2P6U
 
 | |
3GJJ
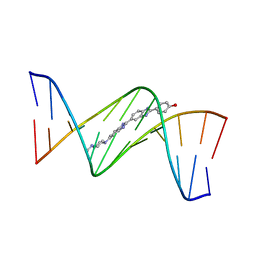 
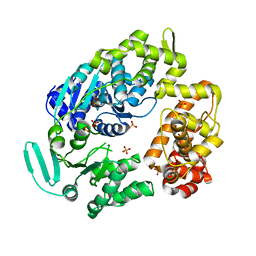
 | | crystal structure of a DNA duplex containing 7,8-dihydropyridol[2,3-d]pyrimidin-2-one | | Descriptor: | 2'-(4-HYDROXYPHENYL)-5-(4-METHYL-1-PIPERAZINYL)-2,5'-BI-BENZIMIDAZOLE, 5'-D(*CP*GP*CP*GP*AP*AP*TP*(B7C)P*CP*GP*CP*G)-3' | | Authors: | Takenaka, A, Juan, E.C.M, Shimizu, S, Haraguchi, T, Xiao, M, Kurose, T, Ohkubo, A, Sekine, M, Shibata, T, Millington, C.L, Williams, D.M. | | Deposit date: | 2009-03-09 | | Release date: | 2010-03-31 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Insights into the stabilizing contributions of bicyclic cytosine analogues: crystal structures of DNA duplexes containing 7,8-dihydropyridol[2,3-d]pyrimidin-2-one
To be Published
|
|
4HPZ
 
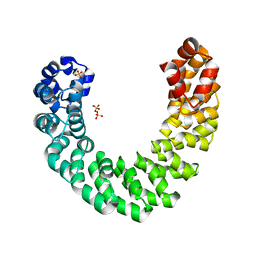 | |
3KTZ
 
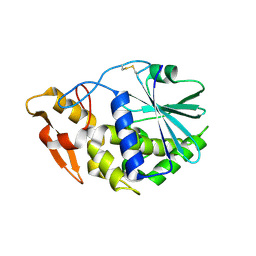 | | Structure of GAP31 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Ribosome-inactivating protein gelonin | | Authors: | Kong, X.-P. | | Deposit date: | 2009-11-26 | | Release date: | 2010-01-26 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A new activity of anti-HIV and anti-tumor protein GAP31: DNA adenosine glycosidase--structural and modeling insight into its functions.
Biochem.Biophys.Res.Commun., 391, 2010
|
|
4QIO
 
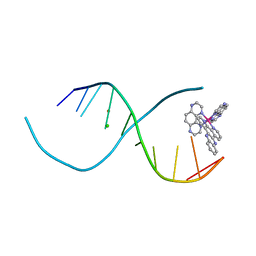 | | Lambda-[Ru(TAP)2(dppz)]2+ bound to d(TCGGCGCCIA) at high resolution | | Descriptor: | 5'-D(*TP*CP*GP*GP*CP*GP*CP*CP*IP*A)-3', BARIUM ION, CHLORIDE ION, ... | | Authors: | Gurung, S.P, Hall, J.P, Cardin, C.J. | | Deposit date: | 2014-06-01 | | Release date: | 2015-06-03 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (0.95 Å) | | Cite: | Inosine Can Increase DNA's Susceptibility to Photo-oxidation by a Ru II Complex due to Structural Change in the Minor Groove.
Chemistry, 23, 2017
|
|
3SM4
 
 | |
2GCJ
 
 | | Crystal Structure of the Pob3 middle domain | | Descriptor: | Hypothetical 63.0 kDa protein in DAK1-ORC1 intergenic region | | Authors: | VanDemark, A.P. | | Deposit date: | 2006-03-14 | | Release date: | 2006-05-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The Structure of the yFACT Pob3-M Domain, Its Interaction with the DNA Replication Factor RPA, and a Potential Role in Nucleosome Deposition.
Mol.Cell, 22, 2006
|
|
4B5G
 
 | | Substrate bound Neisseria AP endonuclease in absence of metal ions (crystal form 2) | | Descriptor: | 5'-D(*CP*GP*AP*TP*GP*GP*GP*TP*AP*GP*CP)-3', 5'-D(*GP*CP*TP*AP*CP*(3DR)P*CP*AP*TP*CP*GP)-3', EXODEOXYRIBONUCLEASE | | Authors: | Lu, D, Silhan, J, MacDonald, J.T, Carpenter, E.P, Jensen, K, Tang, C.M, Baldwin, G.S, Freemont, P.S. | | Deposit date: | 2012-08-03 | | Release date: | 2012-10-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural basis for the recognition and cleavage of abasic DNA in Neisseria meningitidis.
Proc. Natl. Acad. Sci. U.S.A., 109, 2012
|
|
1SXQ
 | |
4B5H
 
 | | Substate bound inactive mutant of Neisseria AP endonuclease in presence of metal ions | | Descriptor: | 5'-D(*CP*GP*AP*TP*GP*GP*GP*TP*AP*GP*CP)-3', 5'-D(*GP*CP*TP*AP*CP*3DRP*CP*AP*TP*CP*GP)-3', MANGANESE (II) ION, ... | | Authors: | Lu, D, Silhan, J, MacDonald, J.T, Carpenter, E.P, Jensen, K, Tang, C.M, Baldwin, G.S, Freemont, P.S. | | Deposit date: | 2012-08-03 | | Release date: | 2012-10-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structural basis for the recognition and cleavage of abasic DNA in Neisseria meningitidis.
Proc. Natl. Acad. Sci. U.S.A., 109, 2012
|
|
1TL6
 
 | | Solution structure of T4 bacteriphage AsiA monomer | | Descriptor: | 10 kDa anti-sigma factor | | Authors: | Lambert, L.J, Wei, Y, Schirf, V, Demeler, B, Werner, M.H. | | Deposit date: | 2004-06-09 | | Release date: | 2005-06-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | T4 AsiA blocks DNA recognition by remodeling sigma70 region 4
Embo J., 23, 2004
|
|
3GPL
 
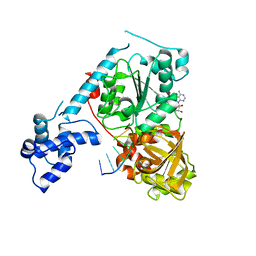 | | Crystal structure of the ternary complex of RecD2 with DNA and ADPNP | | Descriptor: | 5'-D(*T*TP*TP*TP*TP*TP*TP*T)-3', Exodeoxyribonuclease V, subunit RecD, ... | | Authors: | Saikrishnan, K, Cook, N, Wigley, D.B. | | Deposit date: | 2009-03-23 | | Release date: | 2009-06-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mechanistic basis of 5'-3' translocation in SF1B helicases.
Cell(Cambridge,Mass.), 137, 2009
|
|
5C15
 
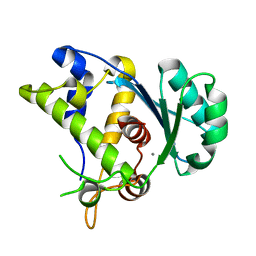 | |
4B5M
 
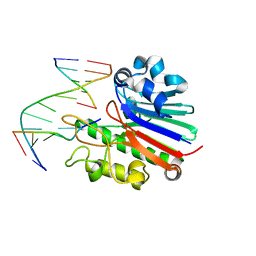 | | Neisseria AP endonuclease bound to the substrate with a cytosine orphan base | | Descriptor: | 5'-D(*3DRP*CP*AP*TP*CP*GP)-3', 5'-D(*CP*GP*AP*TP*GP*CP*GP*TP*AP*GP*CP)-3', 5'-D(*GP*CP*TP*AP*CP)-3', ... | | Authors: | Lu, D, Silhan, J, MacDonald, J.T, Carpenter, E.P, Jensen, K, Tang, C.M, Baldwin, G.S, Freemont, P.S. | | Deposit date: | 2012-08-06 | | Release date: | 2012-10-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.758 Å) | | Cite: | Structural basis for the recognition and cleavage of abasic DNA in Neisseria meningitidis.
Proc. Natl. Acad. Sci. U.S.A., 109, 2012
|
|
5C12
 
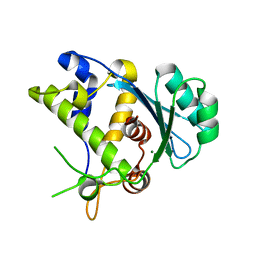 | |
4Y18
 
 | |
5CD4
 
 | | The Type IE CRISPR Cascade complex from E. coli, with two assemblies in the asymmetric unit arranged back-to-back | | Descriptor: | CRISPR system Cascade subunit CasA, CRISPR system Cascade subunit CasB, CRISPR system Cascade subunit CasC, ... | | Authors: | Jackson, R.N, Golden, S.M, Carter, J, Wiedenheft, B. | | Deposit date: | 2015-07-03 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Mechanism of CRISPR-RNA guided recognition of DNA targets in Escherichia coli.
Nucleic Acids Res., 43, 2015
|
|
4Y2G
 
 | |
1SXP
 
 | | BGT in complex with a 13mer DNA containing a central A:G mismatch | | Descriptor: | 5'-D(*A*AP*TP*AP*CP*TP*AP*AP*GP*AP*TP*AP*G)-3', 5'-D(*CP*TP*AP*TP*CP*TP*GP*AP*GP*TP*AP*TP*T)-3', DNA beta-glucosyltransferase, ... | | Authors: | Lariviere, L, Morera, S. | | Deposit date: | 2004-03-31 | | Release date: | 2004-06-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural evidence of a passive base flipping mechanism for {beta}-Glucosyltransferase
J.Biol.Chem., 279, 2004
|
|
5C2F
 
 | | K428A mutant nuclease domain of the large terminase subunit gp2 of bacterial virus Sf6 with Manganese and beta-thujaplicinol | | Descriptor: | 2,7-dihydroxy-4-(propan-2-yl)cyclohepta-2,4,6-trien-1-one, Gene 2 protein, MANGANESE (II) ION | | Authors: | Zhao, H, Tang, L. | | Deposit date: | 2015-06-15 | | Release date: | 2015-10-21 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Two distinct modes of metal ion binding in the nuclease active site of a viral DNA-packaging terminase: insight into the two-metal-ion catalytic mechanism.
Nucleic Acids Res., 43, 2015
|
|
1PGZ
 
 | |
5CR5
 
 | |
6IFO
 
 | | Crystal structure of AcrIIA2-SpyCas9-sgRNA ternary complex | | Descriptor: | AcrIIA2, CRISPR-associated endonuclease Cas9/Csn1, RNA (99-MER) | | Authors: | Liu, L, Wang, Y. | | Deposit date: | 2018-09-20 | | Release date: | 2019-01-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.313 Å) | | Cite: | Phage AcrIIA2 DNA Mimicry: Structural Basis of the CRISPR and Anti-CRISPR Arms Race.
Mol. Cell, 73, 2019
|
|
2J85
 
 | |
2IG0
 
 | | Structure of 53BP1/methylated histone peptide complex | | Descriptor: | Dimethylated Histone H4-K20 peptide, SULFATE ION, Tumor suppressor p53-binding protein 1 | | Authors: | Mer, G. | | Deposit date: | 2006-09-22 | | Release date: | 2007-01-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Basis for the Methylation State-Specific Recognition of Histone H4-K20 by 53BP1 and Crb2 in DNA Repair.
Cell(Cambridge,Mass.), 127, 2006
|
|
