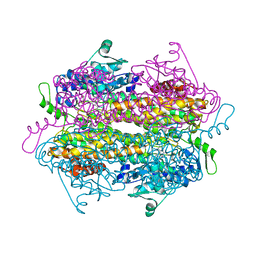
2QVE
 
 | |
3GPU
 
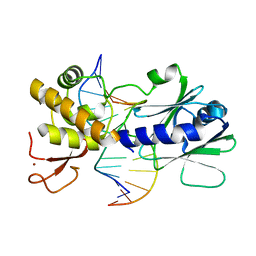 | | MutM encountering an intrahelical 8-oxoguanine (oxoG) lesion in EC4-loop deletion complex | | Descriptor: | DNA (5'-D(*AP*GP*GP*TP*AP*GP*AP*CP*TP*CP*GP*GP*AP*CP*GP*C)-3'), DNA (5'-D(*TP*GP*CP*GP*TP*CP*CP*(8OG)P*AP*GP*TP*CP*TP*AP*CP*C)-3'), DNA glycosylase, ... | | Authors: | Banerjee, A, Qi, Y, Verdine, G.L. | | Deposit date: | 2009-03-23 | | Release date: | 2009-11-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Encounter and extrusion of an intrahelical lesion by a DNA repair enzyme.
Nature, 462, 2009
|
|
3GQ3
 
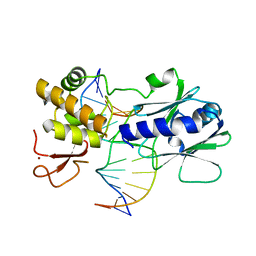 | | MutM encountering an intrahelical 8-oxoguanine (oxoG) lesion in EC5-loop deletion complex | | Descriptor: | DNA (5'-D(*AP*GP*GP*TP*AP*GP*AP*CP*CP*CP*GP*GP*AP*CP*GP*C)-3'), DNA (5'-D(*TP*GP*CP*GP*TP*CP*CP*GP*(8OG)P*GP*TP*CP*TP*AP*CP*C)-3'), DNA glycosylase, ... | | Authors: | Banerjee, A, Qi, Y, Verdine, G.L. | | Deposit date: | 2009-03-23 | | Release date: | 2009-11-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Encounter and extrusion of an intrahelical lesion by a DNA repair enzyme.
Nature, 462, 2009
|
|
2QW6
 
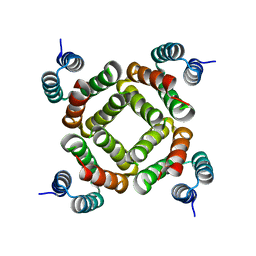 | | Crystal structure of the C-terminal domain of an AAA ATPase from Enterococcus faecium DO | | Descriptor: | AAA ATPase, central region | | Authors: | Bonanno, J.B, Rutter, M, Bain, K.T, Ozyurt, S, Smith, D, Wasserman, S, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2007-08-09 | | Release date: | 2007-08-28 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the C-terminal domain of an AAA ATPase from Enterococcus faecium DO.
To be Published
|
|
2QX6
 
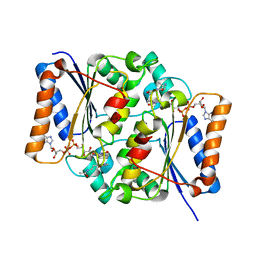 | | Crystal Structure of Quinone Reductase II | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, N-[2-(5-methoxy-1H-indol-3-yl)ethyl]acetamide, Ribosyldihydronicotinamide dehydrogenase [quinone], ... | | Authors: | Calamini, B, Santarsiero, B.D, Boutin, J.A, Mesecar, A.D. | | Deposit date: | 2007-08-10 | | Release date: | 2008-09-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Kinetic, thermodynamic and X-ray structural insights into the interaction of melatonin and analogues with quinone reductase 2.
Biochem.J., 413, 2008
|
|
3TKP
 
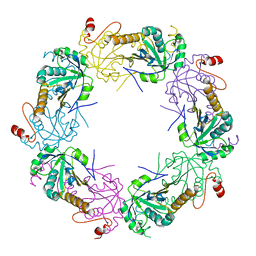 | | Crystal structure of full-length human peroxiredoxin 4 in the reduced form | | Descriptor: | Peroxiredoxin-4 | | Authors: | Wang, X, Wang, L, Wang, X, Sun, F, Wang, C.-C. | | Deposit date: | 2011-08-28 | | Release date: | 2011-10-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Structural insights into the peroxidase activity and inactivation of human peroxiredoxin 4
Biochem.J., 2011
|
|
3TLA
 
 | |
2R1Q
 
 | |
2QDH
 | | Fructose-1,6-bisphosphate aldolase from Leishmania mexicana in complex with mannitol-1,6-bisphosphate, a competitive inhibitor | | Descriptor: | D-MANNITOL-1,6-DIPHOSPHATE, Fructose-1,6-bisphosphate aldolase | | Authors: | Lafrance-Vanasse, J, Sygusch, J. | | Deposit date: | 2007-06-20 | | Release date: | 2007-08-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Carboxy-Terminus Recruitment Induced by Substrate Binding in Eukaryotic Fructose Bis-phosphate Aldolases
Biochemistry, 46, 2007
|
|
2QEW
 
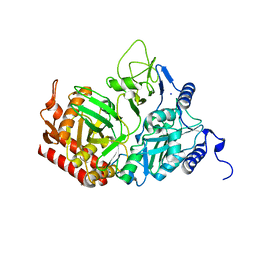 | | Rat cytosolic PEPCK, in complex with manganese ion. | | Descriptor: | MANGANESE (II) ION, Phosphoenolpyruvate carboxykinase, cytosolic [GTP], ... | | Authors: | Sullivan, S.M, Holyoak, T. | | Deposit date: | 2007-06-26 | | Release date: | 2007-08-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structures of rat cytosolic PEPCK: insight into the mechanism of phosphorylation and decarboxylation of oxaloacetic acid.
Biochemistry, 46, 2007
|
|
2R2B
 
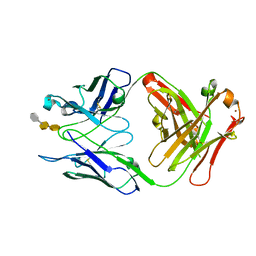 | | Crystal structure of S25-2 Fab in complex with Kdo analogues | | Descriptor: | 3-deoxy-alpha-D-manno-oct-2-ulopyranosonic acid-(2-4)-3-deoxy-alpha-D-manno-oct-2-ulopyranosonic acid-(2-4)-prop-2-en-1-yl 3-deoxy-alpha-D-manno-oct-2-ulopyranosidonic acid, Fab, antibody fragment (IgG1k), ... | | Authors: | Brooks, C.L, Evans, S.V. | | Deposit date: | 2007-08-24 | | Release date: | 2008-12-30 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Exploration of specificity in germline monoclonal antibody recognition of a range of natural and synthetic epitopes.
J.Mol.Biol., 377, 2008
|
|
3GW6
 
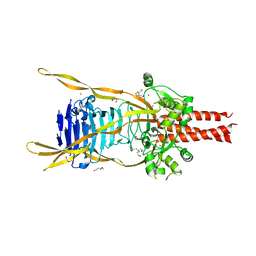 | | Intramolecular Chaperone | | Descriptor: | BROMIDE ION, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Schulz, E.C, Dickmanns, A, Ficner, R. | | Deposit date: | 2009-03-31 | | Release date: | 2010-02-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of an intramolecular chaperone mediating triple-beta-helix folding.
Nat.Struct.Mol.Biol., 17, 2010
|
|
3WHE
 
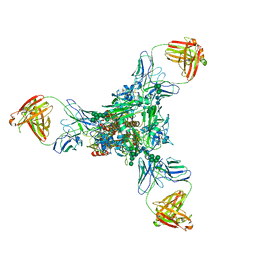 | | A new conserved neutralizing epitope at the globular head of hemagglutinin in H3N2 influenza viruses | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin, alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Fujii, Y, Sumida, T, Shirouzu, M, Yokoyama, S. | | Deposit date: | 2013-08-25 | | Release date: | 2014-04-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Conserved neutralizing epitope at globular head of hemagglutinin in H3N2 influenza viruses.
J.Virol., 88, 2014
|
|
3WIS
 
 | | Crystal structure of Burkholderia xenovorans DmrB in complex with FMN: A Cubic Protein Cage for Redox Transfer | | Descriptor: | FLAVIN MONONUCLEOTIDE, Putative dihydromethanopterin reductase (AfpA), SULFATE ION | | Authors: | Bobik, T.A, Cascio, D, Jorda, J, McNamara, D.E, Bustos, C, Wang, T.C, Rasche, M.E, Yeates, T.O. | | Deposit date: | 2013-09-25 | | Release date: | 2014-02-19 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.901 Å) | | Cite: | Structure of dihydromethanopterin reductase, a cubic protein cage for redox transfer
J.Biol.Chem., 289, 2014
|
|
3GZ6
 
 | |
3TQQ
 
 | | Structure of the methionyl-tRNA formyltransferase (fmt) from Coxiella burnetii | | Descriptor: | Methionyl-tRNA formyltransferase, POTASSIUM ION | | Authors: | Rudolph, M, Cheung, J, Franklin, M.C, Cassidy, M, Gary, E, Burshteyn, F, Love, J. | | Deposit date: | 2011-09-09 | | Release date: | 2011-09-28 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural genomics for drug design against the pathogen Coxiella burnetii.
Proteins, 83, 2015
|
|
2R47
 
 | | Crystal structure of MTH_862 protein of unknown function from Methanothermobacter thermautotrophicus | | Descriptor: | SULFATE ION, Uncharacterized protein MTH_862 | | Authors: | Osipiuk, J, Evdokimova, E, Kudritska, M, Savchenko, A, Edwards, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2007-08-30 | | Release date: | 2007-09-11 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Crystal structure of MTH_862 protein of unknown function from Methanothermobacter thermautotrophicus.
To be Published
|
|
3TS3
 
 | |
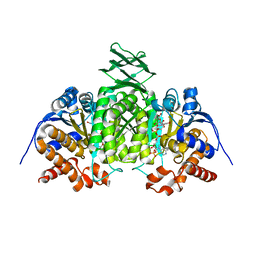
2QFX
 
 | |
3TT7
 
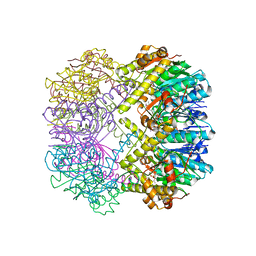 | |
3WO6
 
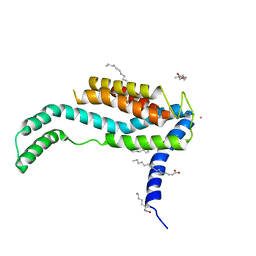 | | Crystal structure of YidC from Bacillus halodurans (form I) | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, CADMIUM ION, Membrane protein insertase YidC 2 | | Authors: | Kumazaki, K, Tsukazaki, T, Ishitani, R, Nureki, O. | | Deposit date: | 2013-12-20 | | Release date: | 2014-04-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.403 Å) | | Cite: | Structural basis of Sec-independent membrane protein insertion by YidC.
Nature, 509, 2014
|
|
2QI8
 
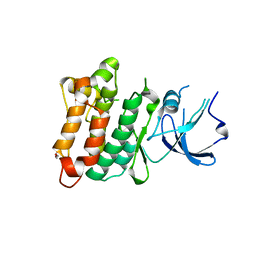 | | Crystal structure of drug resistant SRC kinase domain | | Descriptor: | GLYCEROL, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Michalczyk, A, Rode, H.B, Gruetter, C, Rauh, D. | | Deposit date: | 2007-07-03 | | Release date: | 2008-03-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Structural insights into how irreversible inhibitors can overcome drug resistance in EGFR.
Bioorg.Med.Chem., 16, 2008
|
|
3TUZ
 
 | | Inward facing conformations of the MetNI methionine ABC transporter: CY5 SeMet soak crystal form | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, D-methionine transport system permease protein metI, Methionine import ATP-binding protein MetN, ... | | Authors: | Johnson, E, Nguyen, P, Rees, D.C. | | Deposit date: | 2011-09-19 | | Release date: | 2011-11-30 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Inward facing conformations of the MetNI methionine ABC transporter: Implications for the mechanism of transinhibition.
Protein Sci., 21, 2012
|
|
3H0J
 
 | |
3H5L
 
 | | Crystal structure of a putative branched-chain amino acid ABC transporter from Silicibacter pomeroyi | | Descriptor: | putative Branched-chain amino acid ABC transporter | | Authors: | Bonanno, J.B, Freeman, J, Bain, K.T, Iizuka, M, Sampathkumar, P, Wasserman, S, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-04-22 | | Release date: | 2009-05-05 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of a putative branched-chain amino acid ABC transporter from Silicibacter pomeroyi
To be Published
|
|
