7-(2-methoxyethyl)-1,5,6,7-tetrahydropyrrolo[3,2-c]pyridin-4-one, Epidermal growth factor receptor | | Authors: | Hilbert, B.J, Brooijmans, N, Milgram, B.C, Pagliarini, R.A. | | Deposit date: | 2024-06-17 | | Release date: | 2025-01-29 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (2.495 Å) | | Cite: | Discovery of STX-721, a Covalent, Potent, and Highly Mutant-Selective EGFR/HER2 Exon20 Insertion Inhibitor for the Treatment of Non-Small Cell Lung Cancer.
J.Med.Chem., 68, 2025
|
|
9H37
 
 | |
9H2X
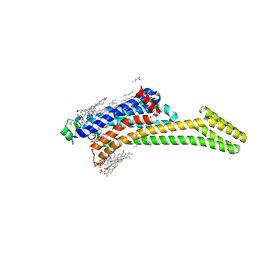 
 | | Crystal structure of stabilized A2A adenosine receptor A2AR-StaR2-bRIL in complex with compound 7, a novel nanomolar A2A receptor antagonist from modern hit-finding with structure-guided de novo design | | Descriptor: | 4-(furan-2-yl)-6-(6-imidazol-1-ylpyridin-2-yl)-1,3,5-triazin-2-amine, Adenosine receptor A2a,Soluble cytochrome b562, CHOLESTEROL, ... | | Authors: | Tian, G, Maja, N. | | Deposit date: | 2024-10-15 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Identification of nanomolar adenosine A2A receptor ligands using reinforcement learning and structure-based drug design
To Be Published
|
|
9GL9
 
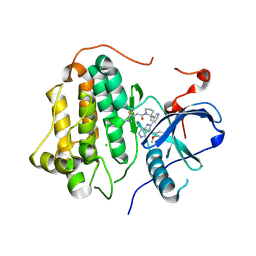 | | Wild-type EGFR bound with STX-721 | | Descriptor: | (2R,3S)-3-[(3-chloranyl-2-methoxy-phenyl)amino]-2-[3-[2-[(2R)-1-[(E)-4-(dimethylamino)but-2-enoyl]-2-methyl-pyrrolidin-2-yl]ethynyl]pyridin-4-yl]-1,2,3,5,6,7-hexahydropyrrolo[3,2-c]pyridin-4-one, CHLORIDE ION, Epidermal growth factor receptor | | Authors: | Hilbert, B.J, Brooijmans, N, Milgram, B.C, Pagliarini, R.A. | | Deposit date: | 2024-08-27 | | Release date: | 2025-05-14 | | Last modified: | 2025-06-11 | | Method: | X-RAY DIFFRACTION (2.147 Å) | | Cite: | STX-721, a Covalent EGFR/HER2 Exon 20 Inhibitor, Utilizes Exon 20-Mutant Dynamic Protein States and Achieves Unique Mutant Selectivity Across Human Cancer Models.
Clin.Cancer Res., 2025
|
|
9GL7
 
 | | EGFR Exon20 insertion mutant NPG bound with (S)-3-((3-chloro-2-methoxyphenyl)amino)-2-(3-((tetrahydrofuran-2-yl)methoxy)pyridin-4-yl)-1,5,6,7-tetrahydro-4H-pyrrolo[3,2-c]pyridin-4-one | | Descriptor: | 3-[(3-chloranyl-2-methoxy-phenyl)amino]-2-[3-[[(2S)-oxolan-2-yl]methoxy]pyridin-4-yl]-1,5,6,7-tetrahydropyrrolo[3,2-c]pyridin-4-one, Epidermal growth factor receptor | | Authors: | Hilbert, B.J, Brooijmans, N, Milgram, B.C, Pagliarini, R.A. | | Deposit date: | 2024-08-27 | | Release date: | 2025-05-14 | | Last modified: | 2025-06-11 | | Method: | X-RAY DIFFRACTION (1.878 Å) | | Cite: | STX-721, a Covalent EGFR/HER2 Exon 20 Inhibitor, Utilizes Exon 20-Mutant Dynamic Protein States and Achieves Unique Mutant Selectivity Across Human Cancer Models.
Clin.Cancer Res., 2025
|
|
9GMC
 
 | | Crystal structure of the complex formed between the radical SAM protein ChlB and the R3A mutant of ChlA | | Descriptor: | ACETATE ION, ChlA R3A mutant, ChlB radical SAM domain, ... | | Authors: | de la Mora, E, Ruel, J, Usclat, A, Martin, L, Amara, P, Morinaka, B, Nicolet, Y. | | Deposit date: | 2024-08-28 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Peptide Recognition and Mechanism of the Radical S -Adenosyl-l-methionine Multiple Cyclophane Synthase ChlB.
J.Am.Chem.Soc., 147, 2025
|
|
9GM3
 
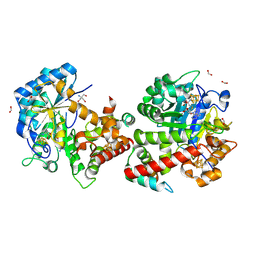 | | Crystal structure of the complex formed between the radical SAM protein ChlB and the leader region of its precursor substrate ChlA | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ChlA, ChlB radical SAM domain, ... | | Authors: | de la Mora, E, Ruel, J, Usclat, A, Martin, L, Amara, P, Morinaka, B, Nicolet, Y. | | Deposit date: | 2024-08-28 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.653 Å) | | Cite: | Peptide Recognition and Mechanism of the Radical S -Adenosyl-l-methionine Multiple Cyclophane Synthase ChlB.
J.Am.Chem.Soc., 147, 2025
|
|
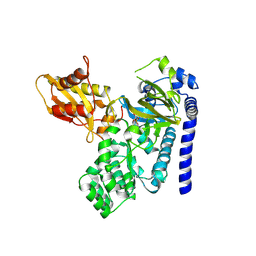
9FWD
 
 | |
