7XV3
 
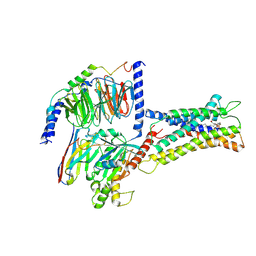 | | Cryo-EM structure of LPS-bound GPR174 in complex with Gs protein | | 分子名称: | (2~{S})-2-$l^{4}-azanyl-3-[[(2~{R})-3-octadecanoyloxy-2-oxidanyl-propoxy]-oxidanyl-oxidanylidene-$l^{6}-phosphanyl]oxy-propanoic acid, Engineered G protein subunit S (mini-Gs), Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | 著者 | He, Y, Liang, J. | | 登録日 | 2022-05-20 | | 公開日 | 2023-02-15 | | 最終更新日 | 2023-03-08 | | 実験手法 | ELECTRON MICROSCOPY (2.76 Å) | | 主引用文献 | Structural basis of lysophosphatidylserine receptor GPR174 ligand recognition and activation.
Nat Commun, 14, 2023
|
|
7ZX2
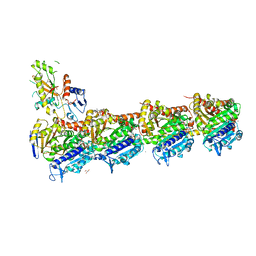 
 | | Tubulin-Pelophen B complex | | 分子名称: | (3R,4S,7S,9S,11S)-3,4,11-trihydroxy-7-((R,Z)-4-(hydroxymethyl)hex-2-en-2-yl)-9-methoxy-12,12-dimethyl-6-oxa-1(1,3)-benzenacyclododecaphan-5-one, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, ... | | 著者 | Estevez-Gallego, J, Diaz, J.F, Van der Eycken, J, Oliva, M.A. | | 登録日 | 2022-05-20 | | 公開日 | 2022-11-23 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Chemical modulation of microtubule structure through the laulimalide/peloruside site.
Structure, 31, 2023
|
|
7ZXD
 
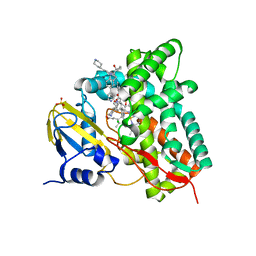 | | Crystal structure of CYP125 from Mycobacterium tuberculosis in complex with an inhibitor | | 分子名称: | 1-[1-(2-piperidin-4-ylethyl)-5-pyridin-4-yl-indol-2-yl]butan-1-one, CHLORIDE ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | 著者 | Snee, M, Katariya, M, Levy, C, Leys, D. | | 登録日 | 2022-05-20 | | 公開日 | 2023-04-05 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (2.09 Å) | | 主引用文献 | Structure Based Discovery of Inhibitors of CYP125 and CYP142 from Mycobacterium tuberculosis.
Chemistry, 29, 2023
|
|
8CWH
 
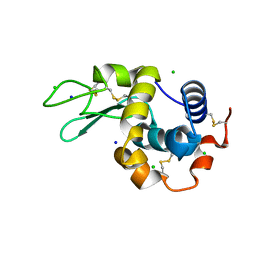 | | 200us Temperature-Jump (Dark2) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWG
 
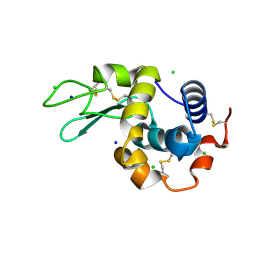 | | 200us Temperature-Jump (Dark1) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWB
 
 | | Laser Off Temperature-Jump XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.51 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWE
 
 | | 20ns Temperature-Jump (Dark2) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWD
 
 | | 20ns Temperature-Jump (Dark1) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWF
 
 | | 200us Temperature-Jump (Light) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CWC
 
 | | 20ns Temperature-Jump (Light) XFEL structure of Lysozyme Bound to N,N'-diacetylchitobiose | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-19 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.48 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CX5
 
 | | Crystal Structure of small molecule alpha,beta-ketoamide 4 covalently bound to K-Ras(G12R) | | 分子名称: | GUANOSINE-5'-DIPHOSPHATE, Isoform 2B of GTPase KRas, MAGNESIUM ION, ... | | 著者 | Zhang, Z, Morstein, J, Ecker, A, Guiley, K.Z, Shokat, K.M. | | 登録日 | 2022-05-19 | | 公開日 | 2022-08-31 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.72 Å) | | 主引用文献 | Chemoselective Covalent Modification of K-Ras(G12R) with a Small Molecule Electrophile.
J.Am.Chem.Soc., 144, 2022
|
|
7ZWH
 
 | | VWF Tubules of D1D2 and D'D3A1 domains | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Javitt, G, Fass, D. | | 登録日 | 2022-05-19 | | 公開日 | 2022-07-13 | | 最終更新日 | 2023-01-11 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | Assembly of von Willebrand factor tubules with in vivo helical parameters requires A1 domain insertion.
Blood, 140, 2022
|
|
7ZWG
 
 | | The Crystal structure of RO4493940 bound to CK2alpha | | 分子名称: | (5~{Z})-5-(quinolin-6-ylmethylidene)-1,3-thiazolidine-2,4-dione, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ACETATE ION, ... | | 著者 | Brear, P, Hyvonen, M. | | 登録日 | 2022-05-19 | | 公開日 | 2023-05-31 | | 最終更新日 | 2024-05-15 | | 実験手法 | X-RAY DIFFRACTION (1.31 Å) | | 主引用文献 | Chemical proteomics reveals the target landscape of 1,000 kinase inhibitors.
Nat.Chem.Biol., 20, 2024
|
|
7ZWE
 
 | | The Crystal structure of GW8695 bound to CK2alpha | | 分子名称: | 7-(1~{H}-indol-2-yl)-5-methyl-~{N}-(3,4,5-trimethoxyphenyl)imidazo[5,1-f][1,2,4]triazin-2-amine, Casein kinase II subunit alpha | | 著者 | Brear, P, Hyvonen, M. | | 登録日 | 2022-05-19 | | 公開日 | 2023-05-31 | | 最終更新日 | 2024-05-15 | | 実験手法 | X-RAY DIFFRACTION (1.47 Å) | | 主引用文献 | Chemical proteomics reveals the target landscape of 1,000 kinase inhibitors.
Nat.Chem.Biol., 20, 2024
|
|
7XUF
 
 | | Cryo-EM structure of the AKT1-AtKC1 complex from Arabidopsis thaliana | | 分子名称: | POTASSIUM ION, Potassium channel AKT1, Potassium channel KAT3 | | 著者 | Yang, G.H, Lu, Y.M, Jia, Y.T, Yang, F, Zhang, Y.M, Xu, X, Li, X.M, Lei, J.L. | | 登録日 | 2022-05-18 | | 公開日 | 2022-11-09 | | 最終更新日 | 2024-07-03 | | 実験手法 | ELECTRON MICROSCOPY (3.3 Å) | | 主引用文献 | Structural basis for the activity regulation of a potassium channel AKT1 from Arabidopsis.
Nat Commun, 13, 2022
|
|
8CW0
 
 | | 20us Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW7
 
 | | 200us Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVU
 
 | | 20ns Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW6
 
 | | 200us Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW8
 
 | | Laser Off Temperature-Jump XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW3
 
 | | 20us Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW1
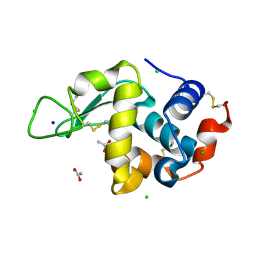 
 | | 20us Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVV
 
 | | 20ns Temperature-Jump (Dark1) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CVW
 
 | | 20ns Temperature-Jump (Dark2) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
8CW5
 
 | | 200us Temperature-Jump (Light) XFEL structure of Lysozyme | | 分子名称: | ACETATE ION, CHLORIDE ION, Lysozyme C, ... | | 著者 | Wolff, A.M, Thompson, M.C, Fraser, J.S, Nango, E. | | 登録日 | 2022-05-18 | | 公開日 | 2022-06-22 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.57 Å) | | 主引用文献 | Mapping protein dynamics at high spatial resolution with temperature-jump X-ray crystallography.
Nat.Chem., 15, 2023
|
|
