1JKP
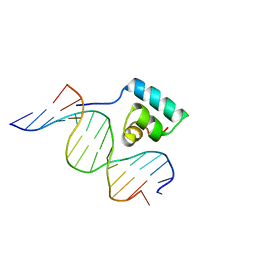 
 | | Testing the Water-Mediated HIN Recombinase DNA Recognition by Systematic Mutations | | Descriptor: | 5'-D(*AP*TP*CP*TP*TP*CP*TP*CP*AP*AP*AP*AP*AP*C)-3', 5'-D(*TP*GP*TP*TP*TP*TP*TP*GP*AP*GP*AP*AP*GP*A)-3', DNA-INVERTASE HIN | | Authors: | Chiu, T.K, Sohn, C, Johnson, R.C, Dickerson, R.E. | | Deposit date: | 2001-07-13 | | Release date: | 2002-02-22 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Testing water-mediated DNA recognition by the Hin recombinase.
EMBO J., 21, 2002
|
|
2F9V
 
 | | HCV NS3 protease domain with NS4a peptide and a ketoamide inhibitor with P1 and P2 cyclopropylalannines | | Descriptor: | (2S,8R,9S,15S)-15-CYCLOHEXYL-9,12-BIS(CYCLOPROPYLMETHYL)-8-HYDROXY-20-METHYL-4,7,11,14,17-PENTAOXO-2-PHENYL-18-OXA-3,6,10,12,13,16-HEXAAZAHENICOSAN-1-OIC ACID, NS3 protease/helicase, ZINC ION, ... | | Authors: | Bogen, S.L, Ruan, S, Liu, R, Agrawal, S, Pichardo, J, Prongay, A, Baroudy, B, Saksena, A, Girijavallabhan, V, Njoroge, F.G. | | Deposit date: | 2005-12-06 | | Release date: | 2007-01-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Depeptidization efforts on P3-P2 a-ketoamide inhibitors of HCV NS3-4A serine protease: Effect on HCV replicon activity.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
1MHT
 
 | |
2CV2
 
 | | Glutamyl-tRNA synthetase from Thermus thermophilus in complex with tRNA(Glu) and an enzyme inhibitor, Glu-AMS | | Descriptor: | CHLORIDE ION, MAGNESIUM ION, O5'-(L-GLUTAMYL-SULFAMOYL)-ADENOSINE, ... | | Authors: | Sekine, S, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-31 | | Release date: | 2006-09-05 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.69 Å) | | Cite: | Structural bases of transfer RNA-dependent amino acid recognition and activation by glutamyl-tRNA synthetase
Structure, 14, 2006
|
|
2FIV
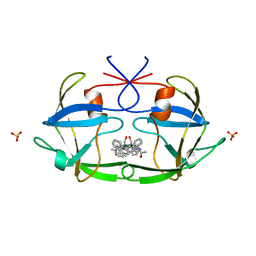 
 | | Crystal structure of feline immunodeficiency virus protease complexed with a substrate | | Descriptor: | ACE-ALN-VAL-STA-GLU-ALN-NH2, FELINE IMMUNODEFICIENCY VIRUS PROTEASE, SULFATE ION | | Authors: | Schalk-Hihi, C, Lubkowski, J, Zdanov, A, Wlodawer, A, Gustchina, A. | | Deposit date: | 1997-07-21 | | Release date: | 1997-11-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of the inactive D30N mutant of feline immunodeficiency virus protease complexed with a substrate and an inhibitor.
Biochemistry, 36, 1997
|
|
2FHI
 
 | | SUBSTRATE ANALOG (IB2) COMPLEX WITH THE HIS 96 ASN SUBSTITUTION OF THE FRAGILE HISTIDINE TRIAD PROTEIN, FHIT | | Descriptor: | FRAGILE HISTIDINE TRIAD PROTEIN, P1-P2-METHYLENE-P3-THIO-DIADENOSINE TRIPHOSPHATE | | Authors: | Pace, H.C, Garrison, P.N, Robinson, A.K, Barnes, L.D, Draganescu, A, Rosler, A, Blackburn, G.M, Siprashvili, Z, Croce, C.M, Heubner, K, Brenner, C. | | Deposit date: | 1998-04-01 | | Release date: | 1998-06-17 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Genetic, biochemical, and crystallographic characterization of Fhit-substrate complexes as the active signaling form of Fhit.
Proc.Natl.Acad.Sci.USA, 95, 1998
|
|
2YBK
 
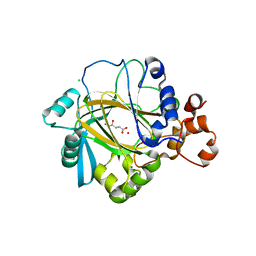 | | JMJD2A COMPLEXED WITH R-2-HYDROXYGLUTARATE | | Descriptor: | (2R)-2-hydroxypentanedioic acid, CHLORIDE ION, LYSINE-SPECIFIC DEMETHYLASE 4A, ... | | Authors: | Chowdhury, R, McDonough, M.A, Schofield, C.J. | | Deposit date: | 2011-03-08 | | Release date: | 2011-04-06 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The oncometabolite 2-hydroxyglutarate inhibits histone lysine demethylases.
EMBO Rep., 12, 2011
|
|
2YQS
 
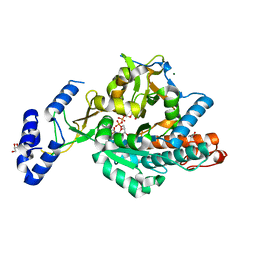 | | Crystal structure of uridine-diphospho-N-acetylglucosamine pyrophosphorylase from Candida albicans, in the product-binding form | | Descriptor: | GLYCEROL, MAGNESIUM ION, SULFATE ION, ... | | Authors: | Miki, K, Maruyama, D, Nishitani, Y, Nonaka, T, Kita, A. | | Deposit date: | 2007-03-30 | | Release date: | 2007-05-22 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Uridine-diphospho-N-acetylglucosamine Pyrophosphorylase from Candida albicans and Catalytic Reaction Mechanism
J.Biol.Chem., 282, 2007
|
|
2YSK
 
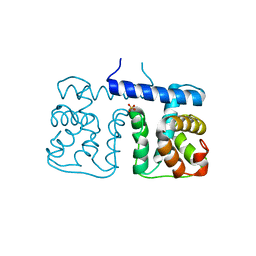 | |
2YUT
 
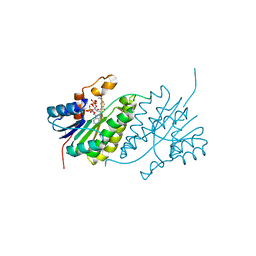 | |
7VSI
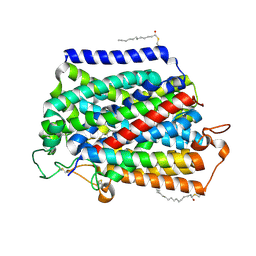 
 | | Structure of human SGLT2-MAP17 complex bound with empagliflozin | | Descriptor: | (2S,3R,4R,5S,6R)-2-[4-chloranyl-3-[[4-[(3S)-oxolan-3-yl]oxyphenyl]methyl]phenyl]-6-(hydroxymethyl)oxane-3,4,5-triol, PALMITIC ACID, PDZK1-interacting protein 1, ... | | Authors: | Chen, L, Niu, Y, Liu, R. | | Deposit date: | 2021-10-26 | | Release date: | 2021-12-15 | | Last modified: | 2022-02-16 | | Method: | ELECTRON MICROSCOPY (2.95 Å) | | Cite: | Structural basis of inhibition of the human SGLT2-MAP17 glucose transporter.
Nature, 601, 2022
|
|
2YVW
 
 | |
2YWC
 
 | | Crystal structure of GMP synthetase from Thermus thermophilus in complex with XMP | | Descriptor: | GMP synthase [glutamine-hydrolyzing], XANTHOSINE-5'-MONOPHOSPHATE | | Authors: | Baba, S, Kanagawa, M, Kuramitsu, S, Yokoyama, S, Sampei, G, Kawai, G, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-04-20 | | Release date: | 2007-10-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of GMP synthetase from Thermus thermophilus
To be Published
|
|
2YT4
 
 | |
2YZM
 
 | |
2Z06
 
 | |
2YWA
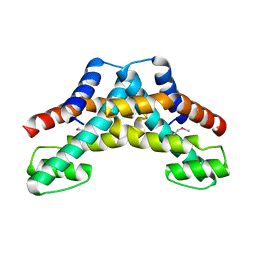 
 | |
2YWN
 
 | | Crystal structure of peroxiredoxin-like protein from Sulfolobus tokodaii | | Descriptor: | Peroxiredoxin-like protein | | Authors: | Ebihara, A, Manzoku, M, Fujimoto, Y, Yokoyama, S, Kuramitsu, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-04-20 | | Release date: | 2007-10-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of peroxiredoxin-like protein from Sulfolobus tokodaii
To be Published
|
|
2YWX
 
 | | Crystal structure of phosphoribosylaminoimidazole carboxylase catalytic subunit from Methanocaldococcus jannaschii | | Descriptor: | Phosphoribosylaminoimidazole carboxylase catalytic subunit | | Authors: | Kanagawa, M, Baba, S, Kuramitsu, S, Yokoyama, S, Kawai, G, Sampei, G, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-04-23 | | Release date: | 2007-10-23 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Crystal structure of phosphoribosylaminoimidazole carboxylase catalytic subunit from Methanocaldococcus jannaschii
To be Published
|
|
2YX5
 
 | | Crystal Structure of Methanocaldococcus jannaschii PurS, One of the Subunits of Formylglycinamide Ribonucleotide Amidotransferase in the Purine Biosynthetic Pathway | | Descriptor: | UPF0062 protein MJ1593 | | Authors: | Kanagawa, M, Baba, S, Agari, Y, Chen, L.Q, Fu, Z.-Q, Chrzas, J, Wang, B.C, Kuramitsu, S, Yokoyama, S, Kawai, G, Sampei, G, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-04-24 | | Release date: | 2007-10-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Methanocaldococcus jannaschii PurS, One of the Subunits of Formylglycinamide Ribonucleotide Amidotransferase in the Purine Biosynthetic Pathway
To be Published
|
|
2YX9
 
 | | Crystal structure of D298K copper amine oxidase from Arthrobacter globiformis | | Descriptor: | COPPER (II) ION, Phenylethylamine oxidase | | Authors: | Murakawa, T, Okajima, T, Yamaguchi, H, Tanizawa, K. | | Deposit date: | 2007-04-25 | | Release date: | 2008-04-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Trapping of a dopaquinone intermediate in the TPQ cofactor biogenesis in a copper-containing amine oxidase from Arthrobacter globiformis.
J.Am.Chem.Soc., 129, 2007
|
|
2YYH
 
 | |
2Z13
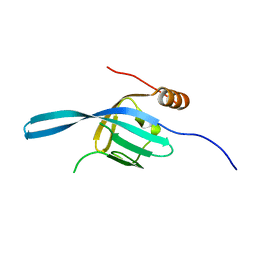 
 | | Crystal structure of the N-terminal DUF1126 in human EF-hand domain | | Descriptor: | EF-hand domain-containing family member C2 | | Authors: | Saito, K, Kishishita, S, Nishino, A, Murayama, K, Terada, T, Shirouzu, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-05-07 | | Release date: | 2007-11-13 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal structure of the N-terminal DUF1126 in human EF-hand domain
To be Published
|
|
1NG7
 
 | |
2YWI
 
 | |
