7PJ1
 
 | |
7PXH
 
 | | Emodepside-bound Drosophila Slo channel | | Descriptor: | (3~{S},6~{R},9~{S},12~{R},15~{S},18~{R},21~{S},24~{R})-4,6,10,16,18,22-hexamethyl-3,9,15,21-tetrakis(2-methylpropyl)-12,24-bis[(4-morpholin-4-ylphenyl)methyl]-1,7,13,19-tetraoxa-4,10,16,22-tetrazacyclotetracosane-2,5,8,11,14,17,20,23-octone, (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSAN-1-AMINIUM 4-OXIDE, CALCIUM ION, ... | | Authors: | Raisch, T, Brockmann, A, Ebbinghaus-Kintscher, U, Freigang, J, Gutbrod, O, Kubicek, J, Maertens, B, Hofnagel, O, Raunser, S. | | Deposit date: | 2021-10-08 | | Release date: | 2021-12-15 | | Last modified: | 2021-12-22 | | Method: | ELECTRON MICROSCOPY (2.59 Å) | | Cite: | Small molecule modulation of the Drosophila Slo channel elucidated by cryo-EM.
Nat Commun, 12, 2021
|
|
7PXG
 
 | | Verruculogen-bound Drosophila Slo channel | | Descriptor: | (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSAN-1-AMINIUM 4-OXIDE, CALCIUM ION, CHOLESTEROL, ... | | Authors: | Raisch, T, Brockmann, A, Ebbinghaus-Kintscher, U, Freigang, J, Gutbrod, O, Kubicek, J, Maertens, B, Hofnagel, O, Raunser, S. | | Deposit date: | 2021-10-08 | | Release date: | 2021-12-15 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (2.73 Å) | | Cite: | Small molecule modulation of the Drosophila Slo channel elucidated by cryo-EM.
Nat Commun, 12, 2021
|
|
7PXF
 
 | | Ca2+ free Drosophila Slo channel | | Descriptor: | Isoform J of Calcium-activated potassium channel slowpoke, MAGNESIUM ION, POTASSIUM ION | | Authors: | Raisch, T, Brockmann, A, Ebbinghaus-Kintscher, U, Freigang, J, Gutbrod, O, Kubicek, J, Maertens, B, Hofnagel, O, Raunser, S. | | Deposit date: | 2021-10-08 | | Release date: | 2021-12-15 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (2.68 Å) | | Cite: | Small molecule modulation of the Drosophila Slo channel elucidated by cryo-EM.
Nat Commun, 12, 2021
|
|
7PXE
 
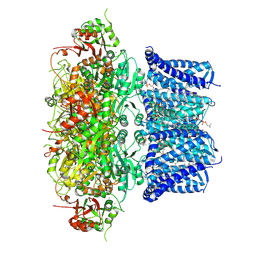 | | Ca2+ bound Drosophila Slo channel | | Descriptor: | (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSAN-1-AMINIUM 4-OXIDE, CALCIUM ION, CHOLESTEROL, ... | | Authors: | Raisch, T, Brockmann, A, Ebbinghaus-Kintscher, U, Freigang, J, Gutbrod, O, Kubicek, J, Maertens, B, Hofnagel, O, Raunser, S. | | Deposit date: | 2021-10-08 | | Release date: | 2021-12-15 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (2.38 Å) | | Cite: | Small molecule modulation of the Drosophila Slo channel elucidated by cryo-EM.
Nat Commun, 12, 2021
|
|
2NRP
 
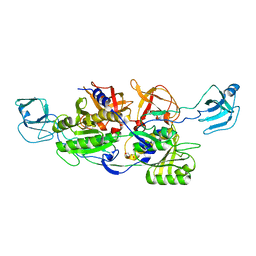 | | MoeA R350A | | Descriptor: | GLYCEROL, Molybdopterin biosynthesis protein moeA | | Authors: | Nicolas, J, Xiang, S, Schindelin, H, Rajagopalan, K.V. | | Deposit date: | 2006-11-02 | | Release date: | 2007-01-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Mutational Analysis of Escherichia coli MoeA: Two Functional Activities Map to the Active Site Cleft.
Biochemistry, 46, 2007
|
|
5T9Y
 
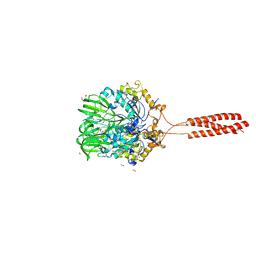 | | Crystal structure of the infectious salmon anemia virus (ISAV) hemagglutinin-esterase protein | | Descriptor: | FORMIC ACID, GLYCEROL, HE protein, ... | | Authors: | Cook, J.D, Sultana, A, Lee, J.E. | | Deposit date: | 2016-09-09 | | Release date: | 2017-03-22 | | Last modified: | 2020-01-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of the infectious salmon anemia virus receptor complex illustrates a unique binding strategy for attachment.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
7PHU
 
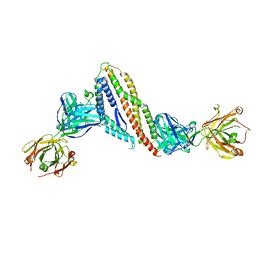 | |
2NRO
 | | MoeA K279Q | | Descriptor: | Molybdopterin biosynthesis protein moeA | | Authors: | Nicolas, J, Xiang, S, Schindelin, H, Rajagopalan, K.V. | | Deposit date: | 2006-11-02 | | Release date: | 2007-01-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mutational Analysis of Escherichia coli MoeA: Two Functional Activities Map to the Active Site Cleft.
Biochemistry, 46, 2007
|
|
5YI7
 
 | |
7Q4I
 
 | | Crystal structure of DmC1GalT1 in complex with UDP-Mn2+ and the APD-TGalNAc-RP | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-alpha-D-galactopyranose, Glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase 1, ... | | Authors: | Gonzalez-Ramirez, A.M, Coelho, H, Companon, I, Grosso, A.S, Yang, Z, Narimatsu, Y, Clausen, H, Marcelo, F, Corzana, F, Hurtado-Guerrero, R. | | Deposit date: | 2021-10-31 | | Release date: | 2022-04-13 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for the synthesis of the core 1 structure by C1GalT1.
Nat Commun, 13, 2022
|
|
2PG1
 
 | |
4C5H
 
 | |
2QDI
 
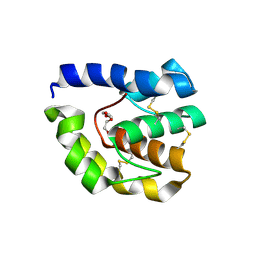 | | Drosophila OBP LUSH D118A mutation | | Descriptor: | General odorant-binding protein lush, TRIETHYLENE GLYCOL | | Authors: | Laughlin, J.D, Jones, D.N.M. | | Deposit date: | 2007-06-20 | | Release date: | 2008-06-03 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Activation of pheromone-sensitive neurons is mediated by conformational activation of pheromone-binding protein.
Cell(Cambridge,Mass.), 133, 2008
|
|
2QKT
 
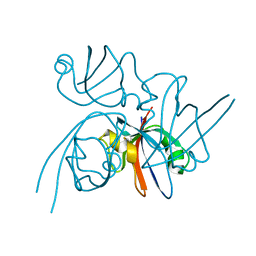 | |
5T96
 
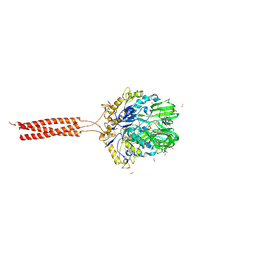 | | Crystal structure of the infectious salmon anemia virus (ISAV) HE viral receptor complex | | Descriptor: | 4-O-acetyl-5-acetamido-3,5-dideoxy-L-glycero-alpha-D-galacto-non-2-ulopyranosonic acid, FORMIC ACID, GLYCEROL, ... | | Authors: | Cook, J.D, Sultana, A, Lee, J.E. | | Deposit date: | 2016-09-09 | | Release date: | 2017-03-22 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the infectious salmon anemia virus receptor complex illustrates a unique binding strategy for attachment.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
2PGH
 | | STRUCTURE DETERMINATION OF AQUOMET PORCINE HEMOGLOBIN AT 2.8 ANGSTROM RESOLUTION | | Descriptor: | HEMOGLOBIN (AQUO MET) (ALPHA CHAIN), HEMOGLOBIN (AQUO MET) (BETA CHAIN), PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Katz, D.S, White, S.P, Huang, W, Kumar, R, Christianson, D.W. | | Deposit date: | 1994-09-16 | | Release date: | 1994-11-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure determination of aquomet porcine hemoglobin at 2.8 A resolution.
J.Mol.Biol., 244, 1994
|
|
4F7U
 
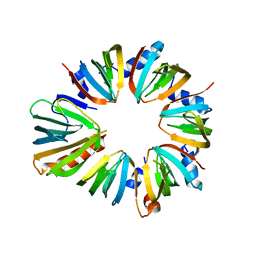 | | The 6S snRNP assembly intermediate | | Descriptor: | HEXAETHYLENE GLYCOL, Methylosome subunit pICln, Small nuclear ribonucleoprotein E, ... | | Authors: | Grimm, C, Pelz, J.P. | | Deposit date: | 2012-05-16 | | Release date: | 2013-01-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Structural Basis of Assembly Chaperone- Mediated snRNP Formation.
Mol.Cell, 49, 2013
|
|
4DBQ
 
 | |
5VJC
 
 | |
5VO5
 
 | |
5W3D
 
 | | The structure of kinesin-14 wild-type Ncd-ADP dimer | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Protein claret segregational | | Authors: | Park, H.W, Ma, Z, Chacko, J, Jiang, S.M, Robinson, R.C, Endow, S.A. | | Deposit date: | 2017-06-07 | | Release date: | 2017-12-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Structural basis of small molecule ATPase inhibition of a human mitotic kinesin motor protein.
Sci Rep, 7, 2017
|
|
5Z81
 
 | |
4EAG
 
 | | Co-crystal structure of an chimeric AMPK core with ATP | | Descriptor: | 5'-AMP-activated protein kinase subunit beta-1, 5'-AMP-activated protein kinase subunit gamma-1, ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Chen, L, Wang, J, Zhang, Y.-Y, Yan, S.F, Neumann, D, Schlattner, U, Wang, Z.-X, Wu, J.-W. | | Deposit date: | 2012-03-22 | | Release date: | 2012-06-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.701 Å) | | Cite: | AMP-activated protein kinase undergoes nucleotide-dependent conformational changes
Nat.Struct.Mol.Biol., 19, 2012
|
|
2QKU
 
 | |
