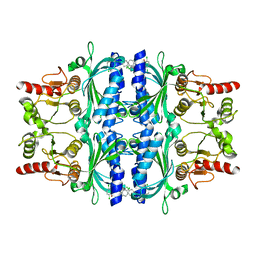
2FHY
 | |
2FMT
 
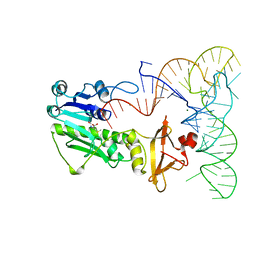 | | METHIONYL-TRNAFMET FORMYLTRANSFERASE COMPLEXED WITH FORMYL-METHIONYL-TRNAFMET | | Descriptor: | FORMYL-METHIONYL-TRNAFMET2, MAGNESIUM ION, METHIONYL-TRNA FMET FORMYLTRANSFERASE, ... | | Authors: | Schmitt, E, Mechulam, Y, Blanquet, S. | | Deposit date: | 1998-07-29 | | Release date: | 1999-07-29 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of methionyl-tRNAfMet transformylase complexed with the initiator formyl-methionyl-tRNAfMet.
EMBO J., 17, 1998
|
|
2FQN
 
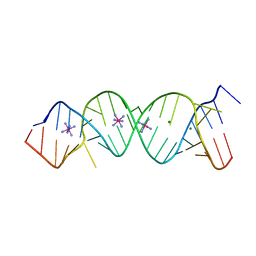 | |
2FV2
 
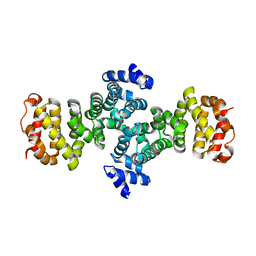 | |
4RC4
 
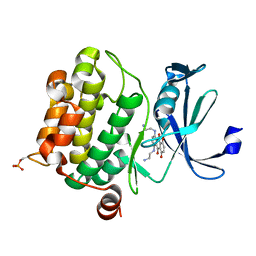 | |
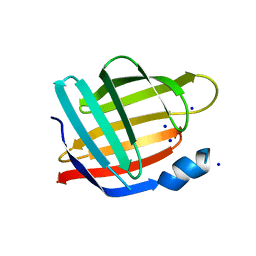
2FRS
 | |
2FSJ
 
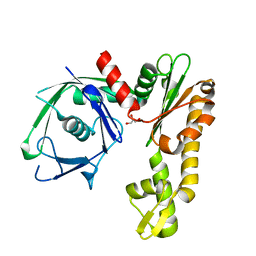 | | Crystal structure of Ta0583, an archaeal actin homolog, native data | | Descriptor: | GLYCEROL, hypothetical protein Ta0583 | | Authors: | Roeben, A, Kofler, C, Nagy, I, Nickell, S, Ulrich Hartl, F, Bracher, A. | | Deposit date: | 2006-01-23 | | Release date: | 2006-04-18 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of an archaeal actin homolog
J.Mol.Biol., 358, 2006
|
|
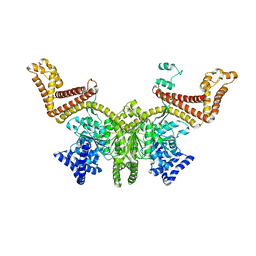
2FSF
 | |
2FSI
 
 | |
6P7W
 
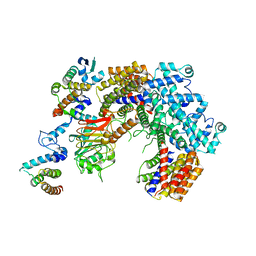 | | Structure of the K. lactis CBF3 core - Ndc10 D1 complex | | Descriptor: | Cep3, Ctf13, Ndc10, ... | | Authors: | Lee, P.D, Wei, H, Tan, D, Harrison, S.C. | | Deposit date: | 2019-06-06 | | Release date: | 2019-09-18 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Structure of the Centromere Binding Factor 3 Complex from Kluyveromyces lactis.
J.Mol.Biol., 431, 2019
|
|
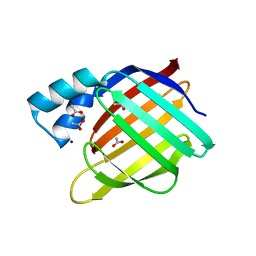
2FS6
 | |
2FSK
 
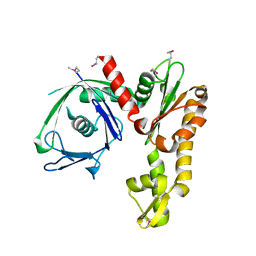 | | Crystal structure of Ta0583, an archaeal actin homolog, SeMet data | | Descriptor: | hypothetical protein Ta0583 | | Authors: | Roeben, A, Kofler, C, Nagy, I, Nickell, S, Ulrich Hartl, F, Bracher, A. | | Deposit date: | 2006-01-23 | | Release date: | 2006-04-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of an archaeal actin homolog
J.Mol.Biol., 358, 2006
|
|
2FMM
 
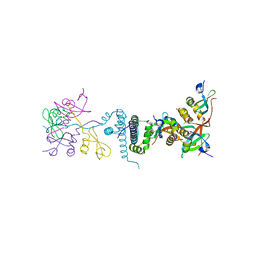 | | Crystal Structure of EMSY-HP1 complex | | Descriptor: | Chromobox protein homolog 1, Protein EMSY, SULFATE ION | | Authors: | Huang, Y. | | Deposit date: | 2006-01-09 | | Release date: | 2006-05-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the HP1-EMSY complex reveals an unusual mode of HP1 binding.
Structure, 14, 2006
|
|
2FSN
 
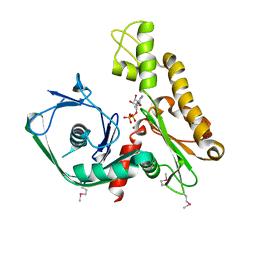 | | Crystal structure of Ta0583, an archaeal actin homolog, complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, hypothetical protein Ta0583 | | Authors: | Roeben, A, Kofler, C, Nagy, I, Nickell, S, Ulrich Hartl, F, Bracher, A. | | Deposit date: | 2006-01-23 | | Release date: | 2006-04-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of an archaeal actin homolog
J.Mol.Biol., 358, 2006
|
|
4RBL
 
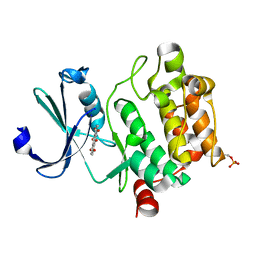 | | Crystal structure of Ser/Thr kinase Pim1 in complex with Mitoxantrone derivatives | | Descriptor: | 1,4,5,8-tetrahydroxyanthracene-9,10-dione, Serine/threonine-protein kinase pim-1 | | Authors: | Zhang, W, Wan, X, Huang, N. | | Deposit date: | 2014-09-12 | | Release date: | 2015-03-18 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of Ser/Thr kinase Pim1 in complex with Mitoxantrone derivatives
To be Published, 2014
|
|
4RC3
 
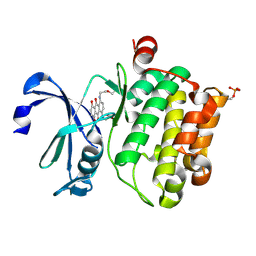 | |
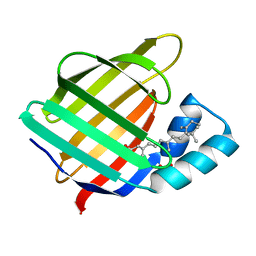
2FR3
 | |
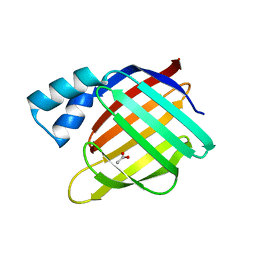
2FS7
 | |
2FXE
 
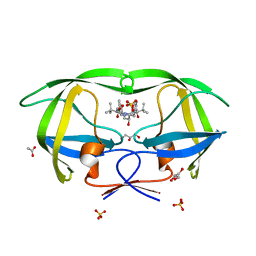 | | X-ray crystal structure of HIV-1 protease CRM mutant complexed with atazanavir (BMS-232632) | | Descriptor: | (3S,8S,9S,12S)-3,12-BIS(1,1-DIMETHYLETHYL)-8-HYDROXY-4,11-DIOXO-9-(PHENYLMETHYL)-6-[[4-(2-PYRIDINYL)PHENYL]METHYL]-2,5, 6,10,13-PENTAAZATETRADECANEDIOIC ACID DIMETHYL ESTER, ACETATE ION, ... | | Authors: | Sheriff, S, Klei, H.E. | | Deposit date: | 2006-02-05 | | Release date: | 2007-02-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structures of human immunodeficiency virus type 1 protease mutants complexed with atazanavir.
J.Virol., 81, 2007
|
|
2FQY
 
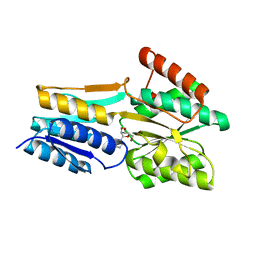 | | PnrA from Treponema pallidum complexed with adenosine. | | Descriptor: | ADENOSINE, Membrane lipoprotein tmpC | | Authors: | Brautigam, C.A, Deka, R.K, Tomchick, D.R, Machius, M, Norgard, M.V. | | Deposit date: | 2006-01-18 | | Release date: | 2006-02-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The PnrA (Tp0319; TmpC) lipoprotein represents a new family of bacterial purine nucleoside receptor encoded within an ATP-binding cassette (ABC)-like operon in Treponema pallidum
J.Biol.Chem., 281, 2006
|
|
7PW8
 
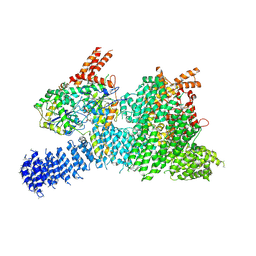 | | Human SMG1-8-9 kinase complex bound to AMPPNP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Langer, L.M, Conti, E. | | Deposit date: | 2021-10-06 | | Release date: | 2021-12-01 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (2.82 Å) | | Cite: | Cryo-EM reconstructions of inhibitor-bound SMG1 kinase reveal an autoinhibitory state dependent on SMG8.
Elife, 10, 2021
|
|
7PW4
 
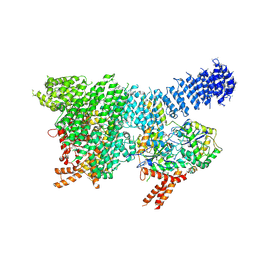 | | Human SMG1-8-9 kinase complex bound to a SMG1 inhibitor | | Descriptor: | 1-[4-[4-[2-[[4-chloranyl-3-(diethylsulfamoyl)phenyl]amino]pyrimidin-4-yl]pyridin-2-yl]phenyl]-3-methyl-urea, ADENOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, ... | | Authors: | Langer, L.M, Conti, E. | | Deposit date: | 2021-10-06 | | Release date: | 2021-12-01 | | Method: | ELECTRON MICROSCOPY (3.27 Å) | | Cite: | Cryo-EM reconstructions of inhibitor-bound SMG1 kinase reveal an autoinhibitory state dependent on SMG8.
Elife, 10, 2021
|
|
7PW9
 
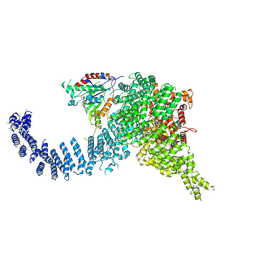 | | Human SMG1-9 kinase complex bound to AMPPNP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Langer, L.M, Conti, E. | | Deposit date: | 2021-10-06 | | Release date: | 2021-12-01 | | Method: | ELECTRON MICROSCOPY (3.12 Å) | | Cite: | Cryo-EM reconstructions of inhibitor-bound SMG1 kinase reveal an autoinhibitory state dependent on SMG8.
Elife, 10, 2021
|
|
7PW7
 
 | | Human SMG1-9 kinase complex bound to a SMG1 inhibitor | | Descriptor: | 1-[4-[4-[2-[[4-chloranyl-3-(diethylsulfamoyl)phenyl]amino]pyrimidin-4-yl]pyridin-2-yl]phenyl]-3-methyl-urea, ADENOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, ... | | Authors: | Langer, L.M, Conti, E. | | Deposit date: | 2021-10-06 | | Release date: | 2021-12-01 | | Method: | ELECTRON MICROSCOPY (3.59 Å) | | Cite: | Cryo-EM reconstructions of inhibitor-bound SMG1 kinase reveal an autoinhibitory state dependent on SMG8.
Elife, 10, 2021
|
|
7PW5
 
 | |
