4I60
 
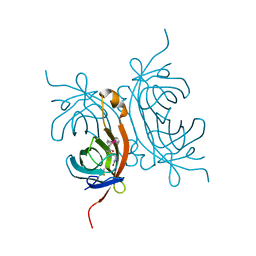 | | Crystal structure of avidin - biotinylruthenocene complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Avidin, [(1,2,3,4,5-eta)-cyclopentadienyl][(1,2,3,4,5-eta)-{5-[(3aS,4S,6aR)-2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl]pentanoyl}cyclopentadienyl]ruthenium | | Authors: | Strzelczyk, P, Bujacz, A, Bujacz, G. | | Deposit date: | 2012-11-29 | | Release date: | 2013-08-14 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural investigation of the interactions of biotinylruthenocene with avidin.
Chem.Biol.Interact, 204, 2013
|
|
9OGK
 
 | | Refinement of PDB-3J5R against EMD-8117 using EMAN2 | | Descriptor: | Transient receptor potential cation channel subfamily V member 1 | | Authors: | Chen, M. | | Deposit date: | 2025-04-30 | | Release date: | 2025-06-04 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Building molecular model series from heterogeneous CryoEM structures using Gaussian mixture models and deep neural networks.
Commun Biol, 8, 2025
|
|
9OMT
 
 | | X-ray structure of paraoxon (POX)-inhibited human acetylcholinesterase (hAChE) in complex with bis-oxime reactivator LG-1922 | | Descriptor: | (2E)-2-(hydroxyimino)-N-{3-[(3R)-3-(2-{[(2E)-2-(hydroxyimino)acetyl]amino}ethyl)piperidin-1-yl]propyl}acetamide, Acetylcholinesterase, DIETHYL PHOSPHONATE, ... | | Authors: | Kovalevsky, A, Radic, Z, Gerlits, O. | | Deposit date: | 2025-05-14 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.704 Å) | | Cite: | Kinetic and structural evidence for specific DMSO interference with reversible binding of uncharged bis-oximes to hAChE and their reactivation kinetics of OP-hAChE.
Chem.Biol.Interact., 419, 2025
|
|
9OMS
 
 | | X-ray structure of human acetylcholinesterase (hAChE) in complex with bis-oxime reactivator LG-1922 | | Descriptor: | 2-(hydroxyimino)-N-{2-[(3S)-1-(3-{[(2E)-2-(hydroxyimino)acetyl]amino}propyl)piperidin-3-yl]ethyl}acetamide, Acetylcholinesterase, GLYCEROL, ... | | Authors: | Kovalevsky, A, Radic, Z, Gerlits, O. | | Deposit date: | 2025-05-14 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Kinetic and structural evidence for specific DMSO interference with reversible binding of uncharged bis-oximes to hAChE and their reactivation kinetics of OP-hAChE.
Chem.Biol.Interact., 419, 2025
|
|
8WZU
 
 | |
5H33
 
 | |
8EHH
 
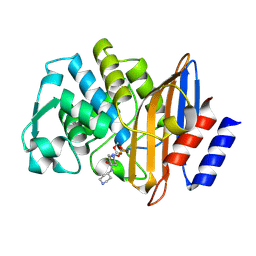 | | Crystal structure of the class A extended-spectrum beta-lactamase CTX-M-96 in complex with relebactam at 1.03 Angstrom resolution | | Descriptor: | (2S,5R)-1-formyl-N-(piperidin-4-yl)-5-[(sulfooxy)amino]piperidine-2-carboxamide, Beta-lactamase | | Authors: | Power, P, Ghiglione, B, Bonomo, R.A, Rodriguez, M.M, Gutkind, G, Klinke, S. | | Deposit date: | 2022-09-14 | | Release date: | 2023-09-20 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.03 Å) | | Cite: | Crystal structure of the class A extended-spectrum beta-lactamase CTX-M-96 in complex with relebactam at 1.03 Angstrom resolution.
Antimicrob.Agents Chemother., 68, 2024
|
|
4UBP
 
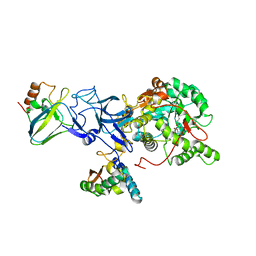 | | STRUCTURE OF BACILLUS PASTEURII UREASE INHIBITED WITH ACETOHYDROXAMIC ACID AT 1.55 A RESOLUTION | | Descriptor: | ACETOHYDROXAMIC ACID, NICKEL (II) ION, PROTEIN (UREASE (CHAIN A)), ... | | Authors: | Benini, S, Rypniewski, W.R, Wilson, K.S, Ciurli, S, Mangani, S. | | Deposit date: | 1999-02-25 | | Release date: | 2000-03-06 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The complex of Bacillus pasteurii urease with acetohydroxamate anion from X-ray data at 1.55 A resolution.
J.Biol.Inorg.Chem., 5, 2000
|
|
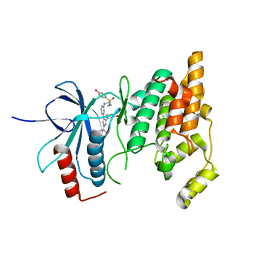
8ENJ
 
 | |
6VIF
 
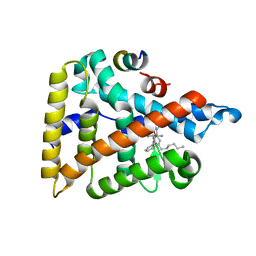 | | Human LRH-1 ligand-binding domain bound to agonist cpd 15 and fragment of coregulator TIF-2 | | Descriptor: | N-[(8beta,11alpha,12alpha)-8-{[methyl(phenyl)amino]methyl}-1,6:7,14-dicycloprosta-1(6),2,4,7(14)-tetraen-11-yl]sulfuric diamide, Nuclear receptor coactivator 2, Nuclear receptor subfamily 5 group A member 2 | | Authors: | Cato, M.L, Ortlund, E.A. | | Deposit date: | 2020-01-13 | | Release date: | 2020-06-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Development of a new class of liver receptor homolog-1 (LRH-1) agonists by photoredox conjugate addition.
Bioorg.Med.Chem.Lett., 30, 2020
|
|
2W3B
 
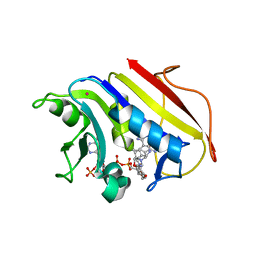 | | HUMAN DIHYDROFOLATE REDUCTASE COMPLEXED WITH NADPH AND A LIPOPHILIC ANTIFOLATE SELECTIVE FOR MYCOBACTERIUM AVIUM DHFR, 6-((2,5- DIETHOXYPHENYL)AMINOMETHYL)-2,4-DIAMINO-5-METHYLPYRIDO(2,3-D) PYRIMIDINE (SRI-8686) | | Descriptor: | 6-{[(2,5-DIETHOXYPHENYL)AMINO]METHYL}-5-METHYLPYRIDO[2,3-D]PYRIMIDINE-2,4-DIAMINE, DIHYDROFOLATE REDUCTASE, FOLIC ACID, ... | | Authors: | Leung, A.K.W, Reynolds, R.C, Borhani, D.W. | | Deposit date: | 2008-11-11 | | Release date: | 2009-11-17 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Structural Basis for Selective Inhibition of Mycobacterium Avium Dihydrofolate Reductase by a Lipophilic Antifolate
To be Published
|
|
9GNS
 
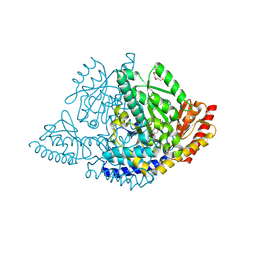 | | X-ray structure of Human holo aromatic L-amino acid decarboxylase (AADC) complex with Carbidopa at physiological pH | | Descriptor: | Aromatic-L-amino-acid decarboxylase, CARBIDOPA, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Perduca, M, Bisello, G, Bertoldi, M. | | Deposit date: | 2024-09-04 | | Release date: | 2025-05-14 | | Last modified: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | alpha-Hydrazino Acids Inhibit Pyridoxal Phosphate-Dependent Decarboxylases via "Catalytically Correct" Ketoenamine Tautomers: A Special Motif for Chemical Biology and Drug Discovery?
Acs Catalysis, 15, 2025
|
|
4PCA
 
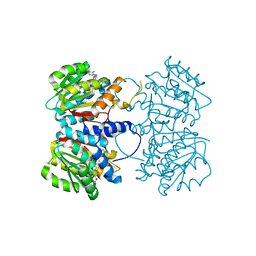 | | X-ray crystal structure of an O-methyltransferase from Anaplasma phagocytophilum bound to SAH and Manganese | | Descriptor: | 1,2-ETHANEDIOL, MANGANESE (II) ION, O-methyltransferase family protein, ... | | Authors: | Fairman, J.W, Abendroth, J, Lorimer, D, Edwards, T.E, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2014-04-14 | | Release date: | 2014-06-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | An O-Methyltransferase Is Required for Infection of Tick Cells by Anaplasma phagocytophilum.
Plos Pathog., 11, 2015
|
|
4PCL
 
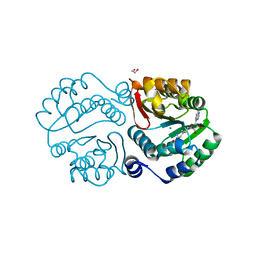 | | X-ray crystal structure of an O-methyltransferase from Anaplasma phagocytophilum bound to SAM and a Manganese ion. | | Descriptor: | 1,2-ETHANEDIOL, MANGANESE (II) ION, O-methyltransferase family protein, ... | | Authors: | Fairman, J.W, Edwards, T.E, Lorimer, D, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2014-04-15 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | An O-Methyltransferase Is Required for Infection of Tick Cells by Anaplasma phagocytophilum.
Plos Pathog., 11, 2015
|
|
8WQ9
 
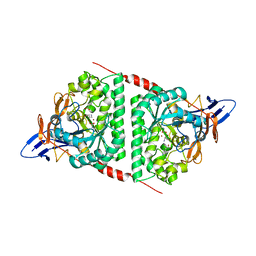 | |
9BYJ
 
 | | Crystal Structure of Hck in complex with the Src-family kinase inhibitor A-419259 | | Descriptor: | 1,2-ETHANEDIOL, 7-[trans-4-(4-methylpiperazin-1-yl)cyclohexyl]-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine, DIMETHYL SULFOXIDE, ... | | Authors: | Selzer, A.M, Alvarado, J.J, Smithgall, T.E. | | Deposit date: | 2024-05-23 | | Release date: | 2024-10-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Cocrystallization of the Src-Family Kinase Hck with the ATP-Site Inhibitor A-419259 Stabilizes an Extended Activation Loop Conformation.
Biochemistry, 63, 2024
|
|
9CQE
 
 | |
2W3A
 
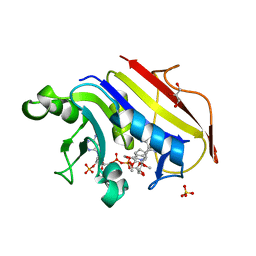 | |
6Q2F
 
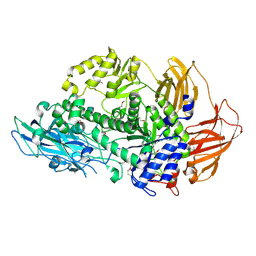 | | Structure of Rhamnosidase from Novosphingobium sp. PP1Y | | Descriptor: | Glycoside hydrolase family protein, SODIUM ION | | Authors: | Terry, B, Ha, J, Izzo, V, Sazinsky, M.H. | | Deposit date: | 2019-08-07 | | Release date: | 2019-11-27 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.20000076 Å) | | Cite: | The crystal structure and insight into the substrate specificity of the alpha-L rhamnosidase RHA-P from Novosphingobium sp. PP1Y.
Arch.Biochem.Biophys., 679, 2019
|
|
9O38
 
 | | Transmembrane domains of the human sweet receptor (TAS1R2 + TAS1R3) from Class 3 particles (rigidly fitted from PDB:9NOX and 9NOR) | | Descriptor: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Nanobody 35 (NB35), ... | | Authors: | Juen, Z, Lu, Z, Yu, R, Chang, A.N, Wang, B, Fitzpatrick, A.W.P, Zuker, C.S. | | Deposit date: | 2025-04-06 | | Release date: | 2025-05-14 | | Last modified: | 2025-08-06 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | The structure of human sweetness.
Cell, 188, 2025
|
|
6PW8
 
 | |
8E0G
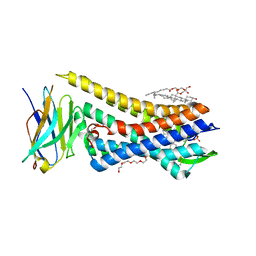 
 | |
8F8O
 
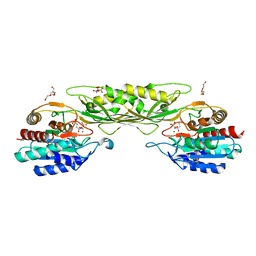 | | Crystal Structure of the Succinyl-diaminopimelate Desuccinylase (DapE) from Acinetobacter baumannii in complex with Succinic and L-Lactic Acids | | Descriptor: | (2S)-2-HYDROXYPROPANOIC ACID, CITRIC ACID, SUCCINIC ACID, ... | | Authors: | Minasov, G, Shuvalova, L, Brunzelle, J.S, Dubrovska, I, Pshenychnyi, S, Satchell, K.J.F, Center for Structural Biology of Infectious Diseases (CSBID), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2022-11-22 | | Release date: | 2022-11-30 | | Last modified: | 2025-09-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and Structural Analysis of the Bacterial Enzyme Succinyl-Diaminopimelate Desuccinylase (DapE) from Acinetobacter baumannii.
Acs Omega, 9, 2024
|
|
5LIT
 
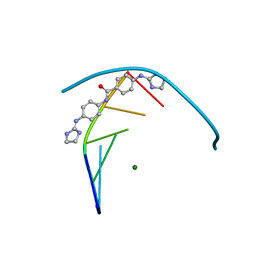 | | Structure of the DNA duplex d(AAATTT)2 with the potential antiparasitic drug 6XV at 1.25 A resolution | | Descriptor: | 4-((4,5-dihydro-1H-imidazol-2-yl)amino)-N-(4-((4,5-dihydro-1H-imidazol-2-yl)amino)phenyl)benzamide dihydrochloride, DNA (5'-D(*AP*AP*AP*TP*TP*T)-3'), MAGNESIUM ION | | Authors: | Millan, C.R, Dardonville, C, de Koning, H.P, Saperas, N, Lourdes Campos, J. | | Deposit date: | 2016-07-15 | | Release date: | 2017-06-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Functional and structural analysis of AT-specific minor groove binders that disrupt DNA-protein interactions and cause disintegration of the Trypanosoma brucei kinetoplast.
Nucleic Acids Res., 45, 2017
|
|
8YP5
 
 | | The structure of MAP2K4 complexed with 5Z7-oxozeaenol | | Descriptor: | (3S,5Z,8S,9S,11E)-8,9,16-trihydroxy-14-methoxy-3-methyl-3,4,9,10-tetrahydro-1H-2-benzoxacyclotetradecine-1,7(8H)-dione, Dual specificity mitogen-activated protein kinase kinase 4, MAGNESIUM ION | | Authors: | Yumura, S, Kinishita, T. | | Deposit date: | 2024-03-15 | | Release date: | 2025-01-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Conserved gatekeeper methionine regulates the binding and access of kinase inhibitors to ATP sites of MAP2K1, 4, and 7: Clues for developing selective inhibitors.
Bioorg.Med.Chem.Lett., 112, 2024
|
|
