6LAX
 | | the mutant SAM-VI riboswitch (U6C) bound to SAM | | 分子名称: | RNA (55-MER), S-ADENOSYLMETHIONINE, U1 small nuclear ribonucleoprotein A | | 著者 | Sun, A, Ren, A. | | 登録日 | 2019-11-13 | | 公開日 | 2020-01-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.7 Å) | | 主引用文献 | SAM-VI riboswitch structure and signature for ligand discrimination.
Nat Commun, 10, 2019
|
|
6LAU
 | | the wildtype SAM-VI riboswitch bound to SAH | | 分子名称: | CESIUM ION, GUANOSINE-5'-TRIPHOSPHATE, RNA (54-MER), ... | | 著者 | Ren, A, Sun, A. | | 登録日 | 2019-11-13 | | 公開日 | 2020-01-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (3.109 Å) | | 主引用文献 | SAM-VI riboswitch structure and signature for ligand discrimination.
Nat Commun, 10, 2019
|
|
6LAS
 
 | | the wildtype SAM-VI riboswitch bound to SAM | | 分子名称: | RNA (55-MER), S-ADENOSYLMETHIONINE, U1 small nuclear ribonucleoprotein A | | 著者 | Ren, A, Sun, A. | | 登録日 | 2019-11-13 | | 公開日 | 2020-01-01 | | 実験手法 | X-RAY DIFFRACTION (2.708 Å) | | 主引用文献 | SAM-VI riboswitch structure and signature for ligand discrimination.
Nat Commun, 10, 2019
|
|
6KOR
 
 | | Crystal structure of the RRM domain of SYNCRIP | | 分子名称: | Heterogeneous nuclear ribonucleoprotein Q | | 著者 | Chen, Y, Chan, J, Chen, W, Jobichen, C. | | 登録日 | 2019-08-12 | | 公開日 | 2020-01-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.602 Å) | | 主引用文献 | SYNCRIP, a new player in pri-let-7a processing.
Rna, 26, 2020
|
|
6JVY
 
 | | Crystal structure of RBM38 in complex with single-stranded DNA | | 分子名称: | DNA (5'-D(*TP*GP*TP*GP*TP*GP*TP*GP*TP*GP*TP*G)-3'), RNA-binding protein 38, SULFATE ION | | 著者 | Qian, K, Li, M, Wang, J, Zhang, M, Wang, M. | | 登録日 | 2019-04-17 | | 公開日 | 2020-01-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.003 Å) | | 主引用文献 | Structural basis for mRNA recognition by human RBM38.
Biochem.J., 477, 2020
|
|
6JVX
 
 | | Crystal structure of RBM38 in complex with RNA | | 分子名称: | RNA (5'-R(*UP*GP*UP*GP*UP*GP*UP*GP*UP*GP*UP*G)-3'), RNA-binding protein 38, SULFATE ION | | 著者 | Qian, K, Li, M, Wang, J, Zhang, M, Wang, M. | | 登録日 | 2019-04-17 | | 公開日 | 2020-01-01 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.301 Å) | | 主引用文献 | Structural basis for mRNA recognition by human RBM38.
Biochem.J., 477, 2020
|
|
6J6Q
 
 | | Cryo-EM structure of the yeast B*-b2 complex at an average resolution of 3.7 angstrom | | 分子名称: | GUANOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, MAGNESIUM ION, ... | | 著者 | Wan, R, Bai, R, Yan, C, Lei, J, Shi, Y. | | 登録日 | 2019-01-15 | | 公開日 | 2019-04-24 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Structures of the Catalytically Activated Yeast Spliceosome Reveal the Mechanism of Branching.
Cell, 177, 2019
|
|
6J6N
 
 | | Cryo-EM structure of the yeast B*-b1 complex at an average resolution of 3.86 angstrom | | 分子名称: | GUANOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, MAGNESIUM ION, ... | | 著者 | Wan, R, Bai, R, Yan, C, Lei, J, Shi, Y. | | 登録日 | 2019-01-15 | | 公開日 | 2019-04-24 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (3.86 Å) | | 主引用文献 | Structures of the Catalytically Activated Yeast Spliceosome Reveal the Mechanism of Branching.
Cell, 177, 2019
|
|
6J6H
 
 | | Cryo-EM structure of the yeast B*-a1 complex at an average resolution of 3.6 angstrom | | 分子名称: | ACT1 pre-mRNA, GUANOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, ... | | 著者 | Wan, R, Bai, R, Yan, C, Lei, J, Shi, Y. | | 登録日 | 2019-01-15 | | 公開日 | 2019-04-24 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (3.6 Å) | | 主引用文献 | Structures of the Catalytically Activated Yeast Spliceosome Reveal the Mechanism of Branching.
Cell, 177, 2019
|
|
6J6G
 
 | | Cryo-EM structure of the yeast B*-a2 complex at an average resolution of 3.2 angstrom | | 分子名称: | ACT1 pre-mRNA, GUANOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, ... | | 著者 | Wan, R, Bai, R, Yan, C, Lei, J, Shi, Y. | | 登録日 | 2019-01-15 | | 公開日 | 2019-04-24 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | Structures of the Catalytically Activated Yeast Spliceosome Reveal the Mechanism of Branching.
Cell, 177, 2019
|
|
6ID1
 
 | | Cryo-EM structure of a human intron lariat spliceosome after Prp43 loaded (ILS2 complex) at 2.9 angstrom resolution | | 分子名称: | 116 kDa U5 small nuclear ribonucleoprotein component, CWF19-like protein 2, Cell division cycle 5-like protein, ... | | 著者 | Zhang, X, Zhan, X, Yan, C, Shi, Y. | | 登録日 | 2018-09-07 | | 公開日 | 2019-03-13 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (2.86 Å) | | 主引用文献 | Structures of the human spliceosomes before and after release of the ligated exon.
Cell Res., 29, 2019
|
|
6ID0
 
 | | Cryo-EM structure of a human intron lariat spliceosome prior to Prp43 loaded (ILS1 complex) at 2.9 angstrom resolution | | 分子名称: | 116 kDa U5 small nuclear ribonucleoprotein component, CWF19-like protein 2, Cell division cycle 5-like protein, ... | | 著者 | Zhang, X, Zhan, X, Yan, C, Shi, Y. | | 登録日 | 2018-09-07 | | 公開日 | 2019-03-13 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (2.9 Å) | | 主引用文献 | Structures of the human spliceosomes before and after release of the ligated exon.
Cell Res., 29, 2019
|
|
6ICZ
 
 | | Cryo-EM structure of a human post-catalytic spliceosome (P complex) at 3.0 angstrom | | 分子名称: | 116 kDa U5 small nuclear ribonucleoprotein component, ADENOSINE-5'-TRIPHOSPHATE, ATP-dependent RNA helicase DHX8, ... | | 著者 | Zhang, X, Zhan, X, Yan, C, Shi, Y. | | 登録日 | 2018-09-07 | | 公開日 | 2019-03-13 | | 最終更新日 | 2020-10-14 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Structures of the human spliceosomes before and after release of the ligated exon.
Cell Res., 29, 2019
|
|
6HPJ
 
 | | Structure of human SRSF1 RRM1 bound to AACAAA RNA | | 分子名称: | Immunoglobulin G-binding protein G,Serine/arginine-rich splicing factor 1, RNA (5'-R(*AP*AP*CP*AP*AP*A)-3') | | 著者 | Allain, F.T.H, Clery, A. | | 登録日 | 2018-09-21 | | 公開日 | 2020-11-18 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structure of SRSF1 RRM1 bound to RNA reveals an unexpected bimodal mode of interaction and explains its involvement in SMN1 exon7 splicing.
Nat Commun, 12, 2021
|
|
6HIP
 
 | | Structure of SPF45 UHM bound to HIV-1 Rev ULM | | 分子名称: | HIV-1 Rev (41-49), SODIUM ION, Splicing factor 45, ... | | 著者 | Pabis, M, Corsini, L, Sattler, M. | | 登録日 | 2018-08-30 | | 公開日 | 2019-03-27 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.2 Å) | | 主引用文献 | Modulation of HIV-1 gene expression by binding of a ULM motif in the Rev protein to UHM-containing splicing factors.
Nucleic Acids Res., 47, 2019
|
|
6GX6
 
 | | Crystal structure of IMP3 RRM12 in complex with RNA (ACAC) | | 分子名称: | 1,2-ETHANEDIOL, Insulin-like growth factor 2 mRNA-binding protein 3, PHOSPHATE ION, ... | | 著者 | Jia, M, Gut, H, Chao, A.J. | | 登録日 | 2018-06-26 | | 公開日 | 2018-09-05 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural basis of IMP3 RRM12 recognition of RNA.
RNA, 24, 2018
|
|
6GSN
 
 | |
6GSM
 
 | | Structure of a partial yeast 48S preinitiation complex in open conformation. | | 分子名称: | 18S ribosomal RNA, 40S ribosomal protein S0, 40S ribosomal protein S1, ... | | 著者 | Llacer, J.L, Hussain, T, Gordiyenko, Y, Ramakrishnan, V. | | 登録日 | 2018-06-14 | | 公開日 | 2019-07-31 | | 最終更新日 | 2024-07-10 | | 実験手法 | ELECTRON MICROSCOPY (5.15 Å) | | 主引用文献 | Large-scale movement of eIF3 domains during translation initiation modulate start codon selection.
Nucleic Acids Res., 2021
|
|
6GML
 
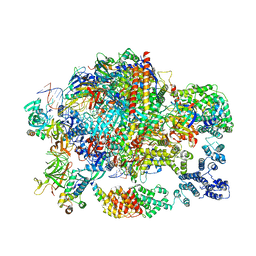 | | Structure of paused transcription complex Pol II-DSIF-NELF | | 分子名称: | DNA-directed RNA polymerase II subunit RPB9, DNA-directed RNA polymerase subunit, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Vos, S.M, Farnung, L, Urlaub, H, Cramer, P. | | 登録日 | 2018-05-27 | | 公開日 | 2018-09-05 | | 最終更新日 | 2024-05-15 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | Structure of paused transcription complex Pol II-DSIF-NELF.
Nature, 560, 2018
|
|
6GD3
 
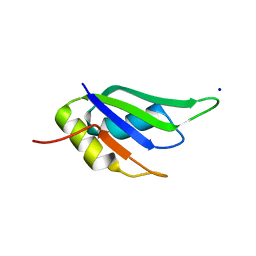 | |
6GD2
 
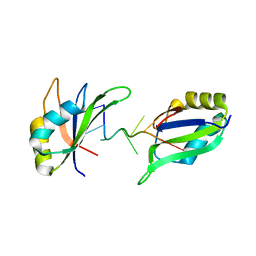 | | Structure of HuR RRM3 in complex with RNA | | 分子名称: | ELAV-like protein 1, RNA (5'-R(P*UP*UP*UP*AP*UP*UP*U)-3') | | 著者 | Pabis, M, Sattler, M. | | 登録日 | 2018-04-21 | | 公開日 | 2018-10-31 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | HuR biological function involves RRM3-mediated dimerization and RNA binding by all three RRMs.
Nucleic Acids Res., 47, 2019
|
|
6GD1
 
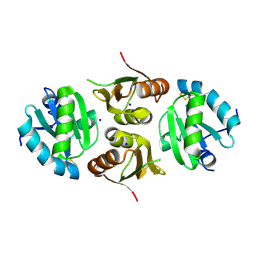 | | Structure of HuR RRM3 | | 分子名称: | SODIUM ION, Thioredoxin 1,ELAV-like protein 1 | | 著者 | Pabis, M, Sattler, M. | | 登録日 | 2018-04-21 | | 公開日 | 2018-10-31 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.01 Å) | | 主引用文献 | HuR biological function involves RRM3-mediated dimerization and RNA binding by all three RRMs.
Nucleic Acids Res., 47, 2019
|
|
6GC5
 
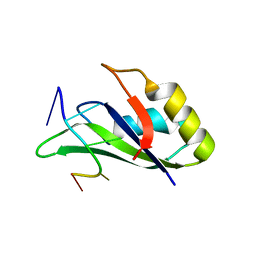 | |
6GBM
 
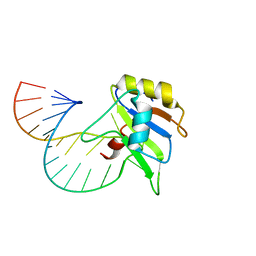 | | Solution structure of FUS-RRM bound to stem-loop RNA | | 分子名称: | RNA (5'-R(*GP*GP*CP*AP*GP*AP*UP*UP*AP*CP*AP*AP*UP*UP*CP*UP*AP*UP*UP*UP*GP*CP*C)-3'), RNA-binding protein FUS | | 著者 | Loughlin, F.E, Allain, F.H.-T. | | 登録日 | 2018-04-15 | | 公開日 | 2019-02-20 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | The Solution Structure of FUS Bound to RNA Reveals a Bipartite Mode of RNA Recognition with Both Sequence and Shape Specificity.
Mol. Cell, 73, 2019
|
|
6G90
 
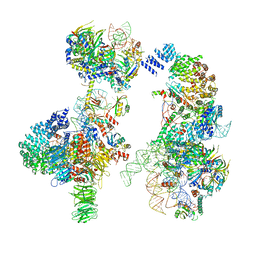 | | Prespliceosome structure provides insight into spliceosome assembly and regulation (map A2) | | 分子名称: | 56 kDa U1 small nuclear ribonucleoprotein component, Cold sensitive U2 snRNA suppressor 1, Pre-mRNA-processing factor 39, ... | | 著者 | Plaschka, C, Lin, P.-C, Charenton, C, Nagai, K. | | 登録日 | 2018-04-10 | | 公開日 | 2018-08-22 | | 実験手法 | ELECTRON MICROSCOPY (4 Å) | | 主引用文献 | Prespliceosome structure provides insights into spliceosome assembly and regulation.
Nature, 559, 2018
|
|
