8TUZ
 
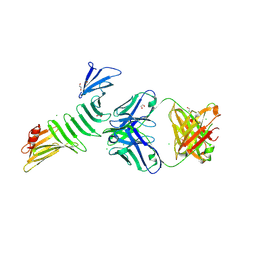 | | Fab 857-2 in complex with OspA | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Heavy chain of human monoclonal antibody 857-2, ... | | Authors: | Rudolph, M.J, Mantis, N. | | Deposit date: | 2023-08-17 | | Release date: | 2024-08-28 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.186 Å) | | Cite: | Fab 857-2 in complex with OspA
To Be Published
|
|
8K09
 
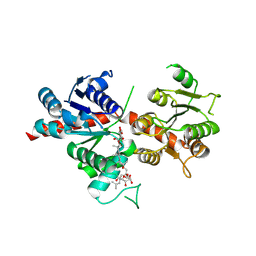 | | Pq3-O-UGT2 with 20(S)-Ginsenoside F2 | | Descriptor: | (2~{R},3~{S},4~{S},5~{R},6~{R})-2-(hydroxymethyl)-6-[[(3~{S},5~{R},8~{R},9~{R},10~{R},12~{R},13~{R},14~{R},17~{S})-17-[(2~{S})-2-[(2~{S},3~{R},4~{S},5~{S},6~{R})-6-(hydroxymethyl)-3,4,5-tris(oxidanyl)oxan-2-yl]oxy-6-methyl-hept-5-en-2-yl]-4,4,8,10,14-pentamethyl-12-oxidanyl-2,3,5,6,7,9,11,12,13,15,16,17-dodecahydro-1~{H}-cyclopenta[a]phenanthren-3-yl]oxy]oxane-3,4,5-triol, Glycosyltransferase | | Authors: | Mei, K.R, Ji, Q.S, Liu, Y.R. | | Deposit date: | 2023-07-07 | | Release date: | 2025-01-29 | | Last modified: | 2025-04-02 | | Method: | X-RAY DIFFRACTION (3.36 Å) | | Cite: | Structural Insights into the Substrate Recognition of Ginsenoside Glycosyltransferase Pq3-O-UGT2.
Adv Sci, 12, 2025
|
|
6OHR
 
 | |
9GQR
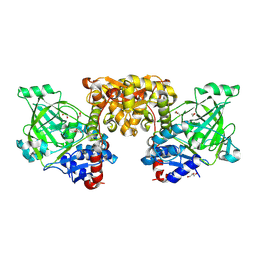 
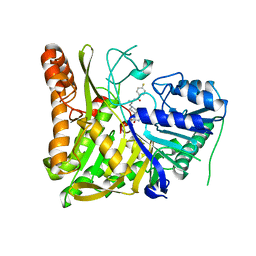
 | | Structure of a consensus-designed decarboxylase (PSC1) | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, decarboxylase | | Authors: | Gavira, J.A, Ramos, J.L, Garcia-Franco, A, de la Torre, J. | | Deposit date: | 2024-09-09 | | Release date: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Sustainable synthesis of styrene exploiting a consensus-designed decarboxylase. Purification and crystal structure of PSC1
Mbio, 2025
|
|
4DZM
 
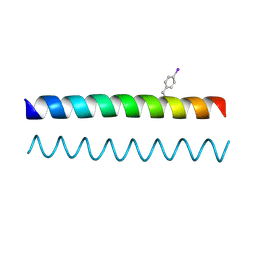 | | A de novo designed Coiled Coil CC-Di | | Descriptor: | COILED-COIL PEPTIDE CC-DI | | Authors: | Bruning, M, Thomson, A.R, Zaccai, N.R, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2012-03-01 | | Release date: | 2012-08-29 | | Last modified: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | A basis set of de novo coiled-coil Peptide oligomers for rational protein design and synthetic biology.
ACS Synth Biol, 1, 2012
|
|
8DYC
 
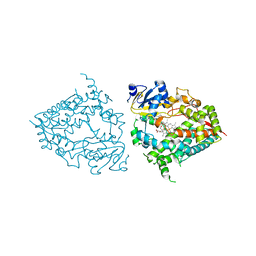 | | Human CYP3A4 bound to a substrate | | Descriptor: | 2-butyl-6-(butylamino)-1H-benzo[de]isoquinoline-1,3(2H)-dione, Cytochrome P450 3A4, GLYCEROL, ... | | Authors: | Sevrioukova, I.F. | | Deposit date: | 2022-08-04 | | Release date: | 2022-10-26 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of CYP3A4 Complexed with Fluorol Identifies the Substrate Access Channel as a High-Affinity Ligand Binding Site.
Int J Mol Sci, 23, 2022
|
|
5YHA
 
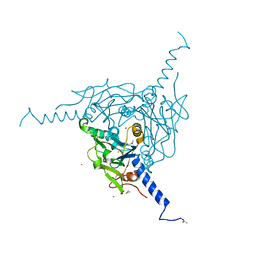 | | Crystal structure of Pd(allyl)/Wild Type Polyhedra | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, PALLADIUM ION, ... | | Authors: | Abe, S, Atsumi, K, Yamashita, K, Hirata, K, Mori, H, Ueno, T. | | Deposit date: | 2017-09-27 | | Release date: | 2017-11-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Structure of in cell protein crystals containing organometallic complexes.
Phys Chem Chem Phys, 20, 2018
|
|
6OSU
 
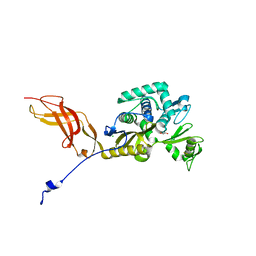 | | Crystal Structure of the D-alanyl-D-alanine carboxypeptidase DacD from Francisella tularensis | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, D-alanyl-D-alanine carboxypeptidase (Penicillin binding protein) family protein | | Authors: | Kim, Y, Stogios, P, Skarina, T, Di, R, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2019-05-02 | | Release date: | 2019-05-15 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Crystal Structure of the D-alanyl-D-alanine carboxypeptidase DacD from Francisella tularensis
To Be Published
|
|
6O88
 
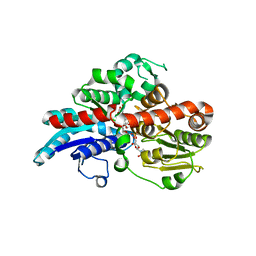 | | Crystal Structure of UDP-dependent glucosyltransferases (UGT) from Stevia rebaudiana in complex with UDP and rebaudioside A | | Descriptor: | (8alpha,9beta,10alpha,13alpha)-13-{[alpha-L-allopyranosyl-(1->2)-[beta-D-mannopyranosyl-(1->3)]-beta-D-allopyranosyl]oxy}kauran-18-oic acid, UDP-glycosyltransferase 76G1, URIDINE-5'-DIPHOSPHATE | | Authors: | Lee, S.G, Jez, J.M. | | Deposit date: | 2019-03-09 | | Release date: | 2019-06-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Molecular basis for branched steviol glucoside biosynthesis.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6OMX
 
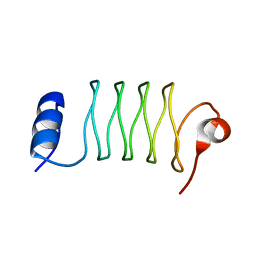 | |
6RY6
 
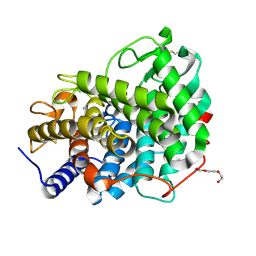 | |
7Q4Q
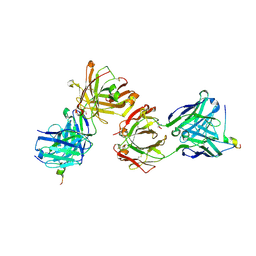 
 | | Magacizumab Fab fragment in complex with human LRG1 epitope | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, LRG1 epitope, Magacizumab heavy chain, ... | | Authors: | Gutierrez-Fernandez, J, Luecke, H. | | Deposit date: | 2021-11-01 | | Release date: | 2022-06-15 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis of human LRG1 recognition by Magacizumab, a humanized monoclonal antibody with therapeutic potential.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
6OWH
 
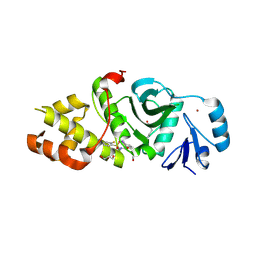 | | Crystal structure of MYST acetyltransferase domain in complex with inhibitor 92 | | Descriptor: | DIMETHYL SULFOXIDE, GLYCEROL, Histone acetyltransferase KAT8, ... | | Authors: | Hermans, S.J, Chung, M.C, Parker, M.W, Thomas, T, Baell, J.B. | | Deposit date: | 2019-05-09 | | Release date: | 2020-04-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of Acylsulfonohydrazide-Derived Inhibitors of the Lysine Acetyltransferase, KAT6A, as Potent Senescence-Inducing Anti-Cancer Agents.
J.Med.Chem., 63, 2020
|
|
6RMW
 
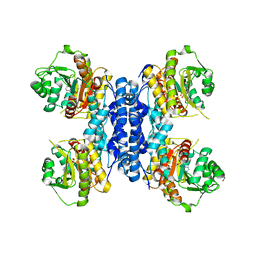 | | Structure of N-terminal truncated IMP bound Plasmodium falciparum IMP-nucleotidase | | Descriptor: | GLYCEROL, IMP-specific 5'-nucleotidase, putative, ... | | Authors: | Carrique, L, Ballut, L, Violot, S, Aghajari, N. | | Deposit date: | 2019-05-07 | | Release date: | 2020-07-15 | | Last modified: | 2025-10-01 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structure and catalytic regulation of Plasmodium falciparum IMP specific nucleotidase.
Nat Commun, 11, 2020
|
|
5Y9R
 
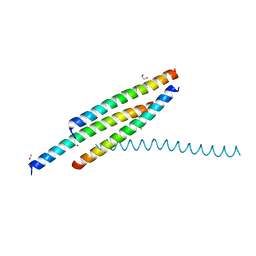 | |
9AV0
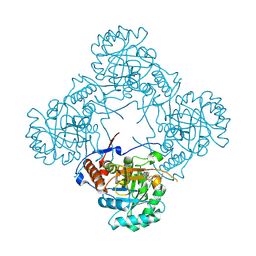 
 | | Crystal structure of S. aureus GuaB dCBS with inhibitor GNE2011 | | Descriptor: | 9-{(1R)-1-[(5P)-5-(4-chloro-1H-imidazol-2-yl)pyridin-3-yl]ethoxy}-1,4-dihydro-2H-pyrano[3,4-c]quinoline, INOSINIC ACID, Inosine-5'-monophosphate dehydrogenase | | Authors: | Harris, S.F, Wu, P. | | Deposit date: | 2024-03-01 | | Release date: | 2024-11-27 | | Last modified: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Discovery of GuaB inhibitors with efficacy against Acinetobacter baumannii infection.
Mbio, 15, 2024
|
|
6RFY
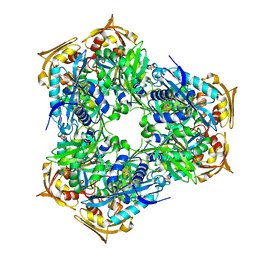 
 | | Crystal structure of Eis2 form Mycobacterium abscessus | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Eis2, SULFATE ION | | Authors: | Blaise, M, Kremer, L, Olieric, V, Alsarraf, H, Ung, K.L. | | Deposit date: | 2019-04-16 | | Release date: | 2019-07-10 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the aminoglycosides N-acetyltransferase Eis2 from Mycobacterium abscessus.
Febs J., 286, 2019
|
|
8UUK
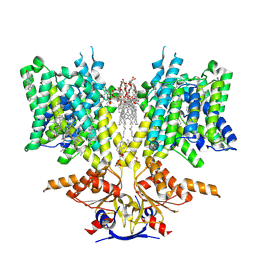 
 | | Pendrin in apo | | Descriptor: | 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, CHOLESTEROL, Lauryl Maltose Neopentyl Glycol, ... | | Authors: | Wang, L, Hoang, A, Zhou, M. | | Deposit date: | 2023-11-01 | | Release date: | 2024-02-07 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Mechanism of anion exchange and small-molecule inhibition of pendrin.
Nat Commun, 15, 2024
|
|
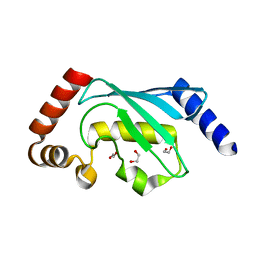
6QH3
 
 | |
9AUV
 
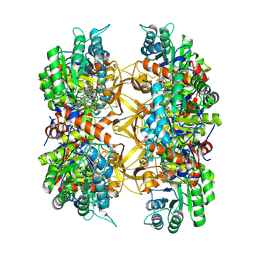 | | Crystal structure of A. baumannii GuaB dCBS with inhibitor GNE9123 | | Descriptor: | INOSINIC ACID, Inosine-5'-monophosphate dehydrogenase, N-(6-chloropyridin-3-yl)-N~2~-(1,4-dihydro-2H-pyrano[3,4-c]quinolin-9-yl)-L-alaninamide | | Authors: | Harris, S.F, Wu, P. | | Deposit date: | 2024-03-01 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Discovery of GuaB inhibitors with efficacy against Acinetobacter baumannii infection.
Mbio, 15, 2024
|
|
6R16
 
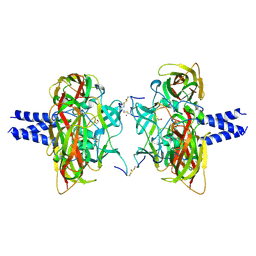 | | Crystal structure of the SUN1-KASH4 6:6 complex | | Descriptor: | 1,2-ETHANEDIOL, Nesprin-4, POTASSIUM ION, ... | | Authors: | Gurusaran, M, Davies, O.R. | | Deposit date: | 2019-03-13 | | Release date: | 2020-07-15 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | A molecular mechanism for LINC complex branching by structurally diverse SUN-KASH 6:6 assemblies.
Elife, 10, 2021
|
|
4Y88
 
 | |
6P41
 
 | |
6STW
 
 | |
6XP3
 
 | |
