4F0I
 
 | | Crystal structure of apo TrkA | | Descriptor: | High affinity nerve growth factor receptor | | Authors: | Liu, J. | | Deposit date: | 2012-05-04 | | Release date: | 2012-09-05 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.302 Å) | | Cite: | The Crystal Structures of TrkA and TrkB Suggest Key Regions for Achieving Selective Inhibition.
J.Mol.Biol., 423, 2012
|
|
4F2G
 
 | | The Crystal Structure of Ornithine carbamoyltransferase from Burkholderia thailandensis E264 | | Descriptor: | 1,2-ETHANEDIOL, Ornithine carbamoyltransferase 1, PHOSPHATE ION | | Authors: | Craig, T.K, Fox, D, Staker, B, Stewart, L, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2012-05-07 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Combining functional and structural genomics to sample the essential Burkholderia structome.
Plos One, 8, 2013
|
|
3LZY
 
 | |
3M51
 
 | | Structure of the 14-3-3/PMA2 complex stabilized by Pyrrolidone1 | | Descriptor: | 14-3-3-like protein C, 2-hydroxy-5-[(5S)-3-hydroxy-5-(4-nitrophenyl)-2-oxo-4-(phenylcarbonyl)-2,5-dihydro-1H-pyrrol-1-yl]benzoic acid, N.plumbaginifolia H+-translocating ATPase mRNA | | Authors: | Ottmann, C, Rose, R, Waldmann, H. | | Deposit date: | 2010-03-12 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Identification and structure of small-molecule stabilizers of 14-3-3 protein-protein interactions
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
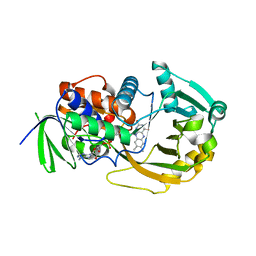
3M0O
 
 | |
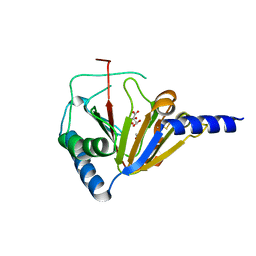
4NRM
 
 | |
3M3I
 
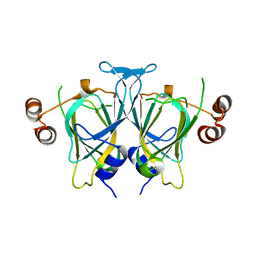 | |
3M8P
 
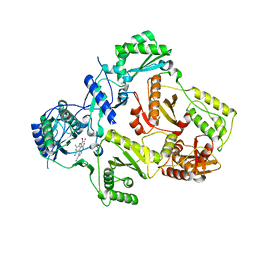 | | HIV-1 RT with NNRTI TMC-125 | | Descriptor: | 4-({6-AMINO-5-BROMO-2-[(4-CYANOPHENYL)AMINO]PYRIMIDIN-4-YL}OXY)-3,5-DIMETHYLBENZONITRILE, Reverse transcriptase/ribonuclease H, p51 RT | | Authors: | Harris, S.F, Villasenor, A. | | Deposit date: | 2010-03-18 | | Release date: | 2010-05-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Discovery of piperidin-4-yl-aminopyrimidines as HIV-1 reverse transcriptase inhibitors. N-benzyl derivatives with broad potency against resistant mutant viruses.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
3M50
 
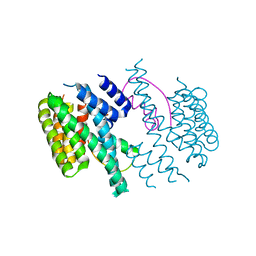 | | Structure of the 14-3-3/PMA2 complex stabilized by Epibestatin | | Descriptor: | 14-3-3-like protein C, N-[(2R,3R)-3-amino-2-hydroxy-4-phenylbutanoyl]-L-leucine, N.plumbaginifolia H+-translocating ATPase mRNA | | Authors: | Ottmann, C, Rose, R, Waldmann, H. | | Deposit date: | 2010-03-12 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Identification and structure of small-molecule stabilizers of 14-3-3 protein-protein interactions
Angew.Chem.Int.Ed.Engl., 49, 2010
|
|
4F1O
 
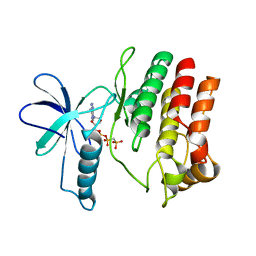 | | Crystal Structure of the L1180T mutant Roco4 Kinase Domain from D. discoideum bound to AppCp | | Descriptor: | PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER, Serine/threonine-protein kinase roco4 | | Authors: | Gilsbach, B.K, Vetter, I.R, Wittinghofer, A, Kortholt, A. | | Deposit date: | 2012-05-07 | | Release date: | 2012-06-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Roco kinase structures give insights into the mechanism of Parkinson disease-related leucine-rich-repeat kinase 2 mutations.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4B9D
 
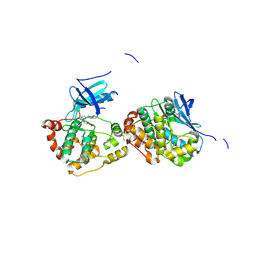 | | Crystal Structure of Human NIMA-related Kinase 1 (NEK1) with inhibitor. | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, SERINE/THREONINE-PROTEIN KINASE NEK1, ... | | Authors: | Elkins, J.M, Hanchuk, T.D.M, Lovato, D.V, Basei, F.L, Meirelles, G.V, Kobarg, J, Szklarz, M, Vollmar, M, Mahajan, P, Rellos, P, Zhang, Y, Krojer, T, Pike, A.C.W, Canning, P, von Delft, F, Raynor, J, Bountra, C, Arrowsmith, C, Edwards, A, Knapp, S. | | Deposit date: | 2012-09-04 | | Release date: | 2012-09-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | NEK1 kinase domain structure and its dynamic protein interactome after exposure to Cisplatin.
Sci Rep, 7, 2017
|
|
4F1T
 
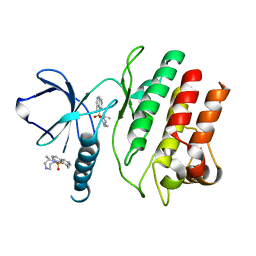 | | Crystal Structure of the Roco4 Kinase Domain from D. discoideum bound to the ROCK Inhibitor H1152 | | Descriptor: | (S)-2-METHYL-1-[(4-METHYL-5-ISOQUINOLINE)SULFONYL]-HOMOPIPERAZINE, Serine/threonine-protein kinase roco4 | | Authors: | Gilsbach, B.K, Vetter, I.R, Wittinghofer, A, Kortholt, A. | | Deposit date: | 2012-05-07 | | Release date: | 2012-06-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Roco kinase structures give insights into the mechanism of Parkinson disease-related leucine-rich-repeat kinase 2 mutations.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3M78
 
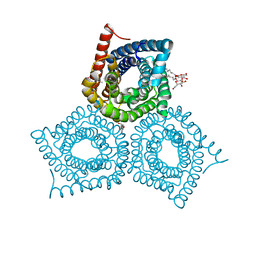 | |
4BAJ
 
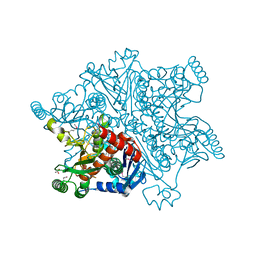 | | MYCOBACTERIUM TUBERCULOSIS CHORISMATE SYNTHASE after exposure to 266nm UV laser | | Descriptor: | ACETATE ION, CHORISMATE SYNTHASE | | Authors: | Pereira, P.J.B, Royant, A, Panjikar, S, de Sanctis, D. | | Deposit date: | 2012-09-14 | | Release date: | 2013-04-17 | | Last modified: | 2019-03-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | In-house UV radiation-damage-induced phasing of selenomethionine-labeled protein structures.
J. Struct. Biol., 181, 2013
|
|
3MCN
 
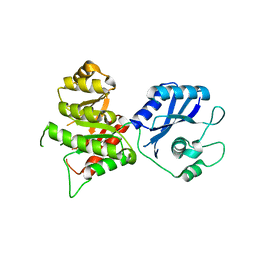 | | Crystal Structure of the 6-hyroxymethyl-7,8-dihydropterin pyrophosphokinase dihydropteroate synthase bifunctional enzyme from Francisella tularensis | | Descriptor: | 2,6-diamino-5-nitropyrimidin-4(3H)-one, 2-amino-4-hydroxy-6-hydroxymethyldihydropteridine pyrophosphokinase/dihydropteroate synthase, MAGNESIUM ION | | Authors: | Pemble IV, C.W, Mehta, P.K, Mehra, S, Li, Z, Lee, R.E, White, S.W. | | Deposit date: | 2010-03-29 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the 6-hydroxymethyl-7,8-dihydropterin pyrophosphokinase.dihydropteroate synthase bifunctional enzyme from Francisella tularensis.
Plos One, 5, 2010
|
|
4BFO
 
 | | Crystal Structure of the Starch-Binding Domain from Rhizopus oryzae Glucoamylase in Complex with isomaltotriose | | Descriptor: | GLUCOAMYLASE, alpha-D-glucopyranose-(1-6)-alpha-D-glucopyranose-(1-6)-alpha-D-glucopyranose | | Authors: | Chu, C.H, Li, K.M, Lin, S.W, Sun, Y.J. | | Deposit date: | 2013-03-21 | | Release date: | 2013-10-23 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.175 Å) | | Cite: | Crystal Structures of Starch Binding Domain from Rhizopus Oryzae Glucoamylase in Complex with Isomaltooligosaccharide: Insights Into Polysaccharide Binding Mechanism of Cbm21 Family.
Proteins, 82, 2014
|
|
3MDZ
 
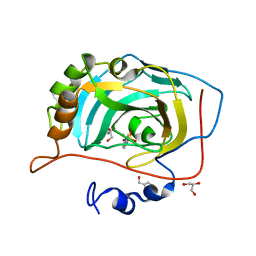 | | Crystal Structure of Human Carbonic Anhydrase VII [isoform 1], CA7 | | Descriptor: | 6-ethoxy-1,3-benzothiazole-2-sulfonamide, Carbonic anhydrase 7, GLYCEROL, ... | | Authors: | Ugochukwu, E, Shafqat, N, Pilka, E, Chaikuad, A, Krojer, T, Muniz, J, Kim, J, Bray, J, Bountra, C, Arrowsmith, C.H, Weigelt, J, Edwards, A, von Delft, F, Carpenter, E.P, Yue, W.W, Oppermann, U, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-03-31 | | Release date: | 2010-06-23 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Crystal Structure of Human Carbonic Anhydrase VII [isoform 1], CA7
to be published
|
|
3M21
 
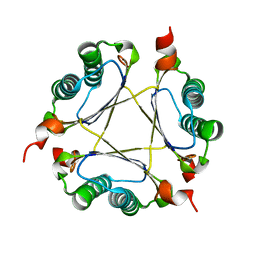 | | Crystal structure of DmpI from Helicobacter pylori Determined to 1.9 Angstroms resolution | | Descriptor: | Probable tautomerase HP_0924 | | Authors: | Hackert, M.L, Whitman, C.P, Almrud, J.J, Dasgupta, R, Kern, A.D, Czerwinski, R.M. | | Deposit date: | 2010-03-06 | | Release date: | 2010-09-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Kinetic and structural characterization of DmpI from Helicobacter pylori and Archaeoglobus fulgidus, two 4-oxalocrotonate tautomerase family members.
Bioorg.Chem., 38, 2010
|
|
4C9B
 
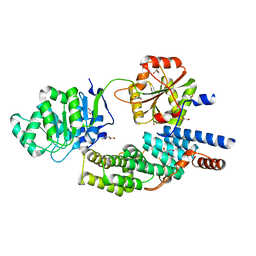 | | Crystal structure of eIF4AIII-CWC22 complex | | Descriptor: | EUKARYOTIC INITIATION FACTOR 4A-III, GLYCEROL, PHOSPHATE ION, ... | | Authors: | Buchwald, G, Schuessler, S, Basquin, C, LeHir, H, Conti, E. | | Deposit date: | 2013-10-02 | | Release date: | 2013-11-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Human Eif4Aiii-Cwc22 Complex Shows How a Dead-Box Protein is Inhibited by a Mif4G Domain
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
3MAC
 
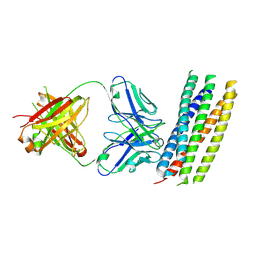 | | crystal structure of GP41-derived protein complexed with fab 8062 | | Descriptor: | Fab8062, Transmembrane glycoprotein | | Authors: | Li, M, Gustchina, E, Louis, J, Gustchina, A, Wlodawer, A, Clore, M. | | Deposit date: | 2010-03-23 | | Release date: | 2010-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural Basis of HIV-1 Neutralization by Affinity Matured Fabs Directed against the Internal Trimeric Coiled-Coil of gp41.
Plos Pathog., 6, 2010
|
|
4NRO
 
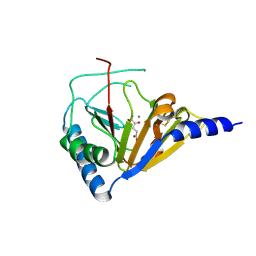 | |
3M6L
 
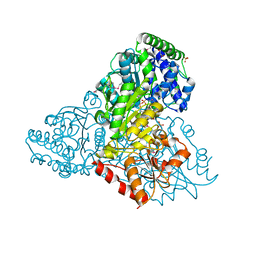 | | Crystal structure of transketolase in complex with thiamine diphosphate, ribose-5-phosphate and calcium ion | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, GLYCEROL, ... | | Authors: | Nocek, B, Makowska-Grzyska, M, Maltseva, N, Joachimiak, A, Anderson, W, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2010-03-15 | | Release date: | 2010-04-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Crystal structure of transketolase in complex with thiamine diphosphate, ribose-5-phosphate and calcium ion
To be Published
|
|
3ME1
 
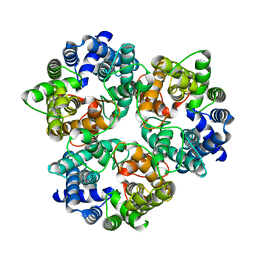 | |
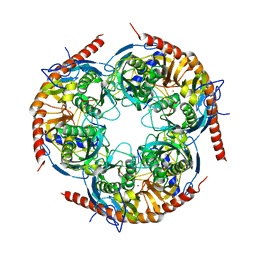
3M85
 
 | |
3MI8
 
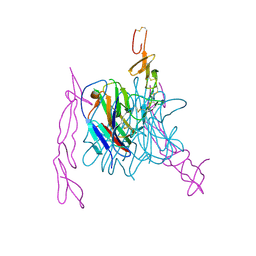 | | The structure of TL1A-DCR3 COMPLEX | | Descriptor: | TUMOR NECROSIS FACTOR LIGAND SUPERFAMILY MEMBER 15, SECRETED FORM, Tumor necrosis factor receptor superfamily member 6B | | Authors: | Zhan, C, Patskovsky, Y, Yan, Q, Li, Z, Ramagopal, U.A, Nathenson, S.G, Almo, S.C. | | Deposit date: | 2010-04-09 | | Release date: | 2011-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.951 Å) | | Cite: | Decoy Strategies: The Structure of TL1A:DcR3 Complex.
Structure, 19, 2011
|
|
