1MNU
 
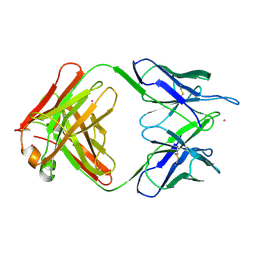 | | UNLIGANDED BACTERICIDAL ANTIBODY AGAINST NEISSERIA MENINGITIDIS | | Descriptor: | CADMIUM ION, PROTEIN (IGG2A-KAPPA ANTIBODY MN12H2 (HEAVY CHAIN)), PROTEIN (IGG2A-KAPPA ANTIBODY MN12H2 (LIGHT CHAIN)) | | Authors: | Van Den Elsen, J, Vandeputte-Rutten, L, Kroon, J, Gros, P. | | Deposit date: | 1999-04-30 | | Release date: | 1999-05-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Bactericidal antibody recognition of meningococcal PorA by induced fit. Comparison of liganded and unliganded Fab structures.
J.Biol.Chem., 274, 1999
|
|
5I8D
 
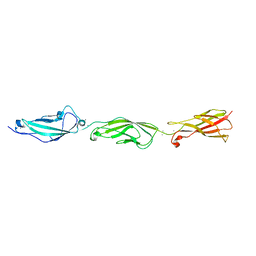 | |
3P1I
 
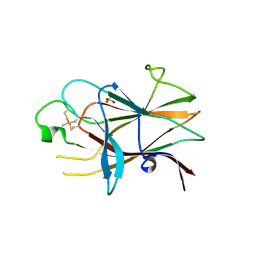 | | Ligand binding domain of human ephrin type-B receptor 3 | | Descriptor: | CHLORIDE ION, Ephrin type-B receptor 3, SULFATE ION | | Authors: | Walker, J.R, Yermekbayeva, L, Seitova, A, Kania, J, Bountra, C, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Dhe-Paganon, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-09-30 | | Release date: | 2011-01-19 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Ligand Binding Domain of Human Ephb3
To be Published
|
|
3V58
 
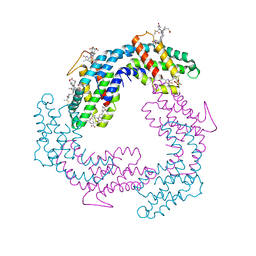 | |
7N6P
 
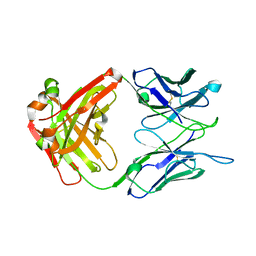 | | Crystal structure of the anti-EBOV and SUDV monoclonal antibody 1C3 Fab | | Descriptor: | 1C3 Fab heavy chain, 1C3 Fab light chain | | Authors: | Milligan, J.C, Yu, X, Buck, T, Saphire, E.O. | | Deposit date: | 2021-06-08 | | Release date: | 2022-04-06 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Asymmetric and non-stoichiometric glycoprotein recognition by two distinct antibodies results in broad protection against ebolaviruses.
Cell, 185, 2022
|
|
3V57
 
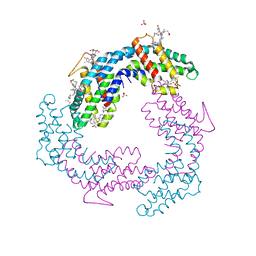 | |
4CGL
 
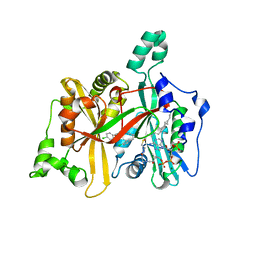 | | Leishmania major N-myristoyltransferase in complex with an aminoacylpyrrolidine inhibitor | | Descriptor: | (3R)-3-azanyl-4-(4-chlorophenyl)-1-[(3S,4R)-3-(4-chlorophenyl)-4-(hydroxymethyl)pyrrolidin-1-yl]butan-1-one, GLYCYLPEPTIDE N-TETRADECANOYLTRANSFERASE, MAGNESIUM ION, ... | | Authors: | Brannigan, J.A, Roberts, S.M, Bell, A.S, Hutton, J.A, Smith, D.F, Tate, E.W, Leatherbarrow, R.J, Wilkinson, A.J. | | Deposit date: | 2013-11-25 | | Release date: | 2014-07-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Diverse Modes of Binding in Structures of Leishmania Major N-Myristoyltransferase with Selective Inhibitors
Iucrj, 1, 2014
|
|
4CGN
 
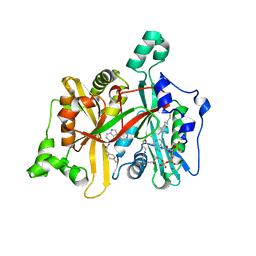 | | Leishmania major N-myristoyltransferase in complex with a piperidinylindole inhibitor | | Descriptor: | 2-(4-fluorophenyl)-N-(3-piperidin-4-yl-1H-indol-5-yl)ethanamide, GLYCYLPEPTIDE N-TETRADECANOYLTRANSFERASE, MAGNESIUM ION, ... | | Authors: | Brannigan, J.A, Roberts, S.M, Bell, A.S, Hutton, J.A, Smith, D.F, Tate, E.W, Leatherbarrow, R.J, Wilkinson, A.J. | | Deposit date: | 2013-11-25 | | Release date: | 2014-07-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Diverse Modes of Binding in Structures of Leishmania Major N-Myristoyltransferase with Selective Inhibitors
Iucrj, 1, 2014
|
|
6KK9
 
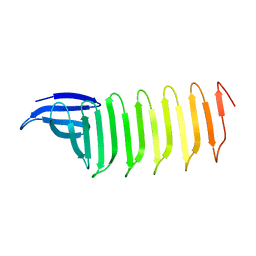 | | A Crystal structure of OspA mutant | | Descriptor: | Outer Surface Protein A | | Authors: | Shiga, S, Makabe, K. | | Deposit date: | 2019-07-24 | | Release date: | 2020-07-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural analysis of the beta-sheet edge of peptide self-assembly using a model protein.
Proteins, 89, 2021
|
|
6AIS
 
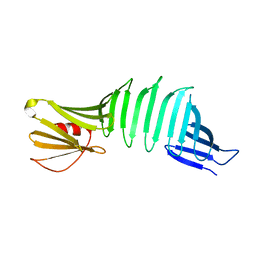 | |
1D1K
 
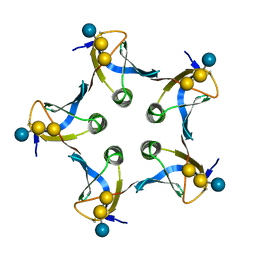 | |
6BEB
 
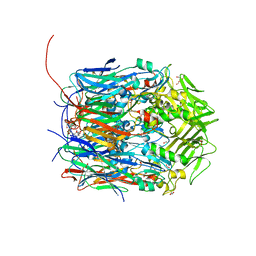 | | Crystal structure of VACV D13 in complex with Rifamycin SV | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, Scaffold protein D13, ... | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6BEF
 
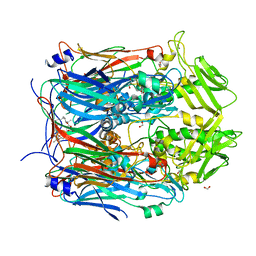 | | Crystal structure of VACV D13 in complex with 3-formyl rifamycin SV | | Descriptor: | (2S,12Z,14E,16S,17S,18R,19R,20R,21S,22R,23S,24E)-8-formyl-5,6,9,17,19-pentahydroxy-23-methoxy-2,4,12,16,18,20,22-heptam ethyl-1,11-dioxo-1,2-dihydro-2,7-(epoxypentadeca[1,11,13]trienoimino)naphtho[2,1-b]furan-21-yl acetate, 1,2-ETHANEDIOL, FORMIC ACID, ... | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.21 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
1CZG
 
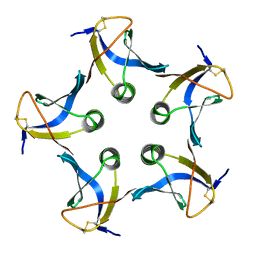 | |
6BEE
 
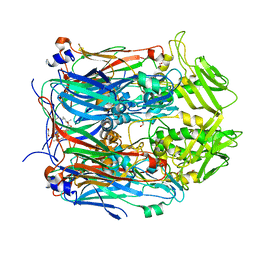 | | Crystal structure of VACV D13 in complex with Rifaximin | | Descriptor: | (2S,16Z,18E,20S,21S,22R,23R,24R,25S,26R,27S,28E)-5,6,21,23-tetrahydroxy-27-methoxy-2,4,11,16,20,22,24,26-octamethyl-1,1 5-dioxo-1,2-dihydro-2,7-(epoxypentadeca[1,11,13]trienoimino)furo[2'',3'':7',8']naphtho[1',2':4,5]imidazo[1,2-a]pyridin-2 5-yl acetate, 1,2-ETHANEDIOL, FORMIC ACID, ... | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
4LE9
 
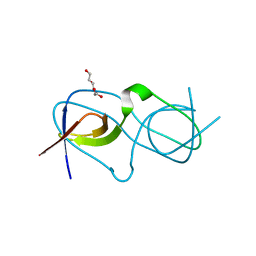 | | Crystal structure of a chimeric c-Src-SH3 domain | | Descriptor: | Proto-oncogene tyrosine-protein kinase Src, TRIETHYLENE GLYCOL | | Authors: | Camara-Artigas, A, Martinez-Rodriguez, S, Ortiz-Salmeron, E, Martin-Garcia, J.M. | | Deposit date: | 2013-06-25 | | Release date: | 2014-05-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.344 Å) | | Cite: | 3D domain swapping in a chimeric c-Src SH3 domain takes place through two hinge loops.
J.Struct.Biol., 186, 2014
|
|
6BED
 
 | | Crystal structure of VACV D13 in complex with Rifampicin | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, RIFAMPICIN, ... | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6BEI
 
 | | Crystal structure of VACV D13 in its apo (unbound) form | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, Scaffold protein D13 | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3KLR
 
 | | Bovine H-protein at 0.88 angstrom resolution | | Descriptor: | GLYCEROL, Glycine cleavage system H protein, SULFATE ION | | Authors: | Higashiura, A, Kurakane, T, Matsuda, M, Suzuki, M, Inaka, K, Sato, M, Tanaka, H, Fujiwara, K, Nakagawa, A. | | Deposit date: | 2009-11-09 | | Release date: | 2010-06-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (0.88 Å) | | Cite: | High-resolution X-ray crystal structure of bovine H-protein at 0.88 A resolution
Acta Crystallogr.,Sect.D, 66, 2010
|
|
6BEC
 
 | | Crystal structure of VACV D13 in complex with Rifabutin | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, RIFABUTIN, ... | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6BEG
 
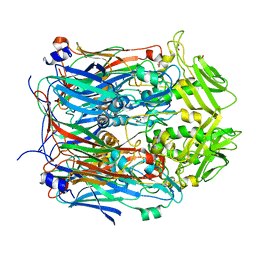 | | Crystal structure of VACV D13 F486A mutant | | Descriptor: | 1,2-ETHANEDIOL, FORMIC ACID, Scaffold protein D13 | | Authors: | Garriga, D, Accurso, C, Coulibaly, F. | | Deposit date: | 2017-10-25 | | Release date: | 2018-07-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural basis for the inhibition of poxvirus assembly by the antibiotic rifampicin.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
5CVZ
 
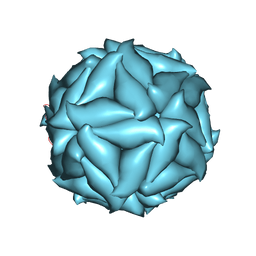 | |
3NF3
 
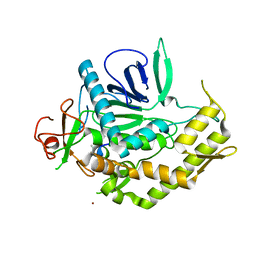 | | Crystal structure of BoNT/A LC with JTH-NB-7239 peptide | | Descriptor: | BoNT/A, JTH-NB72-39 inhibitor, NICKEL (II) ION, ... | | Authors: | Zuniga, J.E. | | Deposit date: | 2010-06-09 | | Release date: | 2010-07-21 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Iterative structure-based peptide-like inhibitor design against the botulinum neurotoxin serotype A.
Plos One, 5, 2010
|
|
3NGG
 
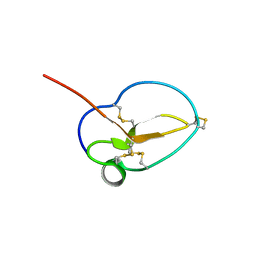 | | X-ray Structure of Omwaprin | | Descriptor: | Omwaprin-a | | Authors: | Banigan, J.R, Mandal, K, Sawaya, M.R, Thammavongsa, V, Hendrickx, A.P.A, Schneewind, O, Yeates, T.O, Kent, S.B.H. | | Deposit date: | 2010-06-11 | | Release date: | 2010-09-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.33 Å) | | Cite: | Determination of the X-ray structure of the snake venom protein omwaprin by total chemical synthesis and racemic protein crystallography.
Protein Sci., 19, 2010
|
|
1ZMC
 
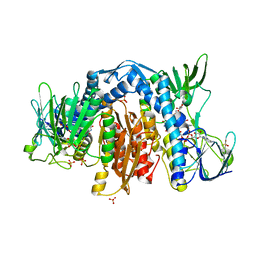 | | Crystal Structure of Human dihydrolipoamide dehydrogenase complexed to NAD+ | | Descriptor: | Dihydrolipoyl dehydrogenase, mitochondrial, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Brautigam, C.A, Chuang, J.L, Tomchick, D.R, Machius, M, Chuang, D.T. | | Deposit date: | 2005-05-10 | | Release date: | 2005-06-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Crystal Structure of Human Dihydrolipoamide Dehydrogenase: NAD(+)/NADH Binding and the Structural Basis of Disease-causing Mutations
J.Mol.Biol., 350, 2005
|
|
