2F9L
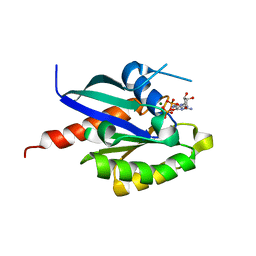 
 | | 3D structure of inactive human Rab11b GTPase | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, RAB11B, ... | | Authors: | Scapin, S.M.N, Guimaraes, B.G, Zanchin, N.I.T. | | Deposit date: | 2005-12-06 | | Release date: | 2006-04-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | The crystal structure of the small GTPase Rab11b reveals critical differences relative to the Rab11a isoform.
J.Struct.Biol., 154, 2006
|
|
2FLK
 
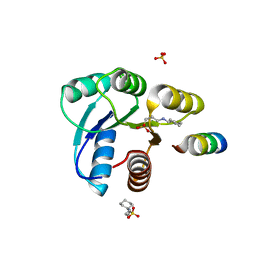 | | Crystal structure of CheY in complex with CheZ(200-214) solved from a F432 crystal grown in CAPS (pH 10.5) | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, C-terminal 15-mer from Chemotaxis protein cheZ, Chemotaxis protein cheY, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-06 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation.
J.Mol.Biol., 359, 2006
|
|
2EU9
 
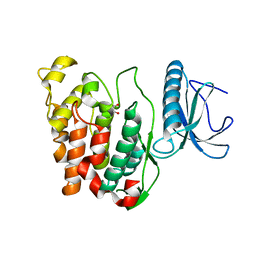 | | Crystal Structure of CLK3 | | Descriptor: | 1,2-ETHANEDIOL, Dual specificity protein kinase CLK3 | | Authors: | Papagrigoriou, E, Rellos, P, Das, S, Ugochukwu, E, Turnbull, A, von Delft, F, Bunkoczi, G, Sobott, F, Bullock, A, Fedorov, O, Gileadi, C, Savitsky, P, Edwards, A, Aerrowsmith, C, Weigelt, J, Sundstrom, M, Knapp, S. | | Deposit date: | 2005-10-28 | | Release date: | 2005-11-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Kinase domain insertions define distinct roles of CLK kinases in SR protein phosphorylation.
Structure, 17, 2009
|
|
2EVP
 
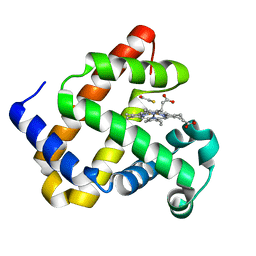 | | The Structures of Thiolate- and Carboxylate-Ligated Ferric H93G Myoglobin: Models for Cytochrome P450 and for Oxyanion-Bound Heme Proteins | | Descriptor: | BETA-MERCAPTOETHANOL, Myoglobin, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Qin, J, Perera, R, Lovelace, L.L, Dawson, J.H, Lebioda, L. | | Deposit date: | 2005-10-31 | | Release date: | 2006-03-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of Thiolate- and Carboxylate-Ligated Ferric H93G Myoglobin: Models for Cytochrome P450 and for Oxyanion-Bound Heme Proteins(,).
Biochemistry, 45, 2006
|
|
2NNP
 
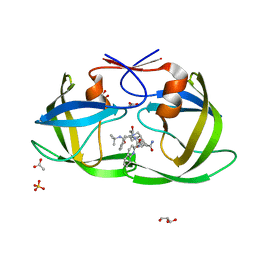 | | Crystal structure analysis of HIV-1 protease mutant I84V with a inhibitor saquinavir | | Descriptor: | (2S)-N-[(2S,3R)-4-[(2S,3S,4aS,8aS)-3-(tert-butylcarbamoyl)-3,4,4a,5,6,7,8,8a-octahydro-1H-isoquinolin-2-yl]-3-hydroxy-1 -phenyl-butan-2-yl]-2-(quinolin-2-ylcarbonylamino)butanediamide, ACETIC ACID, GLYCEROL, ... | | Authors: | Tie, Y, Kovalevsky, A.Y, Boross, P, Wang, Y.F, Ghosh, A.K, Tozser, J, Harrison, R.W, Weber, I.T. | | Deposit date: | 2006-10-24 | | Release date: | 2007-03-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Atomic resolution crystal structures of HIV-1 protease and mutants V82A and I84V with saquinavir.
Proteins, 67, 2007
|
|
2ETR
 
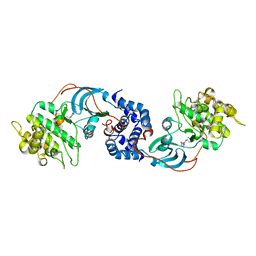 | | Crystal Structure of ROCK I bound to Y-27632 | | Descriptor: | (R)-TRANS-4-(1-AMINOETHYL)-N-(4-PYRIDYL) CYCLOHEXANECARBOXAMIDE, Rho-associated protein kinase 1 | | Authors: | Jacobs, M. | | Deposit date: | 2005-10-27 | | Release date: | 2005-11-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Structure of Dimeric ROCK I Reveals the Mechanism for Ligand Selectivity.
J.Biol.Chem., 281, 2006
|
|
4IX6
 
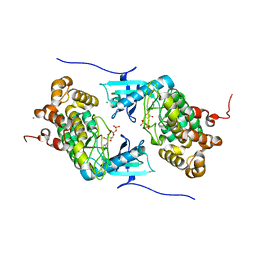 | | Crystal structure of a Stt7 homolog from Micromonas algae soaked with ATP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, MsStt7d protein | | Authors: | Guo, J, Wei, X, Li, M, Pan, X, Chang, W, Liu, Z. | | Deposit date: | 2013-01-24 | | Release date: | 2013-10-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of the catalytic domain of a state transition kinase homolog from Micromonas algae
Protein Cell, 4, 2013
|
|
2EXV
 
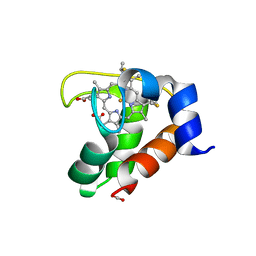 | | Crystal structure of the F7A mutant of the cytochrome c551 from Pseudomonas aeruginosa | | Descriptor: | ACETIC ACID, Cytochrome c-551, HEME C | | Authors: | Bonivento, D, Di Matteo, A, Borgia, A, Travaglini-Allocatelli, C, Brunori, M. | | Deposit date: | 2005-11-09 | | Release date: | 2006-02-07 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Unveiling a Hidden Folding Intermediate in c-Type Cytochromes by Protein Engineering
J.Biol.Chem., 281, 2006
|
|
2EVK
 
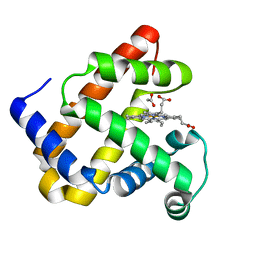 | | The Structures of Thiolate- and Carboxylate-Ligated Ferric H93G Myoglobin: Models for Cytochrome P450 and for Oxyanion-Bound Heme Proteins | | Descriptor: | ACETIC ACID, Myoglobin, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Qin, J, Perera, R, Lovelace, L.L, Dawson, J.H, Lebioda, L. | | Deposit date: | 2005-10-31 | | Release date: | 2006-04-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structures of thiolate- and carboxylate-ligated ferric H93G myoglobin: models for cytochrome P450 and for oxyanion-bound heme proteins.
Biochemistry, 45, 2006
|
|
2EYV
 
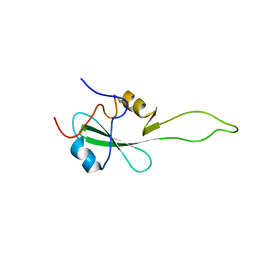 | | SH2 domain of CT10-Regulated Kinase | | Descriptor: | v-crk sarcoma virus CT10 oncogene homolog isoform a | | Authors: | Kobashigawa, Y, Tanaka, S, Inagaki, F. | | Deposit date: | 2005-11-10 | | Release date: | 2006-11-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the transforming activity of human cancer-related signaling adaptor protein CRK.
Nat.Struct.Mol.Biol., 14, 2007
|
|
4IX4
 
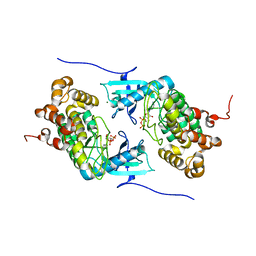 | | Crystal structure of a Stt7 homolog from Micromonas algae in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, MsStt7d protein | | Authors: | Guo, J, Wei, X, Li, M, Pan, X, Chang, W, Liu, Z. | | Deposit date: | 2013-01-24 | | Release date: | 2013-10-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.499 Å) | | Cite: | Structure of the catalytic domain of a state transition kinase homolog from Micromonas algae
Protein Cell, 4, 2013
|
|
4IZY
 
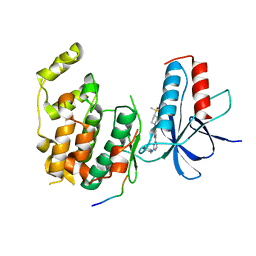 | |
2EWY
 
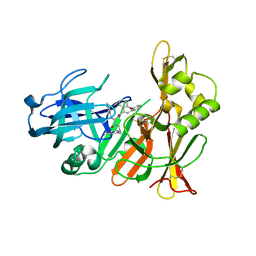 | |
2N82
 
 | |
2NNJ
 
 | | CYP2C8dH complexed with felodipine | | Descriptor: | Cytochrome P450 2C8, FELODIPINE, PALMITIC ACID, ... | | Authors: | Schoch, G.A, Yano, J.K, Stout, C.D, Johnson, E.F. | | Deposit date: | 2006-10-24 | | Release date: | 2007-10-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Determinants of cytochrome P450 2C8 substrate binding: structures of complexes with montelukast, troglitazone, felodipine, and 9-cis-retinoic acid.
J.Biol.Chem., 283, 2008
|
|
2JE4
 
 | | Atomic-resolution crystal structure of chemically-synthesized HIV-1 protease in complex with JG-365 | | Descriptor: | ACETATE ION, GLYCEROL, INHIBITOR MOLECULE JG365, ... | | Authors: | Malito, E, Johnson, E.C.B, Tang, W.J. | | Deposit date: | 2007-01-15 | | Release date: | 2007-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Modular Total Chemical Synthesis of a Human Immunodeficiency Virus Type 1 Protease.
J.Am.Chem.Soc., 129, 2007
|
|
2JM4
 
 | |
2NMZ
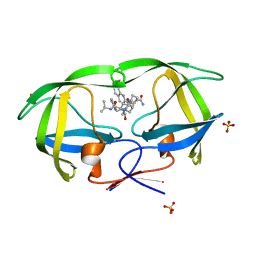 
 | | Crystal structure analysis of HIV-1 protease mutant V82A with a inhibitor saquinavir | | Descriptor: | (2S)-N-[(2S,3R)-4-[(2S,3S,4aS,8aS)-3-(tert-butylcarbamoyl)-3,4,4a,5,6,7,8,8a-octahydro-1H-isoquinolin-2-yl]-3-hydroxy-1 -phenyl-butan-2-yl]-2-(quinolin-2-ylcarbonylamino)butanediamide, PROTEASE, SULFATE ION | | Authors: | Tie, Y, Kovalevsky, A.Y, Boross, P, Wang, Y.F, Ghosh, A.K, Tozser, J, Harrison, R.W, Weber, I.T. | | Deposit date: | 2006-10-23 | | Release date: | 2007-03-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (0.97 Å) | | Cite: | Atomic resolution crystal structures of HIV-1 protease and mutants V82A and I84V with saquinavir.
Proteins, 67, 2007
|
|
2NNK
 
 | | Crystal structure analysis of HIV-1 protease mutant I84V with a inhibitor saquinavir | | Descriptor: | (2S)-N-[(2S,3R)-4-[(2S,3S,4aS,8aS)-3-(tert-butylcarbamoyl)-3,4,4a,5,6,7,8,8a-octahydro-1H-isoquinolin-2-yl]-3-hydroxy-1 -phenyl-butan-2-yl]-2-(quinolin-2-ylcarbonylamino)butanediamide, ACETIC ACID, CHLORIDE ION, ... | | Authors: | Tie, Y, Kovalevsky, A.Y, Boross, P, Wang, Y.F, Ghosh, A.K, Tozser, J, Harrison, R.W, Weber, I.T. | | Deposit date: | 2006-10-24 | | Release date: | 2007-03-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Atomic resolution crystal structures of HIV-1 protease and mutants V82A and I84V with saquinavir.
Proteins, 67, 2007
|
|
2P48
 
 | | Complex of a camelid single-domain vhh antibody fragment with RNASE A at 2.3A resolution: SE5B-tetra crystal form with five se-met sites (L4M, M34, M51, F68M, M83) in vhh scaffold. | | Descriptor: | ANTIBODY CAB-RN05, Ribonuclease pancreatic, SULFATE ION | | Authors: | Tereshko, V, Uysal, S, Koide, A, Margalef, K, Koide, S, Kossiakoff, A.A. | | Deposit date: | 2007-03-11 | | Release date: | 2008-03-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Toward chaperone-assisted crystallography: protein engineering enhancement of crystal packing and X-ray phasing capabilities of a camelid single-domain antibody (VHH) scaffold
Protein Sci., 17, 2008
|
|
2JV1
 
 | |
2JVK
 
 | |
2JW4
 
 | | NMR solution structure of the N-terminal SH3 domain of human Nckalpha | | Descriptor: | Cytoplasmic protein NCK1 | | Authors: | Santiveri, C.M, Borroto, A, Simon, L, Rico, M, Ortiz, A.R, Alarcon, B, Jimenez, M. | | Deposit date: | 2007-10-05 | | Release date: | 2008-08-26 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Interaction between the N-terminal SH3 domain of Nckalpha and CD3epsilon-derived peptides: Non-canonical and canonical recognition motifs
BIOCHEM.BIOPHYS.ACTA PROTEINS & PROTEOMICS, 1794, 2009
|
|
2JZW
 
 | | How the HIV-1 nucleocapsid protein binds and destabilises the (-)primer binding site during reverse transcription | | Descriptor: | DNA (5'-D(*DGP*DTP*DCP*DCP*DCP*DTP*DGP*DTP*DTP*DCP*DGP*DGP*DGP*DC)-3'), HIV-1 nucleocapsid protein NCp7(12-55), ZINC ION | | Authors: | Bourbigot, S, Ramalanjaona, N, Salgado, G.F.J, Mely, Y, Roques, B.P, Bouaziz, S, Morellet, N. | | Deposit date: | 2008-01-21 | | Release date: | 2009-01-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | How the HIV-1 nucleocapsid protein binds and destabilises the (-)primer binding site during reverse transcription.
J.Mol.Biol., 383, 2008
|
|
2K11
 
 | | Solution structure of human pancreatic ribonuclease | | Descriptor: | Pancreatic Ribonuclease | | Authors: | Kover, K.E, Bruix, M, Santoro, J, Batta, G, Laurents, D.V, Rico, M. | | Deposit date: | 2008-02-20 | | Release date: | 2008-06-03 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The solution structure and dynamics of human pancreatic ribonuclease determined by NMR spectroscopy provide insight into its remarkable biological activities and inhibition.
J.Mol.Biol., 379, 2008
|
|
