6AI5
 
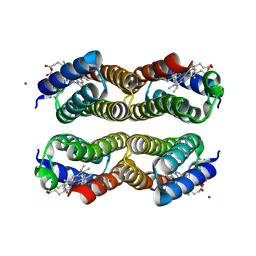 | | Disulfide-free, Zn-directed tetramer of the engineered cyt cb562 variant, C96T/A104AB3 | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Song, W.J, Tezcan, F.A. | | Deposit date: | 2018-08-21 | | Release date: | 2019-09-11 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Disulfide-free, Zn-directed tetramer of the engineered cyt cb562 variant, C96T/A104AB3
To Be Published
|
|
4MIF
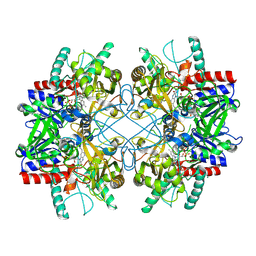 
 | | Pyranose 2-oxidase from Phanerochaete chrysosporium, wild type from natural source | | Descriptor: | DIHYDROFLAVINE-ADENINE DINUCLEOTIDE, MAGNESIUM ION, Pyranose 2-oxidase | | Authors: | Hassan, N, Tan, T.C, Spadiut, O, Pisanelli, I, Fusco, L, Haltrich, D, Peterbauer, C, Divne, C. | | Deposit date: | 2013-08-31 | | Release date: | 2013-12-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structures of Phanerochaete chrysosporium pyranose 2-oxidase suggest that the N-terminus acts as a propeptide that assists in homotetramer assembly.
FEBS Open Bio, 3, 2013
|
|
4GYP
 
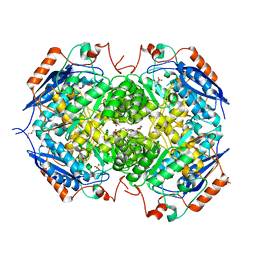 | | Crystal structure of the heterotetrameric complex of GlucD and GlucDRP from E. coli K-12 MG1655 (EFI TARGET EFI-506058) | | Descriptor: | CITRIC ACID, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Lukk, T, Ghasempur, S, Imker, H.J, Gerlt, J.A, Nair, S.K, Enzyme Function Initiative (EFI) | | Deposit date: | 2012-09-05 | | Release date: | 2012-09-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Glucarate dehydratase and its related protein from Escherichia coli form a heterotetrameric complex.
to be published
|
|
3OB8
 
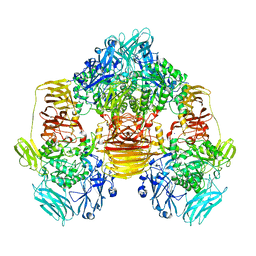 | | Structure of the beta-galactosidase from Kluyveromyces lactis in complex with galactose | | Descriptor: | Beta-galactosidase, MAGNESIUM ION, MANGANESE (II) ION, ... | | Authors: | Fernandez-Leiro, R, Pereira-Rodriguez, A, Becerra, M, Gonzalez-Siso, I, Cerdan, M.E, Sanz-Aparicio, J. | | Deposit date: | 2010-08-06 | | Release date: | 2011-08-17 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural basis of specificity in tetrameric Kluyveromyces lactis beta-galactosidase.
J.Struct.Biol., 177, 2012
|
|
7UJN
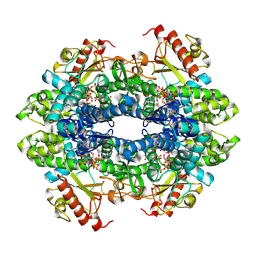 
 | | Structure of Human SAMHD1 with Non-Hydrolysable dGTP Analog | | Descriptor: | 2'-deoxyguanosine-5'-O-(1-thiotriphosphate), Deoxynucleoside triphosphate triphosphohydrolase SAMHD1 | | Authors: | Huynh, K.W, Ammirati, M, Han, S. | | Deposit date: | 2022-03-31 | | Release date: | 2022-07-20 | | Last modified: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (2.89 Å) | | Cite: | Phosphorylation of SAMHD1 Thr592 increases C-terminal domain dynamics, tetramer dissociation and ssDNA binding kinetics.
Nucleic Acids Res., 50, 2022
|
|
3T4P
 
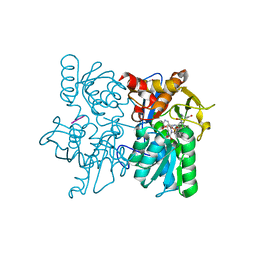 | |
3OBA
 
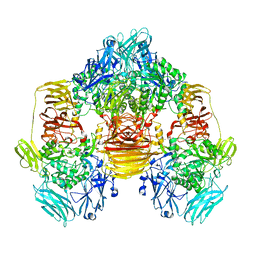 | | Structure of the beta-galactosidase from Kluyveromyces lactis | | Descriptor: | Beta-galactosidase, GLYCEROL, MANGANESE (III) ION | | Authors: | Fernandez-Leiro, R, Pereira-Rodriguez, A, Becerra, M, Gonzalez-Siso, I, Cerdan, M.E, Sanz-Aparicio, J. | | Deposit date: | 2010-08-06 | | Release date: | 2011-08-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural basis of specificity in tetrameric Kluyveromyces lactis beta-galactosidase.
J.Struct.Biol., 177, 2012
|
|
1XOF
 
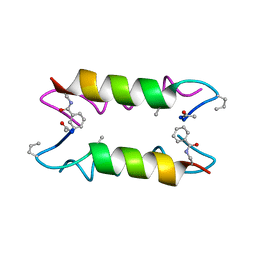 | | Heterooligomeric Beta Beta Alpha Miniprotein | | Descriptor: | BBAhetT1 | | Authors: | Ali, M.H, Taylor, C.M, Grigoryan, G, Allen, K.N, Imperiali, B, Keating, A.E. | | Deposit date: | 2004-10-06 | | Release date: | 2005-02-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Design of a Heterospecific, Tetrameric, 21-Residue Miniprotein with Mixed alpha/beta Structure.
Structure, 13, 2005
|
|
1TJC
 
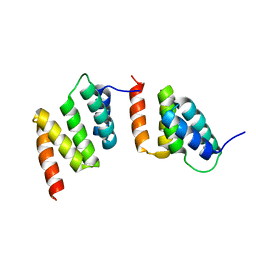 | | Crystal structure of peptide-substrate-binding domain of human type I collagen prolyl 4-hydroxylase | | Descriptor: | Prolyl 4-hydroxylase alpha-1 subunit | | Authors: | Pekkala, M, Hieta, R, Bergmann, U, Kivirikko, K.I, Wierenga, R.K, Myllyharju, J. | | Deposit date: | 2004-06-04 | | Release date: | 2004-10-12 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Peptide-Substrate-binding Domain of Collagen Prolyl 4-Hydroxylases Is a Tetratricopeptide Repeat Domain with Functional Aromatic Residues.
J.Biol.Chem., 279, 2004
|
|
3V15
 
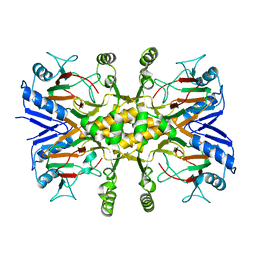 | |
3LW1
 
 | | Binary complex of 14-3-3 sigma and p53 pT387-peptide | | Descriptor: | 14-3-3 protein sigma, CHLORIDE ION, GLYCEROL, ... | | Authors: | Schumacher, B, Mondry, J, Thiel, P, Weyand, M, Ottmann, C. | | Deposit date: | 2010-02-23 | | Release date: | 2010-03-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Structure of the p53 C-terminus bound to 14-3-3: Implications for stabilization of the p53 tetramer
Febs Lett., 584, 2010
|
|
2WEL
 
 | | Crystal structure of SU6656-bound calcium/calmodulin-dependent protein kinase II delta in complex with calmodulin | | Descriptor: | (3Z)-N,N-DIMETHYL-2-OXO-3-(4,5,6,7-TETRAHYDRO-1H-INDOL-2-YLMETHYLIDENE)-2,3-DIHYDRO-1H-INDOLE-5-SULFONAMIDE, 1,2-ETHANEDIOL, CALCIUM ION, ... | | Authors: | Pike, A.C.W, Rellos, P, Salah, E, Burgess-Brown, N, Keates, T, Muniz, J, Sethi, R, Roos, A, Filippakopoulos, P, von Delft, F, Edwards, A, Weigelt, J, Arrowsmith, C.H, Bountra, C, Knapp, S. | | Deposit date: | 2009-03-31 | | Release date: | 2009-04-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the Camkiidelta/Calmodulin Complex Reveals the Molecular Mechanism of Camkii Kinase Activation.
Plos Biol., 8, 2010
|
|
3J6J
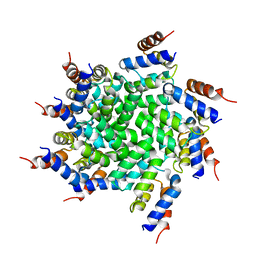 
 | | 3.6 Angstrom resolution MAVS filament generated from helical reconstruction | | Descriptor: | Mitochondrial antiviral-signaling protein | | Authors: | Wu, B, Peisley, A, Li, Z, Egelman, E, Walz, T, Penczek, P, Hur, S. | | Deposit date: | 2014-03-13 | | Release date: | 2014-07-30 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3.64 Å) | | Cite: | Molecular Imprinting as a Signal-Activation Mechanism of the Viral RNA Sensor RIG-I.
Mol.Cell, 55, 2014
|
|
6HXI
 
 | |
4ZUL
 
 | | Structure ALDH7A1 complexed with alpha-aminoadipate | | Descriptor: | 2-AMINOHEXANEDIOIC ACID, Alpha-aminoadipic semialdehyde dehydrogenase, TETRAETHYLENE GLYCOL, ... | | Authors: | Luo, M, Tanner, J.J. | | Deposit date: | 2015-05-16 | | Release date: | 2015-08-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structural Basis of Substrate Recognition by Aldehyde Dehydrogenase 7A1.
Biochemistry, 54, 2015
|
|
6HXJ
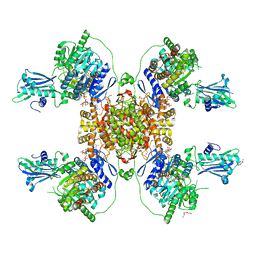 
 | |
3Q5P
 
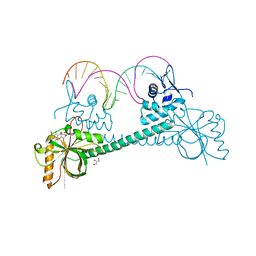 | | Crystal structure of BmrR bound to Tetracycline | | Descriptor: | 23 bp BmrR promoter DNA, GLYCEROL, Multidrug-efflux transporter 1 regulator, ... | | Authors: | Bachas, S, Eginton, C, Gunio, G, Wade, H. | | Deposit date: | 2010-12-29 | | Release date: | 2011-06-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.942 Å) | | Cite: | Structural contributions to multidrug recognition in the multidrug resistance (MDR) gene regulator, BmrR.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
4ARF
 
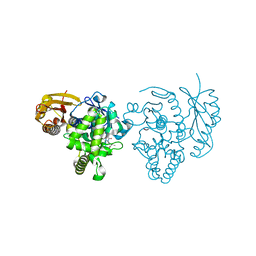 | | CRYSTAL STRUCTURE OF THE PEPTIDASE DOMAIN OF COLLAGENASE H FROM CLOSTRIDIUM HISTOLYTICUM IN COMPLEX WITH THE PEPTIDIC INHIBITOR ISOAMYLPHOSPHONYL-GLY-PRO-ALA AT 1.77 ANGSTROM RESOLUTION. | | Descriptor: | CALCIUM ION, COLH PROTEIN, ISOAMYLPHOSPHONYL-GLY-PRO-ALA, ... | | Authors: | Eckhard, U, Brandstetter, H. | | Deposit date: | 2012-04-23 | | Release date: | 2013-06-05 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structural Basis for Activity Regulation and Substrate Preference of Clostridial Collagenases G, H, and T.
J.Biol.Chem., 288, 2013
|
|
3V17
 
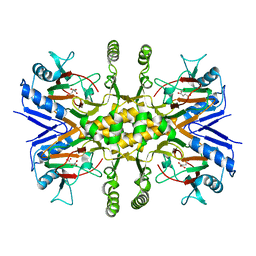 | |
4ZUK
 
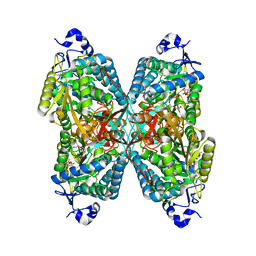 | | Structure ALDH7A1 complexed with NAD+ | | Descriptor: | Alpha-aminoadipic semialdehyde dehydrogenase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, TETRAETHYLENE GLYCOL | | Authors: | Luo, M, Tanner, J.J. | | Deposit date: | 2015-05-16 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Structural Basis of Substrate Recognition by Aldehyde Dehydrogenase 7A1.
Biochemistry, 54, 2015
|
|
3OL3
 
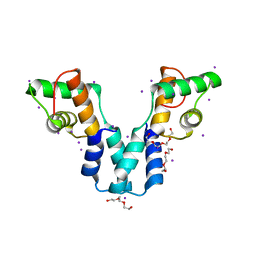 | |
4AWB
 
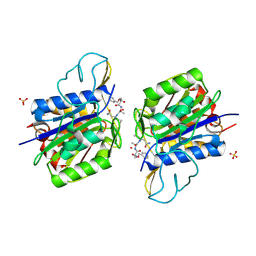 | | Crystal structure of active legumain in complex with AAN-CMK | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, LEGUMAIN, MERCURY (II) ION, ... | | Authors: | Dall, E, Brandstetter, H. | | Deposit date: | 2012-06-01 | | Release date: | 2013-06-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Mechanistic and Structural Studies on Legumain Explain its Zymogenicity, Distinct Activation Pathways, and Regulation
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4MPU
 
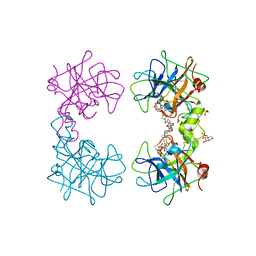 | | Human beta-tryptase co-crystal structure with (6S,8R)-N,N'-bis[3-({4-[3-(aminomethyl)phenyl]piperidin-1-yl}carbonyl)phenyl]-8-hydroxy-6-(1-hydroxycyclobutyl)-5,7-dioxaspiro[3.4]octane-6,8-dicarboxamide | | Descriptor: | (6S,8R)-N,N'-bis[3-({4-[3-(aminomethyl)phenyl]piperidin-1-yl}carbonyl)phenyl]-8-hydroxy-6-(1-hydroxycyclobutyl)-5,7-dioxaspiro[3.4]octane-6,8-dicarboxamide, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, SULFATE ION, ... | | Authors: | White, A, Stein, A.J, Suto, R.K. | | Deposit date: | 2013-09-13 | | Release date: | 2015-03-18 | | Last modified: | 2019-09-25 | | Method: | X-RAY DIFFRACTION (1.649 Å) | | Cite: | Target-Directed Self-Assembly of Homodimeric Drugs Against beta-Tryptase.
Acs Med.Chem.Lett., 9, 2018
|
|
4L0A
 
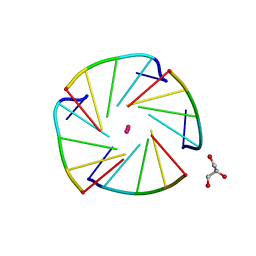 | | X-ray structure of an all LNA quadruplex | | Descriptor: | DNA/RNA (5'-R(*(TLN)P*(LCG)P*(LCG)P*(LCG)P*(TLN))-3'), GLYCEROL, POTASSIUM ION | | Authors: | Russo Krauss, I, Parkinson, G, Merlino, A, Mazzarella, L, Sica, F. | | Deposit date: | 2013-05-31 | | Release date: | 2014-03-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A regular thymine tetrad and a peculiar supramolecular assembly in the first crystal structure of an all-LNA G-quadruplex.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
5AMO
 
 | | Structure of a mouse Olfactomedin-1 disulfide-linked dimer of the Olfactomedin domain and part of the coiled coil | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Pronker, M.F, Bos, T.G.A.A, Sharp, T.H, Thies-Weesie, D.M, Janssen, B.J.C. | | Deposit date: | 2015-03-11 | | Release date: | 2015-04-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Olfactomedin-1 Has a V-Shaped Disulfide-Linked Tetrameric Structure.
J.Biol.Chem., 290, 2015
|
|
