5YI8
 
 | |
5G08
 
 | | Crystal structure of Drosophila NCS-1 bound to chlorpromazine | | Descriptor: | 1,2-ETHANEDIOL, 3-(2-chloro-10H-phenothiazin-10-yl)-N,N-dimethylpropan-1-amine, CALCIUM ION, ... | | Authors: | Chaves-Sanjuan, A, Infantes, L, Sanchez-Barrena, M.J. | | Deposit date: | 2016-03-17 | | Release date: | 2017-01-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Interference of the complex between NCS-1 and Ric8a with phenothiazines regulates synaptic function and is an approach for fragile X syndrome.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
2MAH
 
 | | Solution structure of Smoothened | | Descriptor: | Protein smoothened | | Authors: | Rana, R, Lee, H, Zheng, J.J. | | Deposit date: | 2013-07-09 | | Release date: | 2014-03-05 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural insights into the role of the Smoothened cysteine-rich domain in Hedgehog signalling.
Nat Commun, 4, 2013
|
|
6Y68
 
 | |
3ODN
 
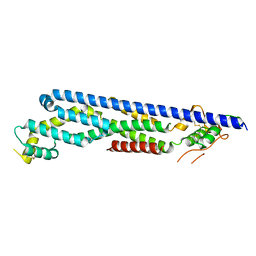 | |
6Y6Q
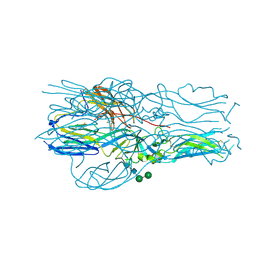 
 | |
7OGC
 
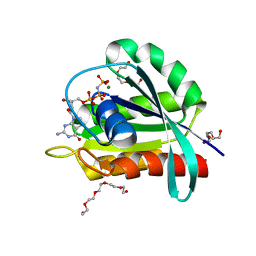 | | GTPase HRAS under 650 MPa pressure | | Descriptor: | GTPase HRas, MAGNESIUM ION, PENTAETHYLENE GLYCOL, ... | | Authors: | Colloc'h, N.C, Girard, E, Prange, T, Kalbitzer, H.R. | | Deposit date: | 2021-05-06 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Equilibria between conformational states of the Ras oncogene protein revealed by high pressure crystallography.
Chem Sci, 13, 2022
|
|
7OGD
 
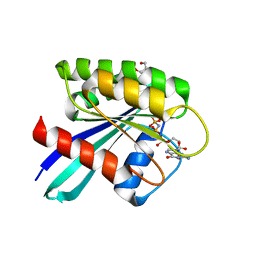 | | GTPase HRAS mutant D33K under ambient pressure | | Descriptor: | GTPase HRas, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Colloc'h, N.C, Girard, E, Prange, T, Kalbitzer, H.R. | | Deposit date: | 2021-05-06 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Equilibria between conformational states of the Ras oncogene protein revealed by high pressure crystallography.
Chem Sci, 13, 2022
|
|
7OGB
 
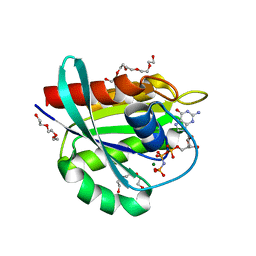 | | GTPase HRAS under 500 MPa pressure | | Descriptor: | GTPase HRas, MAGNESIUM ION, PENTAETHYLENE GLYCOL, ... | | Authors: | Colloc'h, N.C, Girard, E, Prange, T, Kalbitzer, H.R. | | Deposit date: | 2021-05-06 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Equilibria between conformational states of the Ras oncogene protein revealed by high pressure crystallography.
Chem Sci, 13, 2022
|
|
7OGF
 
 | | GTPase HRAS mutant D33K under 900 MPa pressure | | Descriptor: | GTPase HRas, MAGNESIUM ION, PENTAETHYLENE GLYCOL, ... | | Authors: | Colloc'h, N.C, Girard, E, Prange, T, Kalbitzer, H.R. | | Deposit date: | 2021-05-06 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Equilibria between conformational states of the Ras oncogene protein revealed by high pressure crystallography.
Chem Sci, 13, 2022
|
|
7OGE
 | | GTPase HRAS mutant D33K under 200 MPa pressure | | Descriptor: | GTPase HRas, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER, ... | | Authors: | Colloc'h, N.C, Girard, E, Prange, T, Kalbitzer, H.R. | | Deposit date: | 2021-05-06 | | Release date: | 2022-02-09 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Equilibria between conformational states of the Ras oncogene protein revealed by high pressure crystallography.
Chem Sci, 13, 2022
|
|
7NYQ
 
 | | Crystal structure of the Mei-P26 NHL domain | | Descriptor: | Meiotic P26, isoform C | | Authors: | Salerno-Kochan, A, Gaik, M, Medenbach, J, Glatt, S. | | Deposit date: | 2021-03-23 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Molecular insights into RNA recognition and gene regulation by the TRIM-NHL protein Mei-P26.
Life Sci Alliance, 5, 2022
|
|
7OC3
 | |
6YBH
 | | Deoxyribonucleoside Kinase | | Descriptor: | Deoxynucleoside kinase, GLYCEROL, Pyrrolo-dC, ... | | Authors: | Allouche-Arnon, H, Bar-Shir, A, Dym, O. | | Deposit date: | 2020-03-17 | | Release date: | 2021-03-31 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Computationally designed dual-color MRI reporters for noninvasive imaging of transgene expression.
Nat.Biotechnol., 40, 2022
|
|
6Y4H
 
 | |
5YI7
 
 | |
3MN9
 
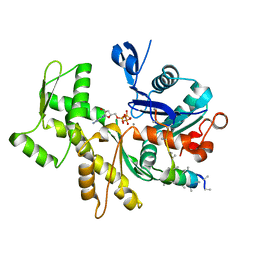 | | Structures of actin-bound WH2 domains of Spire and the implication for filament nucleation | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin-5C, CALCIUM ION, ... | | Authors: | Ducka, A.M, Sitar, T, Popowicz, G.M, Huber, R, Holak, T.A. | | Deposit date: | 2010-04-21 | | Release date: | 2010-05-26 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of actin-bound Wiskott-Aldrich syndrome protein homology 2 (WH2) domains of Spire and the implication for filament nucleation.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2NMB
 
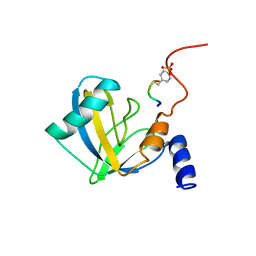 | | DNUMB PTB DOMAIN COMPLEXED WITH A PHOSPHOTYROSINE PEPTIDE, NMR, ENSEMBLE OF STRUCTURES. | | Descriptor: | PROTEIN (GPPY PEPTIDE), PROTEIN (NUMB PROTEIN) | | Authors: | Li, S.-C, Zwahlen, C, Vincent, S.J.F, McGlade, C.J, Pawson, T, Forman-Kay, J.D. | | Deposit date: | 1998-10-29 | | Release date: | 1998-11-04 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure of a Numb PTB domain-peptide complex suggests a basis for diverse binding specificity.
Nat.Struct.Biol., 5, 1998
|
|
4X8F
 
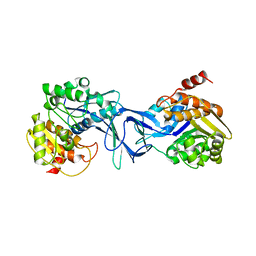 | | Vibrio cholerae O395 Ribokinase in apo form | | Descriptor: | Ribokinase | | Authors: | Paul, R, Patra, M.D, Sen, U. | | Deposit date: | 2014-12-10 | | Release date: | 2015-01-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Crystal Structure of Apo and Ligand
Bound Vibrio choleraeRibokinase (Vc-RK):
Role of Monovalent Cation Induced Activation
and Structural Flexibility in Sugar
Phosphorylation
Adv Exp Med Biol., 842, 2015
|
|
4X24
 | | Crystal structure of Vibrio cholerae 5'-methylthioadenosine/S-adenosyl homocysteine nucleosidase (MTAN) complexed with methylthio-DADMe-Immucillin-A | | Descriptor: | (3R,4S)-1-[(4-AMINO-5H-PYRROLO[3,2-D]PYRIMIDIN-7-YL)METHYL]-4-[(METHYLSULFANYL)METHYL]PYRROLIDIN-3-OL, 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase, TRIETHYLENE GLYCOL | | Authors: | Cameron, S.A, Thomas, K, Almo, S.C, Schramm, V.L. | | Deposit date: | 2014-11-25 | | Release date: | 2015-08-19 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Active site and remote contributions to catalysis in methylthioadenosine nucleosidases.
Biochemistry, 54, 2015
|
|
2ND6
 
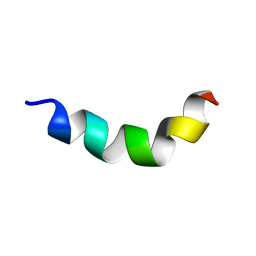 | | Structure of DK17 in GM1 LUVS | | Descriptor: | Cell penetrating peptide | | Authors: | Bera, S, Bhunia, A. | | Deposit date: | 2016-05-11 | | Release date: | 2017-03-22 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural Elucidation of the Cell-Penetrating Penetratin Peptide in Model Membranes at the Atomic Level: Probing Hydrophobic Interactions in the Blood-Brain Barrier
Biochemistry, 55, 2016
|
|
5F85
 
 | |
4WSF
 
 | | Falafel EVH1 domain bound to CENP-C FIM | | Descriptor: | Cenp-C, SULFATE ION, Serine/threonine-protein phosphatase 4 regulatory subunit 3 | | Authors: | Lefevre, S.R, Singleton, M.R. | | Deposit date: | 2014-10-27 | | Release date: | 2014-12-10 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.501 Å) | | Cite: | Centromeric binding and activity of Protein Phosphatase 4.
Nat Commun, 6, 2015
|
|
4WSI
 
 | | Crystal Structure of PALS1/Crb complex | | Descriptor: | MAGUK p55 subfamily member 5, Protein crumbs | | Authors: | Wei, Z, Li, Y, Zhang, M. | | Deposit date: | 2014-10-28 | | Release date: | 2014-11-26 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structure of Crumbs tail in complex with the PALS1 PDZ-SH3-GK tandem reveals a highly specific assembly mechanism for the apical Crumbs complex.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
3DH9
 
 | |
