6QDT
 
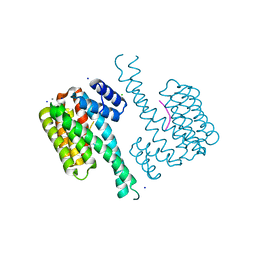 | | Crystal structure of 14-3-3sigma in complex with a RapGef2 pT740 phosphopeptide | | Descriptor: | 14-3-3 protein sigma, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Andrei, S.A, Kaplan, A, Fournier, A.E, Ottman, C. | | Deposit date: | 2019-01-02 | | Release date: | 2020-01-29 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.702 Å) | | Cite: | Polypharmacological Perturbation of the 14-3-3 Adaptor Protein Interactome Stimulates Neurite Outgrowth.
Cell Chem Biol, 27, 2020
|
|
4NAW
 
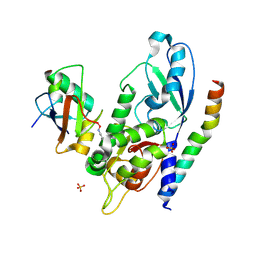 | | Crystal Structure of Human ATG12~ATG5-ATG16N in complex with a fragment of ATG3 | | Descriptor: | Autophagy protein 5, Autophagy-related protein 16-1, SODIUM ION, ... | | Authors: | Metlagel, Z, Otomo, C, Takaesu, G, Otomo, T. | | Deposit date: | 2013-10-22 | | Release date: | 2013-11-06 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.195 Å) | | Cite: | Structural basis of ATG3 recognition by the autophagic ubiquitin-like protein ATG12.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
5DW3
 
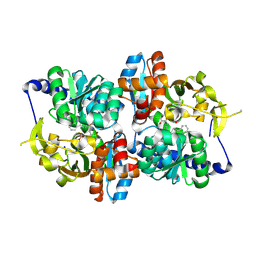 | |
5DVZ
 
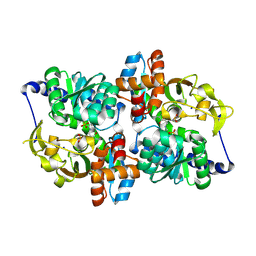 | | Holo TrpB from Pyrococcus furiosus | | Descriptor: | PHOSPHATE ION, SODIUM ION, Tryptophan synthase beta chain 1 | | Authors: | Buller, A.R, Arnold, F.H. | | Deposit date: | 2015-09-21 | | Release date: | 2016-02-03 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Directed evolution of the tryptophan synthase beta-subunit for stand-alone function recapitulates allosteric activation.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
6D1P
 
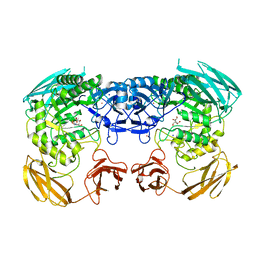 | | Apo structure of Bacteroides uniformis beta-glucuronidase 3 | | Descriptor: | GLYCEROL, Glycosyl hydrolases family 2, sugar binding domain protein, ... | | Authors: | Walton, W.G, Pellock, S.J, Redinbo, M.R. | | Deposit date: | 2018-04-12 | | Release date: | 2018-10-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Three structurally and functionally distinct beta-glucuronidases from the human gut microbeBacteroides uniformis.
J. Biol. Chem., 293, 2018
|
|
8SF6
 
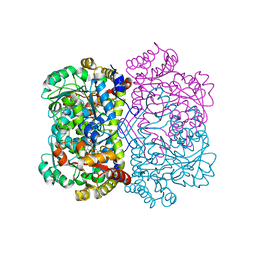 | | Promiscuous amino acid gamma synthase from Caldicellulosiruptor hydrothermalis in closed conformation | | Descriptor: | 2-(2-ETHOXYETHOXY)ETHANOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-{2-[2-2-(METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, ... | | Authors: | Buller, A.R, Zmich, A.P, Bingman, C.A. | | Deposit date: | 2023-04-10 | | Release date: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.702 Å) | | Cite: | Multiplexed Assessment of Promiscuous Non-Canonical Amino Acid Synthase Activity in a Pyridoxal Phosphate-Dependent Protein Family.
Acs Catalysis, 13, 2023
|
|
4NLV
 
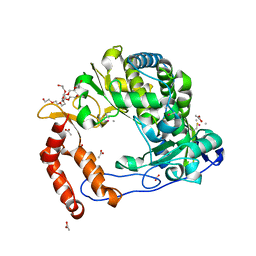 | | Poliovirus Polymerase - G289A/C290F Loop Mutant | | Descriptor: | ACETIC ACID, PENTAETHYLENE GLYCOL, RNA-directed RNA polymerase 3D-POL, ... | | Authors: | Sholders, A.J, Peersen, O.B. | | Deposit date: | 2013-11-14 | | Release date: | 2014-01-22 | | Last modified: | 2014-03-26 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Distinct conformations of a putative translocation element in poliovirus polymerase.
J.Mol.Biol., 426, 2014
|
|
4NLW
 
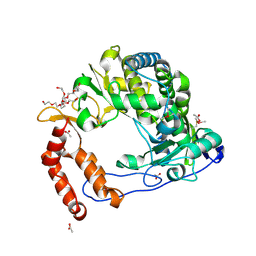 | | Poliovirus Polymerase - G289A/C290I Loop Mutant | | Descriptor: | ACETIC ACID, PENTAETHYLENE GLYCOL, RNA-directed RNA polymerase 3D-POL, ... | | Authors: | Sholders, A.J, Peersen, O.B. | | Deposit date: | 2013-11-14 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Distinct conformations of a putative translocation element in poliovirus polymerase.
J.Mol.Biol., 426, 2014
|
|
4NLY
 
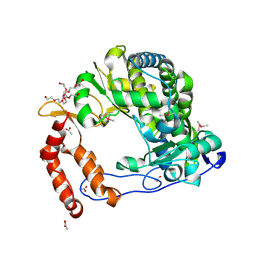 | | Poliovirus Polymerase - C290E Loop Mutant | | Descriptor: | ACETIC ACID, PENTAETHYLENE GLYCOL, RNA-directed RNA polymerase 3D-POL, ... | | Authors: | Sholders, A.J, Peersen, O.B. | | Deposit date: | 2013-11-14 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Distinct conformations of a putative translocation element in poliovirus polymerase.
J.Mol.Biol., 426, 2014
|
|
5MLO
 
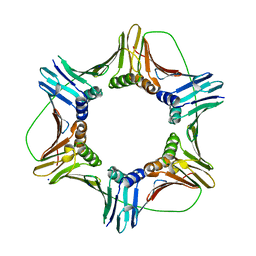 | |
5MXE
 
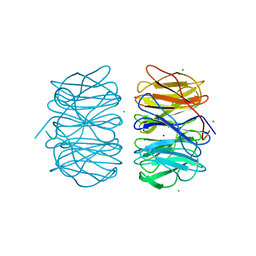 | | Photorhabdus asymbiotica lectin (PHL) in free form | | Descriptor: | CHLORIDE ION, Photorhabdus asymbiotica lectin PHL, SODIUM ION | | Authors: | Jancarikova, G, Houser, J, Demo, G, Wimmerova, M. | | Deposit date: | 2017-01-23 | | Release date: | 2017-08-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Characterization of novel bangle lectin from Photorhabdus asymbiotica with dual sugar-binding specificity and its effect on host immunity.
PLoS Pathog., 13, 2017
|
|
5DDR
 
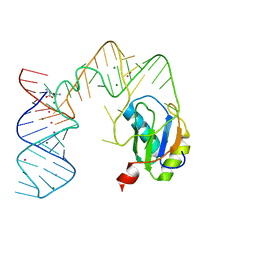 | | L-glutamine riboswitch bound with L-glutamine soaked with Cs+ | | Descriptor: | CESIUM ION, GLUTAMINE, L-glutamine riboswitch RNA (61-MER), ... | | Authors: | Ren, A, Patel, D.J. | | Deposit date: | 2015-08-25 | | Release date: | 2015-12-23 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.605 Å) | | Cite: | Structural and Dynamic Basis for Low-Affinity, High-Selectivity Binding of L-Glutamine by the Glutamine Riboswitch.
Cell Rep, 13, 2015
|
|
3K13
 
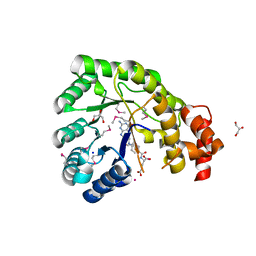 | | Structure of the pterin-binding domain MeTr of 5-methyltetrahydrofolate-homocysteine methyltransferase from Bacteroides thetaiotaomicron | | Descriptor: | 5-methyltetrahydrofolate-homocysteine methyltransferase, GLYCEROL, N-[4-({[(6S)-2-AMINO-4-HYDROXY-5-METHYL-5,6,7,8-TETRAHYDROPTERIDIN-6-YL]METHYL}AMINO)BENZOYL]-L-GLUTAMIC ACID, ... | | Authors: | Cuff, M.E, Li, H, Cobb, G, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-09-25 | | Release date: | 2009-12-22 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the pterin-binding domain MeTr of 5-methyltetrahydrofolate-homocysteine methyltransferase from Bacteroides thetaiotaomicron
TO BE PUBLISHED
|
|
7D0Z
 
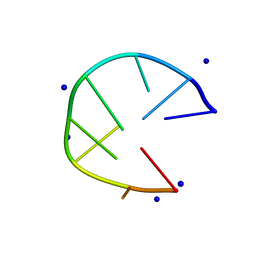 | |
5M66
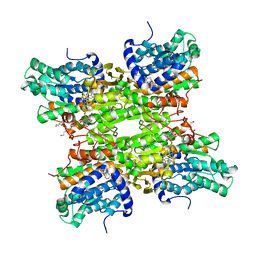 
 | |
5JXH
 
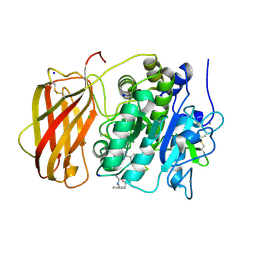 | | Structure the proprotein convertase furin in complex with meta-guanidinomethyl-Phac-RVR-Amba at 2.0 Angstrom resolution. | | Descriptor: | 2UC-ARG-VAL-ARG-00S, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Dahms, S.O, Arciniega, M, Steinmetzer, T, Huber, R, Than, M.E. | | Deposit date: | 2016-05-13 | | Release date: | 2016-10-05 | | Last modified: | 2024-07-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the unliganded form of the proprotein convertase furin suggests activation by a substrate-induced mechanism.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
5K0C
 
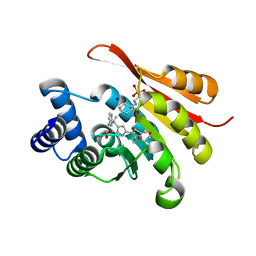 | | Crystal Structure of COMT in complex with 2,4-dimethyl-5-[3-(2-phenylpropan-2-yl)-1H-pyrazol-5-yl]-1,3-thiazole | | Descriptor: | 1,2-ETHANEDIOL, 2,4-dimethyl-5-[3-(2-phenylpropan-2-yl)-1H-pyrazol-5-yl]-1,3-thiazole, 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, ... | | Authors: | Ehler, A, Rodriguez-Sarmiento, R.M, Rudolph, M.G. | | Deposit date: | 2016-05-17 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Design of Potent and Druglike Nonphenolic Inhibitors for Catechol O-Methyltransferase Derived from a Fragment Screening Approach Targeting the S-Adenosyl-l-methionine Pocket.
J. Med. Chem., 59, 2016
|
|
5K0G
 
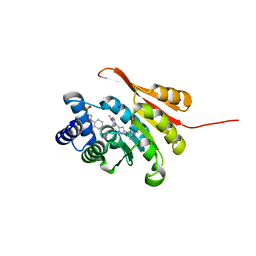 | | Crystal Structure of COMT in complex with 4-[5-[1-(4-methoxyphenyl)ethyl]-1H-pyrazol-3-yl]-1,3-dimethylpyrazole | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, 5-[(1S)-1-(4-methoxyphenyl)ethyl]-1',3'-dimethyl-1'H,2H-3,4'-bipyrazole, Catechol O-methyltransferase, ... | | Authors: | Ehler, A, rodriguez-Sarmiento, R.M, Rudolph, M.G. | | Deposit date: | 2016-05-17 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Design of Potent and Druglike Nonphenolic Inhibitors for Catechol O-Methyltransferase Derived from a Fragment Screening Approach Targeting the S-Adenosyl-l-methionine Pocket.
J. Med. Chem., 59, 2016
|
|
3KQD
 
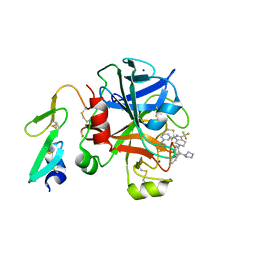 | | Factor xa in complex with the inhibitor 1-(3-(5-oxo-4,5- dihydro-1h-1,2,4-triazol-3-yl)phenyl)-6-(2'-(pyrrolidin-1- ylmethyl)biphenyl-4-yl)-3-(trifluoromethyl)-5,6-dihydro- 1h-pyrazolo[3,4-c]pyridin-7(4h)-one | | Descriptor: | 1-(3-(5-OXO-4,5-DIHYDRO-1H-1,2,4-TRIAZOL-3-YL)PHENYL)-6-(2'-(PYRROLIDIN-1-YLMETHYL)BIPHENYL-4-YL)-3-(TRIFLUOROMETHYL)-5,6-DIHYDRO-1H-PYRAZOLO[3,4-C]PYRIDIN-7(4H)-ONE, SODIUM ION, factor Xa heavy chain, ... | | Authors: | Sheriff, S. | | Deposit date: | 2009-11-17 | | Release date: | 2010-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Phenyltriazolinones as potent factor Xa inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
5MIM
 
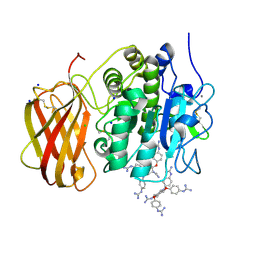 | | Xray structure of human furin bound with the 2,5-dideoxystreptamine derived small molecule inhibitor 1n | | Descriptor: | 1-[(1~{R},2~{R},4~{S},5~{S})-2,4-bis(4-carbamimidamidophenoxy)-5-[(4-carbamimidamidophenyl)amino]cyclohexyl]guanidine, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Dahms, S.O, Guan-Sheng, J, Than, M.E. | | Deposit date: | 2016-11-28 | | Release date: | 2017-05-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Studies Revealed Active Site Distortions of Human Furin by a Small Molecule Inhibitor.
ACS Chem. Biol., 12, 2017
|
|
5K0J
 
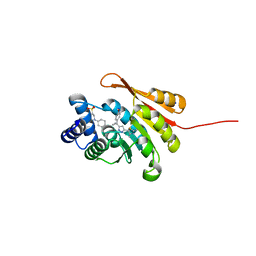 | | Crystal Structure of COMT in complex with 5-[5-[1-(4-methoxyphenyl)cyclopropyl]-1H-pyrazol-3-yl]-2,4-dimethyl-1,3-thiazole | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, 5-{3-[1-(4-methoxyphenyl)cyclopropyl]-1H-pyrazol-5-yl}-2,4-dimethyl-1,3-thiazole, Catechol O-methyltransferase, ... | | Authors: | Ehler, A, Rodriguez-Sarmiento, R.M, Rudolph, M.G. | | Deposit date: | 2016-05-17 | | Release date: | 2016-09-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Design of Potent and Druglike Nonphenolic Inhibitors for Catechol O-Methyltransferase Derived from a Fragment Screening Approach Targeting the S-Adenosyl-l-methionine Pocket.
J. Med. Chem., 59, 2016
|
|
2OQD
 
 | | Crystal Structure of BthTX-II | | Descriptor: | Phospholipase A2 | | Authors: | Correa, L.C, Marchi-Salvador, D.P, Cintra, A.C.O, Soares, A.M, Fontes, M.R.M. | | Deposit date: | 2007-01-31 | | Release date: | 2008-02-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Crystal structure of a myotoxic Asp49-phospholipase A(2) with low catalytic activity: Insights into Ca(2+)-independent catalytic mechanism.
Biochim.Biophys.Acta, 1784, 2008
|
|
5M4J
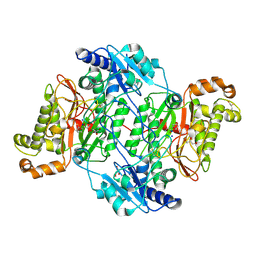 
 | | Crystal Structure of Wild-Type Human Prolidase with GlyPro ligand | | Descriptor: | GLYCEROL, GLYCINE, PROLINE, ... | | Authors: | Wilk, P, Weiss, M.S, Mueller, U, Dobbek, H. | | Deposit date: | 2016-10-18 | | Release date: | 2017-07-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Substrate specificity and reaction mechanism of human prolidase.
FEBS J., 284, 2017
|
|
3KQE
 
 | | Factor xa in complex with the inhibitor 3-methyl-1-(3-(5- oxo-4,5-dihydro-1h-1,2,4-triazol-3-yl)phenyl)-6-(2'- (pyrrolidin-1-ylmethyl)biphenyl-4-yl)-5,6-dihydro-1h- pyrazolo[3,4-c]pyridin-7(4h)-one | | Descriptor: | 3-METHYL-1-(3-(5-OXO-4,5-DIHYDRO-1H-1,2,4-TRIAZOL-3-YL)PHENYL)-6-(2'-(PYRROLIDIN-1-YLMETHYL)BIPHENYL-4-YL)-5,6-DIHYDRO-1H-PYRAZOLO[3,4-C]PYRIDIN-7(4H)-ONE, SODIUM ION, factor Xa heavy chain, ... | | Authors: | Sheriff, S. | | Deposit date: | 2009-11-17 | | Release date: | 2010-02-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Phenyltriazolinones as potent factor Xa inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
5LHL
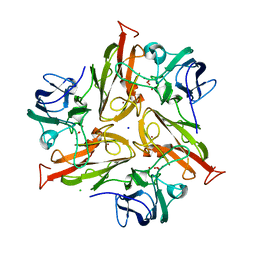 
 | |
