8OTB
 
 | |
3AUV
 
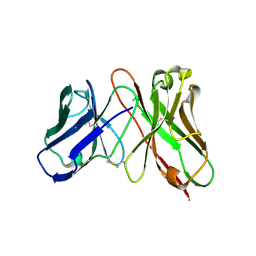 | | Predicting Amino Acid Preferences in the Complementarity Determining Regions of an Antibody-Antigen Recognition Interface | | Descriptor: | sc-dsFv derived from the G6-Fab | | Authors: | Yu, C.M, Peng, H.P, Chen, I.C, Lee, Y.C, Chen, J.B, Tsai, K.C, Chen, C.T, Chang, J.Y, Yang, E.W, Hsu, P.C, Jian, J.W, Hsu, H.J, Chang, H.J, Hsu, W.L, Huang, K.F, Ma, A.C, Yang, A.S. | | Deposit date: | 2011-02-16 | | Release date: | 2012-02-22 | | Last modified: | 2012-04-25 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Rationalization and design of the complementarity determining region sequences in an antibody-antigen recognition interface
Plos One, 7, 2012
|
|
1FL0
 
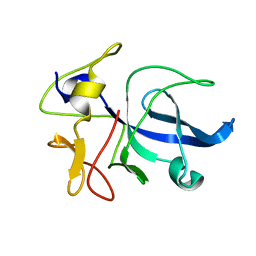 | | CRYSTAL STRUCTURE OF THE EMAP2/RNA-BINDING DOMAIN OF THE P43 PROTEIN FROM HUMAN AMINOACYL-TRNA SYNTHETASE COMPLEX | | Descriptor: | ENDOTHELIAL-MONOCYTE ACTIVATING POLYPEPTIDE II | | Authors: | Renault, L, Kerjan, P, Pasqualato, S, Menetrey, J, Robinson, J.-C, Kawaguchi, S, Vassylyev, D.G, Yokoyama, S, Mirande, M, Cherfils, J. | | Deposit date: | 2000-08-11 | | Release date: | 2000-12-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure of the EMAPII domain of human aminoacyl-tRNA synthetase complex reveals evolutionary dimer mimicry.
EMBO J., 20, 2001
|
|
2F3T
 
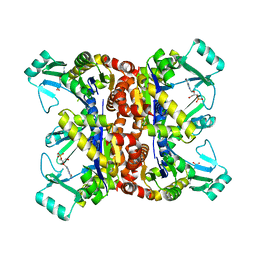 | |
2F3R
 
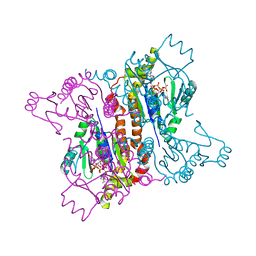 | |
3IQ6
 
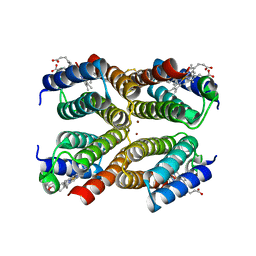 | |
3VOW
 
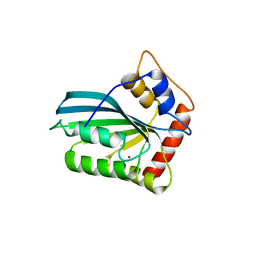 | | Crystal Structure of the Human APOBEC3C having HIV-1 Vif-binding Interface | | Descriptor: | CHLORIDE ION, Probable DNA dC->dU-editing enzyme APOBEC-3C, ZINC ION | | Authors: | Kitamura, S, Suzuki, A, Watanabe, N, Iwatani, Y. | | Deposit date: | 2012-02-22 | | Release date: | 2012-10-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The APOBEC3C crystal structure and the interface for HIV-1 Vif binding.
Nat.Struct.Mol.Biol., 19, 2012
|
|
3C3T
 
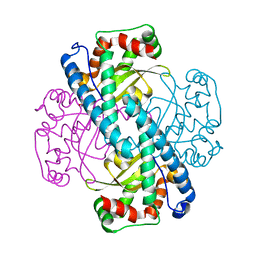 | | Role of a Glutamate Bridge Spanning the Dimeric Interface of Human Manganese Superoxide Dismutase | | Descriptor: | MANGANESE (II) ION, SULFATE ION, Superoxide dismutase [Mn] | | Authors: | Quint, P.S, Domsic, J.F, Cabelli, D.E, McKenna, R, Silverman, D.N. | | Deposit date: | 2008-01-28 | | Release date: | 2008-04-22 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Role of a glutamate bridge spanning the dimeric interface of human manganese superoxide dismutase.
Biochemistry, 47, 2008
|
|
3SXU
 
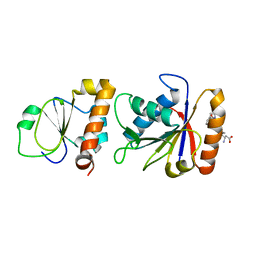 | |
4B4N
 
 | |
3M79
 
 | |
1R9F
 
 | | Crystal structure of p19 complexed with 19-bp small interfering RNA | | Descriptor: | 5'-R(*CP*GP*UP*AP*CP*GP*CP*GP*GP*AP*AP*UP*AP*CP*UP*UP*CP*GP*AP*UP*U)-3', 5'-R(*UP*CP*GP*AP*AP*GP*UP*AP*UP*UP*CP*CP*GP*CP*GP*UP*AP*CP*GP*UP*U)-3', Core protein P19, ... | | Authors: | Ye, K, Malinina, L, Patel, D.J. | | Deposit date: | 2003-10-28 | | Release date: | 2004-01-27 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Recognition of small interfering RNA by a viral suppressor of RNA
Nature, 426, 2003
|
|
4LC5
 
 | | Structural basis of substrate specificity of CDA superfamily guanine deaminase | | Descriptor: | 1,2-ETHANEDIOL, 9-METHYLGUANINE, Cytidine and deoxycytidylate deaminase zinc-binding region, ... | | Authors: | Bitra, A, Biswas, A, Anand, R. | | Deposit date: | 2013-06-21 | | Release date: | 2014-01-22 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural basis of the substrate specificity of cytidine deaminase superfamily Guanine deaminase
Biochemistry, 52, 2013
|
|
1UR5
 
 | | Stabilization of a Tetrameric Malate Dehydrogenase by Introduction of a Disulfide Bridge at the Dimer/Dimer Interface | | Descriptor: | CADMIUM ION, CHLORIDE ION, MALATE DEHYDROGENASE, ... | | Authors: | Bjork, A, Dalhus, B, Mantzilas, D, Eijsink, V.G.H, Sirevag, R. | | Deposit date: | 2003-10-27 | | Release date: | 2003-11-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Stabilization of a Tetrameric Malate Dehydrogenase by Introduction of a Disulfide Bridge at the Dimer-Dimer Interface
J.Mol.Biol., 334, 2003
|
|
2Q1E
 
 | |
2RB5
 
 | |
2RAR
 
 | |
2RAV
 
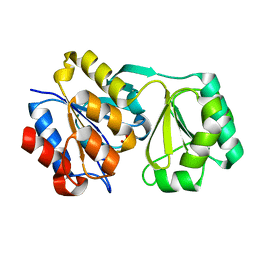 | |
2RBK
 
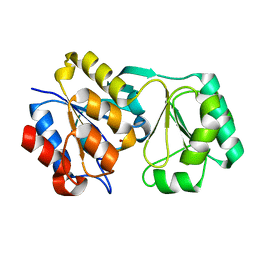 | |
1ORO
 
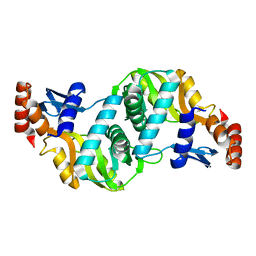 | | A FLEXIBLE LOOP AT THE DIMER INTERFACE IS A PART OF THE ACTIVE SITE OF THE ADJACENT MONOMER OF ESCHERICHIA COLI OROTATE PHOSPHORIBOSYLTRANSFERASE | | Descriptor: | OROTATE PHOSPHORIBOSYLTRANSFERASE, SULFATE ION | | Authors: | Henriksen, A, Aghajari, N, Jensen, K.F, Gajhede, M. | | Deposit date: | 1995-09-11 | | Release date: | 1996-04-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | A flexible loop at the dimer interface is a part of the active site of the adjacent monomer of Escherichia coli orotate phosphoribosyltransferase.
Biochemistry, 35, 1996
|
|
3Q3F
 
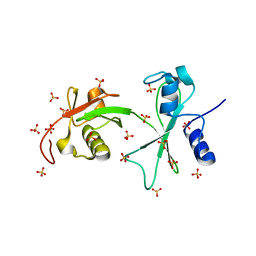 | | Engineering Domain-Swapped Binding Interfaces by Mutually Exclusive Folding: Insertion of Ubiquitin into position 103 of Barnase | | Descriptor: | Ribonuclease/Ubiquitin chimeric protein, SULFATE ION | | Authors: | Ha, J.-H, Karchin, J.M, Walker-Kopp, N, Huang, L.-S, Berry, E.A, Loh, S.N. | | Deposit date: | 2010-12-21 | | Release date: | 2012-01-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.169 Å) | | Cite: | Engineering domain-swapped binding interfaces by mutually exclusive folding.
J.Mol.Biol., 416, 2012
|
|
2BD1
 
 | | A possible role of the second calcium ion in interfacial binding: Atomic and medium resolution crystal structures of the quadruple mutant of phospholipase A2 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCIUM ION, Phospholipase A2 | | Authors: | Sekar, K, Velmurugan, D, Tsai, M.D. | | Deposit date: | 2005-10-19 | | Release date: | 2006-07-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Suggestive evidence for the involvement of the second calcium and surface loop in interfacial binding: monoclinic and trigonal crystal structures of a quadruple mutant of phospholipase A(2).
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2BCN
 
 | |
5TGL
 
 | | A MODEL FOR INTERFACIAL ACTIVATION IN LIPASES FROM THE STRUCTURE OF A FUNGAL LIPASE-INHIBITOR COMPLEX | | Descriptor: | LIPASE, N-HEXYLPHOSPHONATE ETHYL ESTER | | Authors: | Brzozowski, A.M, Derewenda, U, Derewenda, Z.S, Dodson, G.G, Lawson, D, Turkenburg, J.P, Bjorkling, F, Huge-Jensen, B, Patkar, S.R, Thim, L. | | Deposit date: | 1991-10-30 | | Release date: | 1994-01-31 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | A model for interfacial activation in lipases from the structure of a fungal lipase-inhibitor complex.
Nature, 351, 1991
|
|
2BCH
 
 | | A possible of Second calcium ion in interfacial binding: Atomic and Medium resolution crystal structures of the quadruple mutant of phospholipase A2 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Sekar, K, Yogavel, M, Velmurugan, D, Poi, M.J, Dauter, Z, Tsai, M.D. | | Deposit date: | 2005-10-19 | | Release date: | 2006-07-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Suggestive evidence for the involvement of the second calcium and surface loop in interfacial binding: monoclinic and trigonal crystal structures of a quadruple mutant of phospholipase A(2).
Acta Crystallogr.,Sect.D, 62, 2006
|
|
