4UZQ
 
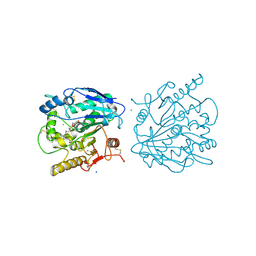 | |
5L6Z
 
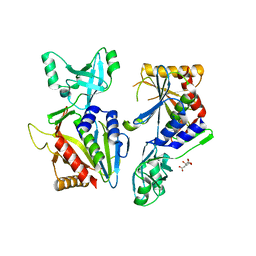 | |
5L3E
 | | LSD1-CoREST1 in complex with quinazoline-derivative reversible inhibitor | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Lysine-specific histone demethylase 1A, N~4~-(1-benzylpiperidin-4-yl)-N~2~-[3-(dimethylamino)propyl]-6,7-dimethoxyquinazoline-2,4-diamine, ... | | Authors: | Speranzini, V, Rotili, D, Ciossani, G, Pilotto, S, Forgione, M, Lucidi, A, Forneris, F, Velankar, S, Mai, A, Mattevi, A. | | Deposit date: | 2016-04-10 | | Release date: | 2016-09-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Polymyxins and quinazolines are LSD1/KDM1A inhibitors with unusual structural features.
Sci Adv, 2, 2016
|
|
5KSN
 | |
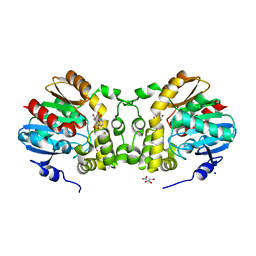
7CLZ
 
 | |
3LP9
 
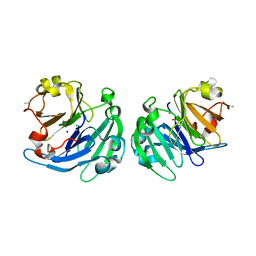 | | Crystal structure of LS24, A Seed Albumin from Lathyrus sativus | | Descriptor: | CALCIUM ION, CHLORIDE ION, LS-24, ... | | Authors: | Gaur, V, Qureshi, I.A, Singh, A, Chanana, V, Salunke, D.M. | | Deposit date: | 2010-02-05 | | Release date: | 2010-02-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure and functional insights of hemopexin fold protein from grass pea
Plant Physiol., 152, 2010
|
|
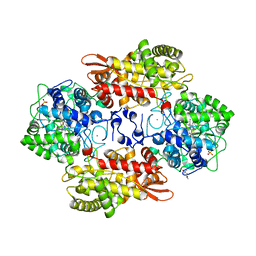
5KQ2
 
 | |
5KQI
 | |
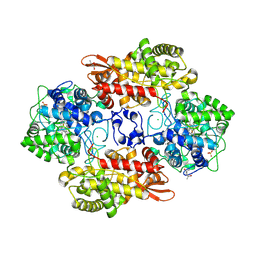
6UBE
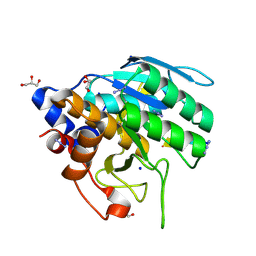 
 | | Azide-triggered subtilisin SUBT_BACAM complexed with the peptide LFRAL | | Descriptor: | AZIDE ION, GLYCEROL, Peptide LFRAL, ... | | Authors: | Toth, E.A, Bryan, P.N, Orban, J, Gallagher, D.T, Custer, G. | | Deposit date: | 2019-09-11 | | Release date: | 2020-09-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Engineering subtilisin proteases that specifically degrade active RAS.
Commun Biol, 4, 2021
|
|
5KSG
 
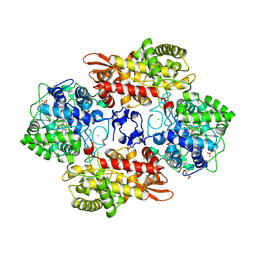 | |
3LWL
 
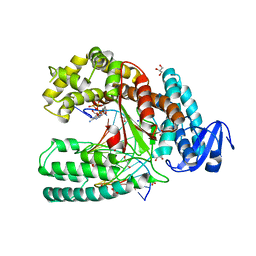 | | Structure of Klenow fragment of Taq polymerase in complex with an abasic site | | Descriptor: | 2',3'-dideoxyadenosine triphosphate, ACETATE ION, DNA (5'-D(*AP*AP*AP*(3DR)P*TP*GP*CP*GP*CP*CP*GP*TP*GP*GP*TP*C)-3'), ... | | Authors: | Marx, A, Diederichs, K, Obeid, S. | | Deposit date: | 2010-02-24 | | Release date: | 2010-05-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Replication through an abasic DNA lesion: structural basis for adenine selectivity
Embo J., 29, 2010
|
|
5KXL
 
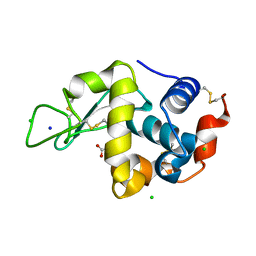 | | Hen Egg White Lysozyme at 100K, Data set 2 | | Descriptor: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | Authors: | Russi, S, Gonzalez, A, Kenner, L.R, Keedy, D.A, Fraser, J.S, van den Bedem, H. | | Deposit date: | 2016-07-20 | | Release date: | 2016-09-07 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.198 Å) | | Cite: | Conformational variation of proteins at room temperature is not dominated by radiation damage.
J Synchrotron Radiat, 24, 2017
|
|
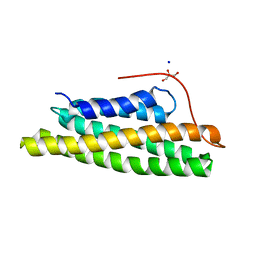
5L0I
 
 | |
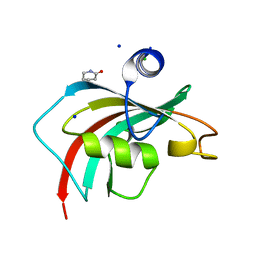
6TX7
 
 | |
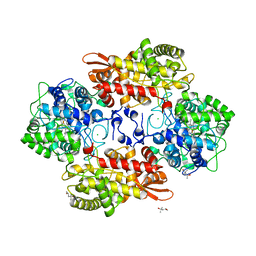
5KT9
 
 | |
5L02
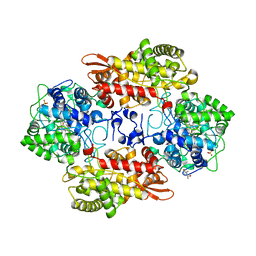 
 | | S324T variant of B. pseudomallei KatG | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Catalase-peroxidase, PHOSPHATE ION, ... | | Authors: | Loewen, P.C. | | Deposit date: | 2016-07-26 | | Release date: | 2016-08-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural characterization of the Ser324Thr variant of the catalase-peroxidase (KatG) from Burkholderia pseudomallei
J. Mol. Biol., 345, 2005
|
|
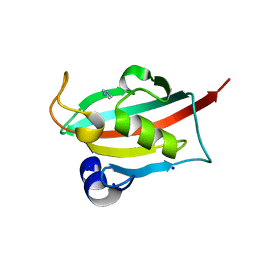
6TX8
 
 | |
5L9F
 
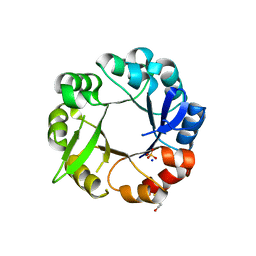 | | S. enterica HisA mutant - D10G, G11D, dup13-15, G44E, G102A | | Descriptor: | 1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino] imidazole-4-carboxamide isomerase, SODIUM ION, SULFATE ION | | Authors: | Guo, X, Soderholm, A, Selmer, M. | | Deposit date: | 2016-06-10 | | Release date: | 2017-04-19 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.594 Å) | | Cite: | Structural and functional innovations in the real-time evolution of new ( beta alpha )8 barrel enzymes.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
6U2O
 
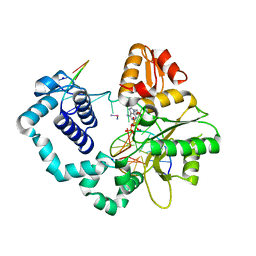 | |
8CPX
 
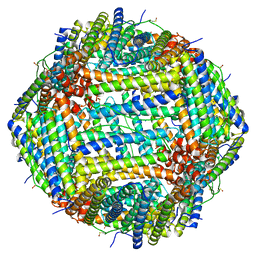 | | Human apoferritin after 488 nm laser exposure in presence of rsEGFP2 | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.76 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPM
 
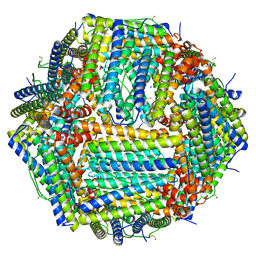 | | Human apoferritin after 405 nm laser exposure | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.81 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPV
 
 | | Human apoferritin | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.76 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPU
 
 | | Human apoferritin after 561 nm laser exposure | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.76 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
8CPS
 
 | | Human apoferritin | | Descriptor: | Ferritin heavy chain, N-terminally processed, MAGNESIUM ION, ... | | Authors: | Last, M.G.F, Noteborn, W.E.M, Sharp, T.H. | | Deposit date: | 2023-03-03 | | Release date: | 2023-03-15 | | Last modified: | 2023-11-22 | | Method: | ELECTRON MICROSCOPY (1.82 Å) | | Cite: | Super-resolution fluorescence imaging of cryosamples does not limit achievable resolution in cryoEM.
J.Struct.Biol., 215, 2023
|
|
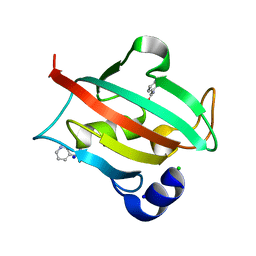
6TX6
 | |
