2H8L
 
 | |
1ZR0
 
 | | Crystal Structure of Kunitz Domain 1 of Tissue Factor Pathway Inhibitor-2 with Bovine Trypsin | | Descriptor: | CALCIUM ION, Cationic trypsin, Tissue factor pathway inhibitor 2 | | Authors: | Schmidt, A.E, Chand, H.S, Cascio, D, Kisiel, W, Bajaj, S.P. | | Deposit date: | 2005-05-18 | | Release date: | 2005-06-07 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.802 Å) | | Cite: | Crystal structure of Kunitz domain 1 (KD1) of tissue factor pathway inhibitor-2 in complex with trypsin. Implications for KD1 specificity of inhibition
J.Biol.Chem., 280, 2005
|
|
2H8Q
 
 | |
1K1X
 
 | | Crystal structure of 4-alpha-glucanotransferase from thermococcus litoralis | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 4-ALPHA-GLUCANOTRANSFERASE, CALCIUM ION | | Authors: | Imamura, H, Fushinobu, S, Kumasaka, T, Yamamoto, M, Jeon, B.S, Wakagi, T, Matsuzawa, H. | | Deposit date: | 2001-09-26 | | Release date: | 2003-06-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structures of 4-alpha-glucanotransferase from Thermococcus litoralis and its complex with an inhibitor
J.BIOL.CHEM., 278, 2003
|
|
2GQR
 
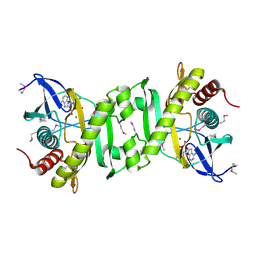 | | SAICAR Synthetase Complexed with ADP-Mg2+ | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Phosphoribosylaminoimidazole-succinocarboxamide synthase | | Authors: | Ginder, N.D, Honzatko, R.B. | | Deposit date: | 2006-04-21 | | Release date: | 2006-05-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Nucleotide Complexes of Escherichia coli Phosphoribosylaminoimidazole Succinocarboxamide Synthetase.
J.Biol.Chem., 281, 2006
|
|
2HDZ
 
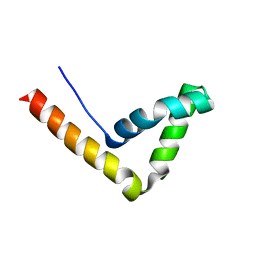 | | Crystal Structure Analysis of the UBF HMG box5 | | Descriptor: | Nucleolar transcription factor 1 | | Authors: | Rong, H, Teng, M.K, Niu, L.W. | | Deposit date: | 2006-06-21 | | Release date: | 2007-06-26 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of human upstream binding factor HMG box 5 and site for binding of the cell-cycle regulatory factor TAF1
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2GUV
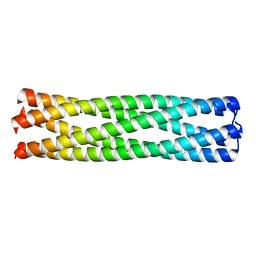 
 | | Conformational Transition between Four- and Five-stranded Phenylalanine Zippers Determined by a Local Packing Interaction | | Descriptor: | Major outer membrane lipoprotein | | Authors: | Liu, J, Zheng, Q, Deng, Y, Kallenbach, N.R, Lu, M. | | Deposit date: | 2006-05-01 | | Release date: | 2006-07-25 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Conformational Transition between Four and Five-stranded Phenylalanine Zippers Determined by a Local Packing Interaction.
J.Mol.Biol., 361, 2006
|
|
2GUS
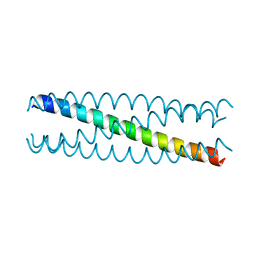 
 | | Conformational Transition between Four- and Five-stranded Phenylalanine Zippers Determined by a Local Packing Interaction | | Descriptor: | Major outer membrane lipoprotein | | Authors: | Liu, J, Zheng, Q, Deng, Y, Kallenbach, N.R, Lu, M. | | Deposit date: | 2006-05-01 | | Release date: | 2006-07-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Conformational Transition between Four and Five-stranded Phenylalanine Zippers Determined by a Local Packing Interaction.
J.Mol.Biol., 361, 2006
|
|
1V0J
 
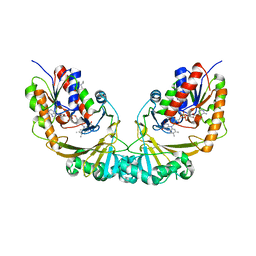 | | Udp-galactopyranose mutase from Mycobacterium tuberculosis | | Descriptor: | BICINE, FLAVIN-ADENINE DINUCLEOTIDE, UDP-GALACTOPYRANOSE MUTASE | | Authors: | Beis, K, Naismith, J.H. | | Deposit date: | 2004-03-30 | | Release date: | 2005-01-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal structures of Mycobacteria tuberculosis and Klebsiella pneumoniae UDP-galactopyranose mutase in the oxidised state and Klebsiella pneumoniae UDP-galactopyranose mutase in the (active) reduced state.
J. Mol. Biol., 348, 2005
|
|
1LKY
 
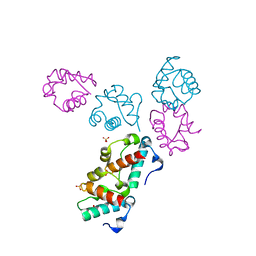 | | Structure of the wild-type TEL-SAM polymer | | Descriptor: | SULFATE ION, TRANSCRIPTION FACTOR ETV6 | | Authors: | Tran, H.H, Kim, C.A, Faham, S, Bowie, J.U. | | Deposit date: | 2002-04-26 | | Release date: | 2002-06-12 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Native interface of the SAM domain polymer of TEL.
BMC STRUCT.BIOL., 2, 2002
|
|
1YO6
 
 | |
2HMD
 
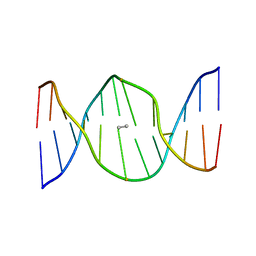 | | Stereochemistry Modulates Stability of Reduced Inter-Strand Cross-Links Arising From R- and S-alpha-methyl-gamma-OH-1,N2-propano-2'-Deoxyguanine in the 5'-CpG-3' DNA Sequence | | Descriptor: | DNA dodecamer with interstrand cross-link | | Authors: | Cho, Y.-J, Kozekov, I.D, Harris, T.M, Rizzo, C.J, Stone, M.P. | | Deposit date: | 2006-07-11 | | Release date: | 2007-03-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Stereochemistry Modulates the Stability of Reduced Interstrand Cross-Links Arising from R- and S-alpha-CH(3)-gamma-OH-1,N(2)-Propano-2'-deoxyguanosine in the 5'-CpG-3' DNA Sequence
Biochemistry, 46, 2007
|
|
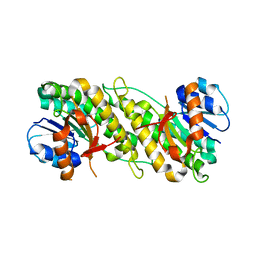
1YNL
 
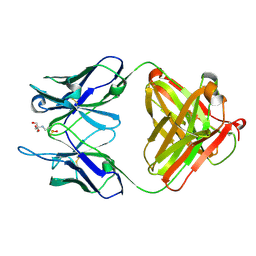 | | Identification of Key residues of the NC6.8 Fab antibody fragment binding to synthetic sweeterners: Crystal structure of NC6.8 co-crystalized with high potency sweetener compound SC45647 | | Descriptor: | 2-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-ETHANESULFONIC ACID, Ig gamma heavy chain, Ig gamma light chain | | Authors: | Gokulan, K, Khare, S, Ronning, D.R, Linthicum, S.D, Sacchettini, J.C, Rupp, B. | | Deposit date: | 2005-01-24 | | Release date: | 2005-08-16 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Cocrystal Structures of NC6.8 Fab Identify Key Interactions for High Potency Sweetener Recognition: Implications for the Design of Synthetic Sweeteners
Biochemistry, 44, 2005
|
|
2HO1
 
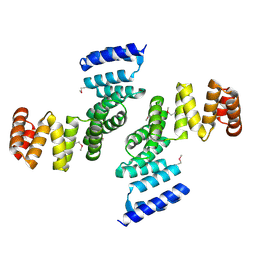 | | Functional Characterization of Pseudomonas Aeruginosa pilF | | Descriptor: | Type 4 fimbrial biogenesis protein PilF | | Authors: | Koo, J. | | Deposit date: | 2006-07-13 | | Release date: | 2006-07-25 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | PilF is an outer membrane lipoprotein required for multimerization and localization of the Pseudomonas aeruginosa Type IV pilus secretin.
J.Bacteriol., 190, 2008
|
|
1SZM
 
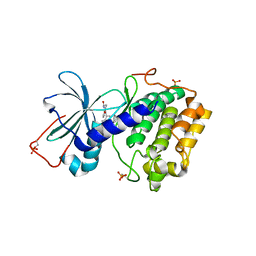 | | DUAL BINDING MODE OF BISINDOLYLMALEIMIDE 2 TO PROTEIN KINASE A (PKA) | | Descriptor: | 3-(1H-INDOL-3-YL)-4-{1-[2-(1-METHYLPYRROLIDIN-2-YL)ETHYL]-1H-INDOL-3-YL}-1H-PYRROLE-2,5-DIONE, cAMP-dependent protein kinase, alpha-catalytic subunit | | Authors: | Gassel, M, Breitenlechner, C.B, Koenig, N, Huber, R, Engh, R.A, Bossemeyer, D. | | Deposit date: | 2004-04-06 | | Release date: | 2004-06-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The protein kinase C inhibitor bisindolyl maleimide 2 binds with reversed orientations to different conformations of protein kinase a.
J.Biol.Chem., 279, 2004
|
|
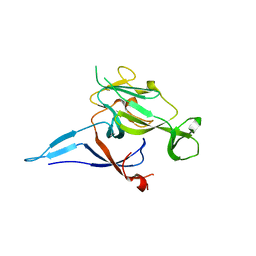
1KN9
 | |
2KYU
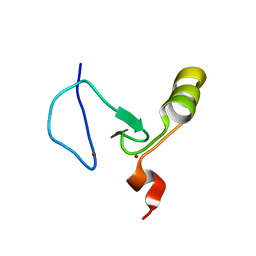 
 | | The solution structure of the PHD3 finger of MLL | | Descriptor: | Histone-lysine N-methyltransferase MLL, ZINC ION | | Authors: | Park, S, Bushweller, J.H. | | Deposit date: | 2010-06-08 | | Release date: | 2010-08-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The PHD3 domain of MLL acts as a CYP33-regulated switch between MLL-mediated activation and repression.
Biochemistry, 49, 2010
|
|
2KV6
 
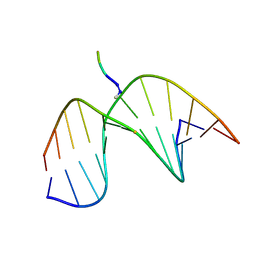 | | Tetrapeptide KWKK conjugated to oligonucleotide duplex by a trimethylene tether | | Descriptor: | 5'-D(*GP*CP*TP*AP*GP*CP*GP*AP*GP*TP*CP*C)-3', 5'-D(*GP*GP*AP*CP*TP*CP*GP*CP*TP*AP*GP*C)-3', KWKK Tetrapeptide | | Authors: | Huang, H, Kozekov, I.D, Kozekova, A, Rizzo, C.J, McCullough, A, LLoyd, R.S, Stone, M.P. | | Deposit date: | 2010-03-08 | | Release date: | 2010-07-21 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Minor Groove Orientation of the KWKK Peptide Tethered via the N-Terminal Amine to the Acrolein-Derived 1,N(2)-gamma-Hydroxypropanodeoxyguanosine Lesion with a Trimethylene Linkage .
Biochemistry, 49, 2010
|
|
2KFS
 
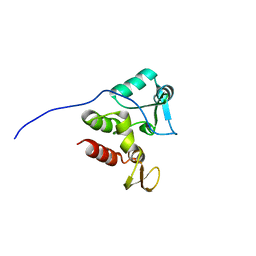 | | NMR structure of Rv2175c | | Descriptor: | Conserved hypothetical regulatory protein | | Authors: | Barthe, P, Cohen-Gonsaud, M, Roumestand, C, Molle, V. | | Deposit date: | 2009-02-27 | | Release date: | 2009-05-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | The Mycobacterium tuberculosis Ser/Thr Kinase Substrate Rv2175c Is a DNA-binding Protein Regulated by Phosphorylation.
J.Biol.Chem., 284, 2009
|
|
1II5
 
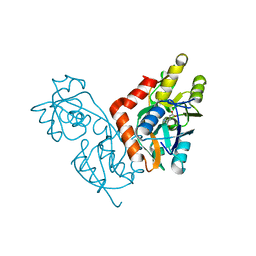 | |
1T95
 
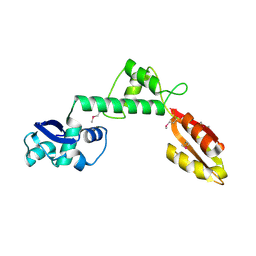 | |
2I67
 
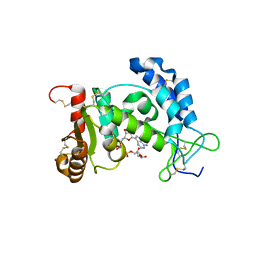 | | Structural Basis for the Mechanistic Understanding Human CD38 Controlled Multiple Catalysis | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, ADP-ribosyl cyclase 1 | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Munshi, C, Lee, H.C, Hao, Q. | | Deposit date: | 2006-08-28 | | Release date: | 2006-09-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Structural basis for the mechanistic understanding of human CD38-controlled multiple catalysis.
J.Biol.Chem., 281, 2006
|
|
2I66
 
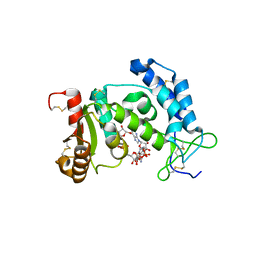 | | Structural Basis for the Mechanistic Understanding Human CD38 Controlled Multiple Catalysis | | Descriptor: | ADP-ribosyl cyclase 1, [(2R,3R,4R,5R)-5-(2-AMINO-6-OXO-1,6-DIHYDRO-9H-PURIN-9-YL)-3,4-DIHYDROXYTETRAHYDROFURAN-2-YL]METHYL [(2R,3S,4R,5S)-3,4,5-TRIHYDROXYTETRAHYDROFURAN-2-YL]METHYL DIHYDROGEN DIPHOSPHATE, [(2R,3R,4R,5R)-5-(2-AMINO-6-OXO-1,6-DIHYDRO-9H-PURIN-9-YL)-3,4-DIHYDROXYTETRAHYDROFURAN-2-YL]METHYL [(2R,3S,4S)-3,4-DIHYDROXYTETRAHYDROFURAN-2-YL]METHYL DIHYDROGEN DIPHOSPHATE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Munshi, C, Lee, H.C, Hao, Q. | | Deposit date: | 2006-08-28 | | Release date: | 2006-09-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for the mechanistic understanding of human CD38-controlled multiple catalysis.
J.Biol.Chem., 281, 2006
|
|
1K34
 
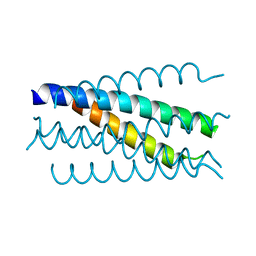 | | Crystal structure analysis of gp41 core mutant | | Descriptor: | Transmembrane glycoprotein GP41 | | Authors: | Shu, W, Lu, M. | | Deposit date: | 2001-10-01 | | Release date: | 2001-10-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Interhelical interactions in the gp41 core: implications for activation of HIV-1 membrane fusion.
Biochemistry, 41, 2002
|
|
2H32
 
 | | Crystal structure of the pre-B cell receptor | | Descriptor: | Immunoglobulin heavy chain, Immunoglobulin iota chain, Immunoglobulin omega chain, ... | | Authors: | Bankovich, A.J. | | Deposit date: | 2006-05-22 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural Insight into Pre-B Cell Receptor Function
Science, 316, 2007
|
|
