8YE9
 
 | |
8YE6
 
 | |
8YEO
 
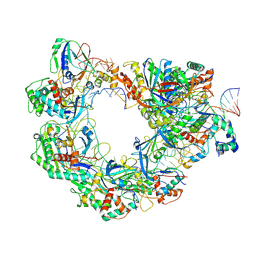 | | Type I-FHNH Cascade-dsDNA R-loop complex | | Descriptor: | 60-nt crRNA, Cas5f, Cas6f, ... | | Authors: | Li, Z. | | Deposit date: | 2024-02-22 | | Release date: | 2024-07-31 | | Last modified: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (3.44 Å) | | Cite: | Mechanisms for HNH-mediated target DNA cleavage in type I CRISPR-Cas systems.
Mol.Cell, 84, 2024
|
|
8S55
 
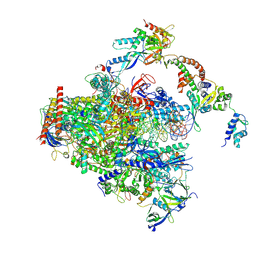 | | RNA polymerase II early elongation complex bound to TFIIE and TFIIF - state a (composite structure) | | Descriptor: | DNA-directed RNA polymerase II subunit E, DNA-directed RNA polymerase II subunit RPB11-a, DNA-directed RNA polymerase II subunit RPB3, ... | | Authors: | Zhan, Y, Grabbe, F, Oberbeckmann, E, Dienemann, C, Cramer, P. | | Deposit date: | 2024-02-22 | | Release date: | 2024-04-17 | | Last modified: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Three-step mechanism of promoter escape by RNA polymerase II.
Mol.Cell, 84, 2024
|
|
8S51
 
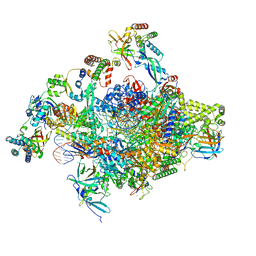 | | RNA polymerase II core initially transcribing complex with an ordered RNA of 8 nt | | Descriptor: | DNA-directed RNA polymerase II subunit E, DNA-directed RNA polymerase II subunit RPB11-a, DNA-directed RNA polymerase II subunit RPB3, ... | | Authors: | Zhan, Y, Grabbe, F, Oberbeckmann, E, Dienemann, C, Cramer, P. | | Deposit date: | 2024-02-22 | | Release date: | 2024-04-17 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Three-step mechanism of promoter escape by RNA polymerase II.
Mol.Cell, 84, 2024
|
|
8S52
 
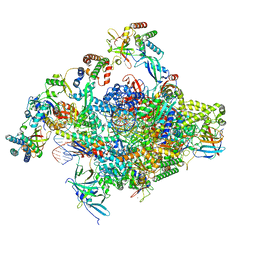 | | RNA polymerase II core initially transcribing complex with an ordered RNA of 10 nt | | Descriptor: | DNA-directed RNA polymerase II subunit E, DNA-directed RNA polymerase II subunit RPB11-a, DNA-directed RNA polymerase II subunit RPB3, ... | | Authors: | Zhan, Y, Grabbe, F, Oberbeckmann, E, Dienemann, C, Cramer, P. | | Deposit date: | 2024-02-22 | | Release date: | 2024-04-17 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Three-step mechanism of promoter escape by RNA polymerase II.
Mol.Cell, 84, 2024
|
|
8S54
 
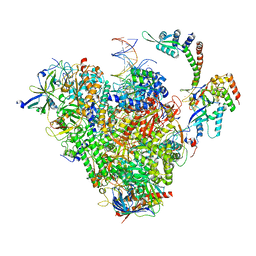 | | RNA polymerase II early elongation complex bound to TFIIE and TFIIF - state b (composite structure) | | Descriptor: | DNA-directed RNA polymerase II subunit E, DNA-directed RNA polymerase II subunit RPB11-a, DNA-directed RNA polymerase II subunit RPB3, ... | | Authors: | Zhan, Y, Grabber, F, Oberbeckmann, E, Dienemann, C, Cramer, P. | | Deposit date: | 2024-02-22 | | Release date: | 2024-04-17 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Three-step mechanism of promoter escape by RNA polymerase II.
Mol.Cell, 84, 2024
|
|
8S4T
 
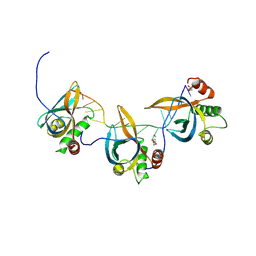 | |
8S4S
 
 | | PrgE from plasmid pCF10 | | Descriptor: | POTASSIUM ION, PrgE, SULFATE ION | | Authors: | Breidenstein, A, Berntsson, R.P.-A. | | Deposit date: | 2024-02-22 | | Release date: | 2024-05-29 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | PrgE: an OB-fold protein from plasmid pCF10 with striking differences to prototypical bacterial SSBs.
Life Sci Alliance, 7, 2024
|
|
8W35
 
 | | Aca2 from Pectobacterium phage ZF40 bound to RNA | | Descriptor: | Anti-CRISPR associated (Aca) protein, Aca2, IR2 and IR-RBS RNA | | Authors: | Wilkinson, M.E, Birkholz, N, Kimanius, D, Fineran, P.C. | | Deposit date: | 2024-02-21 | | Release date: | 2024-07-24 | | Last modified: | 2024-08-14 | | Method: | ELECTRON MICROSCOPY (2.61 Å) | | Cite: | Phage anti-CRISPR control by an RNA- and DNA-binding helix-turn-helix protein.
Nature, 631, 2024
|
|
8W2Q
 
 | | BsaXI-DNA complex II | | Descriptor: | DNA (41-MER), RM.BsaXI, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Shen, B.W, Stoddard, B.L, Xu, S. | | Deposit date: | 2024-02-21 | | Release date: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Structural and biochemical analysis of subunit assembly, DNA recognition and cleavage by a Type IIB restriction-modification enzyme: BsaXI
To Be Published
|
|
8YE4
 
 | | The complex of TCR NYN-I and HLA-A24 bound to SARS-CoV-2 Spike448-456 peptide NYNYLYRLF | | Descriptor: | Beta-2-microglobulin, MHC class I antigen precusor, Spike protein S1, ... | | Authors: | Deng, S.S, Jin, T.C, Xu, Z.H, Wang, M.H. | | Deposit date: | 2024-02-21 | | Release date: | 2024-08-21 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural insights into immune escape at killer T cell epitope by SARS-CoV-2 Spike Y453F variants.
J.Biol.Chem., 300, 2024
|
|
8W2W
 
 | | Structure of transthyretin synthetic mutation A120L | | Descriptor: | Transthyretin | | Authors: | Yang, K, Sun, X, Ferguson, J.A, Stanfield, R.L, Wright, P.E. | | Deposit date: | 2024-02-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-08-28 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Mispacking of the F87 sidechain drives aggregation-promoting conformational fluctuations in the subunit interfaces of the transthyretin tetramer.
Protein Sci., 33, 2024
|
|
8W2P
 
 | | BsaXI-DNA complex I | | Descriptor: | CALCIUM ION, DNA (41-MER) Complementary strand of BsaXI DNA substrate, DNA (41-MER) Top strand of BsaXI cognate DNA substrate, ... | | Authors: | Shen, B.W, Stoddard, B.L, Xu, S. | | Deposit date: | 2024-02-20 | | Release date: | 2024-10-30 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | Structural and biochemical analysis of subunit assembly, DNA recognition and cleavage by a Type IIB restriction-modification enzyme: BsaXI
To Be Published
|
|
8YDC
 
 | | Crystal structure of a hammerhead ribozyme with pseudoknot | | Descriptor: | DNA/RNA (5'-R(*AP*CP*AP*UP*GP*UP*CP*U)-D(P*C)-R(P*UP*GP*GP*GP*A)-3'), GUANOSINE-5'-TRIPHOSPHATE, ribozyme strand | | Authors: | Liu, Y, Zhan, X. | | Deposit date: | 2024-02-20 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2.89 Å) | | Cite: | The structure and catalytic mechanism of a pseudoknot-containing hammerhead ribozyme.
Nat Commun, 15, 2024
|
|
8W2O
 
 | | Yeast U1 snRNP with humanized U1C Zinc-Finger domain | | Descriptor: | 56 kDa U1 small nuclear ribonucleoprotein component, ACT1 pre-mRNA, Pre-mRNA-processing factor 39, ... | | Authors: | Shi, S.S, Kuang, Z.L, Zhao, R. | | Deposit date: | 2024-02-20 | | Release date: | 2024-05-15 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (3.49 Å) | | Cite: | Selected humanization of yeast U1 snRNP leads to global suppression of pre-mRNA splicing and mitochondrial dysfunction in the budding yeast.
Rna, 30, 2024
|
|
8S33
 
 | | Malic semialdehyde dehydrogenase (MSA-DH) from Acinetobacter baumannii | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Succinate-semialdehyde dehydrogenase [NAD(P)+] | | Authors: | Piskol, F, Lukat, P, Blankenfeldt, W, Jahn, D, Moser, J. | | Deposit date: | 2024-02-19 | | Release date: | 2024-08-14 | | Last modified: | 2024-09-11 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Biochemical and structural elucidation of the L-carnitine degradation pathway of the human pathogen Acinetobacter baumannii .
Front Microbiol, 15, 2024
|
|
8S37
 | | DNA-bound Type IV-A3 CRISPR effector in complex with DinG helicase from K. pneumoniae (state III) | | Descriptor: | CRISPR type AFERR-associated protein Csf1, CRISPR type AFERR-associated protein Csf2, CRISPR type AFERR-associated protein Csf3, ... | | Authors: | Skorupskaite, A, Ragozius, V, Cepaite, R, Klein, N, Randau, L, Malinauskaite, L, Pausch, P. | | Deposit date: | 2024-02-19 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural variation of types IV-A1- and IV-A3-mediated CRISPR interference.
Nat Commun, 15, 2024
|
|
8YD8
 
 | |
8YD7
 
 | | Structure of FADD/Caspase-8/cFLIP death effector domain assembly | | Descriptor: | CASP8 and FADD-like apoptosis regulator subunit p12, Caspase-8, FAS-associated death domain protein, ... | | Authors: | Lin, S.-C, Yang, C.-Y. | | Deposit date: | 2024-02-19 | | Release date: | 2024-05-15 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Deciphering DED assembly mechanisms in FADD-procaspase-8-cFLIP complexes regulating apoptosis.
Nat Commun, 15, 2024
|
|
8YD3
 
 | | Crystal structure of human Cu-Zn Superoxide Dismutase 1 in complex with 1,2,10-Decanetriol | | Descriptor: | (2S)-decane-1,2,10-triol, GLYCEROL, Superoxide dismutase [Cu-Zn], ... | | Authors: | Aouti, S, Padmanabhan, B. | | Deposit date: | 2024-02-19 | | Release date: | 2024-08-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Crystal structure of human Cu-Zn Superoxide Dismutase 1 in complex with 1,2,10-Decanetriol
To Be Published
|
|
8S35
 | | DNA-bound Type IV-A3 CRISPR effector in complex with DinG helicase from K. pneumoniae (state I) | | Descriptor: | CRISPR type AFERR-associated protein Csf1, CRISPR type AFERR-associated protein Csf2, CRISPR type AFERR-associated protein Csf3, ... | | Authors: | Skorupskaite, A, Ragozius, V, Cepaite, R, Klein, N, Randau, L, Malinauskaite, L, Pausch, P. | | Deposit date: | 2024-02-19 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural variation of types IV-A1- and IV-A3-mediated CRISPR interference.
Nat Commun, 15, 2024
|
|
8S36
 
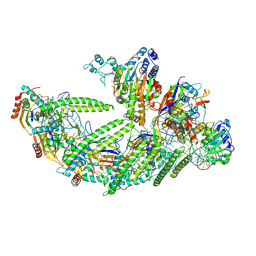 | | DNA-bound Type IV-A3 CRISPR effector in complex with DinG helicase from K. pneumoniae (state II) | | Descriptor: | CRISPR type AFERR-associated protein Csf1, CRISPR type AFERR-associated protein Csf2, CRISPR type AFERR-associated protein Csf3, ... | | Authors: | Skorupskaite, A, Ragozius, V, Cepaite, R, Klein, N, Randau, L, Malinauskaite, L, Pausch, P. | | Deposit date: | 2024-02-19 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural variation of types IV-A1- and IV-A3-mediated CRISPR interference.
Nat Commun, 15, 2024
|
|
8YDB
 
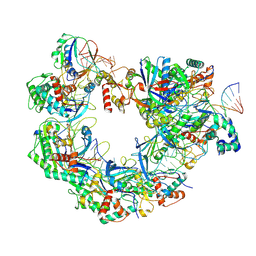 | | Type I-FHNH Cascade-dsDNA intermediate complex | | Descriptor: | 60-nt crRNA, Cas5f, Cas6f, ... | | Authors: | Li, Z. | | Deposit date: | 2024-02-19 | | Release date: | 2024-07-31 | | Last modified: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Mechanisms for HNH-mediated target DNA cleavage in type I CRISPR-Cas systems.
Mol.Cell, 84, 2024
|
|
8S32
 
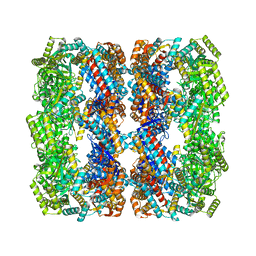 | | GroEL with bound GroTAC peptide | | Descriptor: | Chaperonin GroEL, GroTAC | | Authors: | Wroblewski, K, Izert-Nowakowska, M.A, Goral, T.K, Klimecka, M.M, Kmiecik, S, Gorna, M.W. | | Deposit date: | 2024-02-19 | | Release date: | 2024-02-28 | | Method: | ELECTRON MICROSCOPY (2.45 Å) | | Cite: | Depletion of essential GroEL protein in Escherichia coli using Clp-Interacting Peptidic Protein Erasers (CLIPPERs)
To Be Published
|
|
