2O8S
 
 | | X-ray Crystal Structure of Protein AGR_C_984 from Agrobacterium tumefaciens. Northeast Structural Genomics Consortium Target AtR120. | | Descriptor: | AGR_C_984p, SULFATE ION | | Authors: | Seetharaman, J, Forouhar, F, Su, M, Zhao, L, Cunningham, K, Ma, L.-C, Janjua, H, Xiao, R, Liu, J, Baran, M.C, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-12-12 | | Release date: | 2007-01-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of the Hypothetical Protein AGR_C_984 from Agrobacterium tumefaciens, Northeast Structural Genomics Target AtR120.
To be Published
|
|
2O8T
 
 | | Crystal Structure and Binding Epitopes of Urokinase-type Plasminogen Activator (C122A/N145Q) in complex with Inhibitors | | Descriptor: | DI(HYDROXYETHYL)ETHER, PENTAETHYLENE GLYCOL, SULFATE ION, ... | | Authors: | Zhao, G, Yuan, C, Jiang, L, Huang, Z, Huang, M. | | Deposit date: | 2006-12-12 | | Release date: | 2007-12-25 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Crystal Structure and Binding Epitopes of Urokinase-type Plasminogen Activator (C122A/N145Q/S195A) in complex with Inhibitors
To be Published
|
|
2O8U
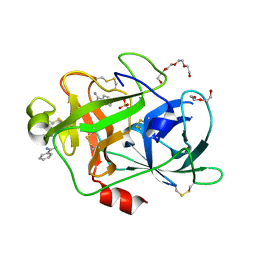 
 | | Crystal Structure and Binding Epitopes of Urokinase-type Plasminogen Activator (C122A/N145Q/S195A) in complex with Inhibitors | | Descriptor: | BENZAMIDINE, DI(HYDROXYETHYL)ETHER, SULFATE ION, ... | | Authors: | Zhao, G, Yuan, C, Jiang, L, Huang, Z, Huang, M. | | Deposit date: | 2006-12-12 | | Release date: | 2007-12-25 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure and Binding Epitopes of Urokinase-type Plasminogen Activator (C122A/N145Q/S195A) in complex with Inhibitors
To be Published
|
|
2O8V
 
 | | PAPS reductase in a covalent complex with thioredoxin C35A | | Descriptor: | Phosphoadenosine phosphosulfate reductase, Thioredoxin 1 | | Authors: | Chartron, J, Shiau, C, Stout, C.D, Carroll, K.S. | | Deposit date: | 2006-12-12 | | Release date: | 2007-03-27 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | 3'-Phosphoadenosine-5'-phosphosulfate Reductase in Complex with Thioredoxin: A Structural Snapshot in the Catalytic Cycle.
Biochemistry, 46, 2007
|
|
2O8W
 
 | | Crystal Structure and Binding Epitopes of Urokinase-type Plasminogen Activator (C122A/N145Q/S195A) in complex with Inhibitors | | Descriptor: | 1-phenylguanidine, SULFATE ION, TETRAETHYLENE GLYCOL, ... | | Authors: | Zhao, G, Yuan, C, Jiang, L, Huang, Z, Huang, M. | | Deposit date: | 2006-12-12 | | Release date: | 2007-12-25 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Crystal Structure and Binding Epitopes of Urokinase-type Plasminogen Activator (C122A/N145Q/S195A) in complex with Inhibitors
To be Published
|
|
2O8X
 
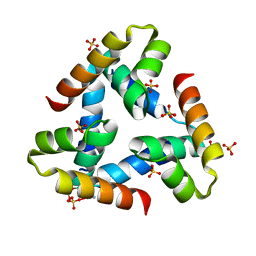 | |
2O8Y
 
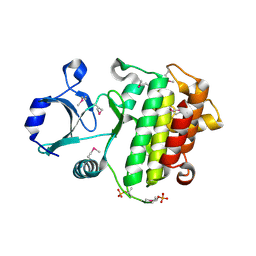 | | Apo IRAK4 Kinase Domain | | Descriptor: | Interleukin-1 receptor-associated kinase 4 | | Authors: | Boriack-Sjodin, P.A, Mol, C. | | Deposit date: | 2006-12-12 | | Release date: | 2007-12-25 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structures of the apo and inhibited IRAK4 kinase domain
To be Published
|
|
2O8Z
 
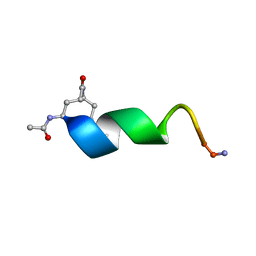 | | Bound Structure of CRF1 Extracellular Domain Antagonist | | Descriptor: | cCRF(30-41) Peptide | | Authors: | Mesleh, M.F, Shirley, W.A, Heise, C.E, Ling, N, Maki, R.A, Laura, R.P. | | Deposit date: | 2006-12-12 | | Release date: | 2006-12-26 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR structural characterization of a minimal peptide antagonist bound to the extracellular domain of the corticotropin-releasing factor1 receptor.
J.Biol.Chem., 282, 2007
|
|
2O90
 
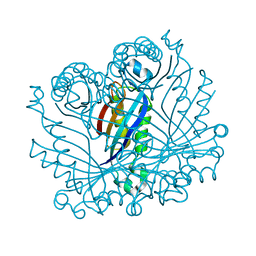 | |
2O92
 
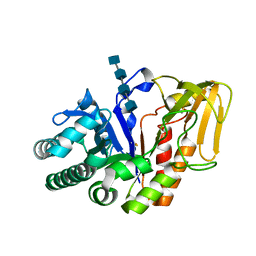 | | Crystal structure of a signalling protein (SPG-40) complex with tetrasaccharide at 3.0A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Chitinase-3-like protein 1, alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Sharma, P, Singh, N, Sharma, S, Kaur, P, Singh, T.P. | | Deposit date: | 2006-12-13 | | Release date: | 2006-12-26 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of a signalling protein (SPG-40) complex with tetrasaccharide at 3.0A resolution
To be Published
|
|
2O93
 
 | |
2O94
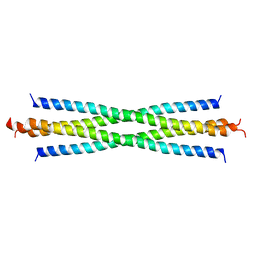 
 | |
2O95
 
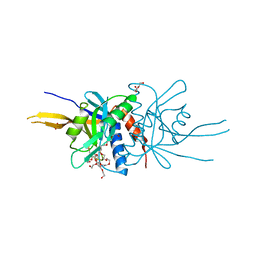 | | Crystal Structure of the Metal-Free Dimeric Human Mov34 MPN domain (residues 1-186) | | Descriptor: | 2-{2-[2-2-(METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, 26S proteasome non-ATPase regulatory subunit 7, DODECAETHYLENE GLYCOL, ... | | Authors: | Sanches, M, Alves, B.S.C, Zanchin, N.I.T, Guimaraes, B.G. | | Deposit date: | 2006-12-13 | | Release date: | 2007-07-17 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Crystal Structure of the Human Mov34 MPN Domain Reveals a Metal-free Dimer
J.Mol.Biol., 370, 2007
|
|
2O96
 
 | |
2O97
 
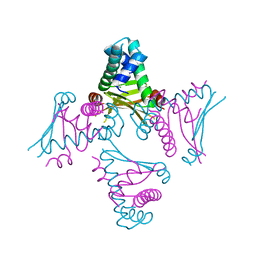 | | Crystal Structure of E. coli HU heterodimer | | Descriptor: | CHLORIDE ION, DNA-binding protein HU-alpha, DNA-binding protein HU-beta, ... | | Authors: | Guo, F, Adhya, S. | | Deposit date: | 2006-12-13 | | Release date: | 2007-03-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Spiral structure of Escherichia coli HU{alpha}beta provides foundation for DNA supercoiling.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2O98
 
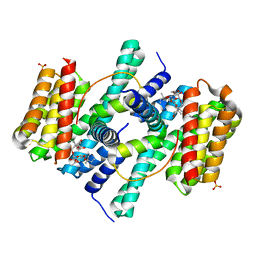 | | Structure of the 14-3-3 / H+-ATPase plant complex | | Descriptor: | 14-3-3-like protein C, FUSICOCCIN, Plasma membrane H+ ATPase, ... | | Authors: | Ottmann, C, Weyand, M, Wittinghofer, A, Oecking, C. | | Deposit date: | 2006-12-13 | | Release date: | 2007-04-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of a 14-3-3 coordinated hexamer of the plant plasma membrane H+ -ATPase by combining X-ray crystallography and electron cryomicroscopy
Mol.Cell, 25, 2007
|
|
2O99
 
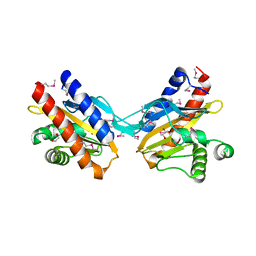 | | The crystal structure of E.coli IclR C-terminal fragment in complex with glyoxylate | | Descriptor: | 1,2-ETHANEDIOL, Acetate operon repressor, GLYCOLIC ACID | | Authors: | Lunin, V.V, Ezersky, A, Evdokimova, E, Kudritska, M, Savchenko, A. | | Deposit date: | 2006-12-13 | | Release date: | 2007-04-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Glyoxylate and Pyruvate Are Antagonistic Effectors of the Escherichia coli IclR Transcriptional Regulator.
J.Biol.Chem., 282, 2007
|
|
2O9A
 
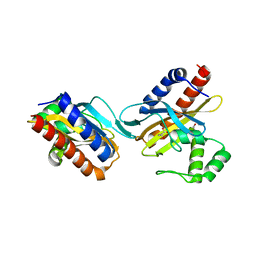 | | The crystal structure of the E.coli IclR C-terminal fragment in complex with pyruvate. | | Descriptor: | 1,2-ETHANEDIOL, Acetate operon repressor, PYRUVIC ACID | | Authors: | Lunin, V.V, Ezersky, A, Evdokimova, E, Kudritska, M, Savchenko, A. | | Deposit date: | 2006-12-13 | | Release date: | 2007-04-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Glyoxylate and Pyruvate Are Antagonistic Effectors of the Escherichia coli IclR Transcriptional Regulator.
J.Biol.Chem., 282, 2007
|
|
2O9B
 
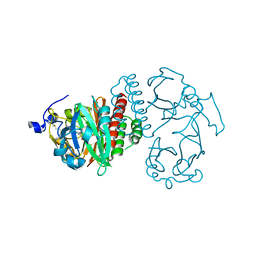 | | Crystal Structure of Bacteriophytochrome chromophore binding domain | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome | | Authors: | Wagner, J.R, Brunzelle, J.S, Vierstra, R.D, Forest, K.T. | | Deposit date: | 2006-12-13 | | Release date: | 2007-03-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | High resolution structure of deinococcus bacteriophytochrome yields new insights into phytochrome architecture and evolution.
J.Biol.Chem., 282, 2007
|
|
2O9C
 
 | | Crystal Structure of Bacteriophytochrome chromophore binding domain at 1.45 angstrom resolution | | Descriptor: | 3-[2-[(Z)-[3-(2-carboxyethyl)-5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-4-methyl-pyrrol-1-ium -2-ylidene]methyl]-5-[(Z)-[(3E)-3-ethylidene-4-methyl-5-oxidanylidene-pyrrolidin-2-ylidene]methyl]-4-methyl-1H-pyrrol-3- yl]propanoic acid, Bacteriophytochrome | | Authors: | Wagner, J.R, Brunzelle, J.S, Vierstra, R.D, Forest, K.T. | | Deposit date: | 2006-12-13 | | Release date: | 2007-03-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | High resolution structure of deinococcus bacteriophytochrome yields new insights into phytochrome architecture and evolution.
J.Biol.Chem., 282, 2007
|
|
2O9D
 
 | |
2O9E
 
 | |
2O9F
 
 | |
2O9G
 
 | |
2O9I
 
 | | Crystal Structure of the Human Pregnane X Receptor LBD in complex with an SRC-1 coactivator peptide and T0901317 | | Descriptor: | N-(2,2,2-TRIFLUOROETHYL)-N-{4-[2,2,2-TRIFLUORO-1-HYDROXY-1-(TRIFLUOROMETHYL)ETHYL]PHENYL}BENZENESULFONAMIDE, Nuclear Receptor Coactivator 1 isoform 3, Orphan nuclear receptor PXR | | Authors: | Xue, Y, Redinbo, M.R. | | Deposit date: | 2006-12-13 | | Release date: | 2007-01-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the PXR-T1317 complex provides a scaffold to examine the potential for receptor antagonism.
Bioorg.Med.Chem., 15, 2007
|
|
