2LX1
 
 | |
7EDU
 
 | | RNA duplex containing C-Au-C base pair | | Descriptor: | GOLD ION, RNA (5'-R(*GP*GP*AP*CP*UP*CP*GP*AP*CP*UP*CP*C)-3') | | Authors: | Kondo, J, Iwase, E. | | Deposit date: | 2021-03-17 | | Release date: | 2022-03-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | RNA duplex containing C-Au-C base pair
To Be Published
|
|
7ECK
 
 | | RNA duplex containing C-Ag-A and U-Ag-A base pairs | | Descriptor: | DNA/RNA (5'-R(*GP*GP*AP*CP*U)-D(P*(CBR))-R(P*GP*AP*AP*UP*CP*C)-3'), SILVER ION | | Authors: | Kondo, J, Uchida, Y. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | RNA duplex containing C-Ag-A and U-Ag-A base pairs
To Be Published
|
|
7ECJ
 
 | | RNA duplex containing C-A base pairs | | Descriptor: | DNA/RNA (5'-R(*GP*GP*AP*CP*U)-D(P*(CBR))-R(P*GP*AP*AP*UP*CP*C)-3') | | Authors: | Kondo, J, Uchida, Y. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | RNA duplex containing C-A base pairs
To Be Published
|
|
7ECM
 
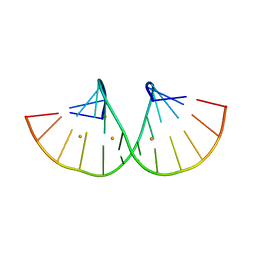 | | RNA duplex containing C-A base pair | | Descriptor: | BARIUM ION, RNA (5'-R(*GP*GP*GP*CP*CP*CP*GP*GP*AP*CP*CP*C)-3') | | Authors: | Kondo, J, Tsudura, A. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | RNA duplex containing C-A base pair
To Be Published
|
|
7ECL
 
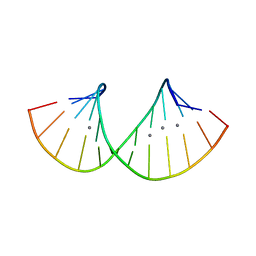 | | RNA duplex containing C-Ag-A and U-Ag-A base pair | | Descriptor: | DNA/RNA (5'-R(*GP*GP*AP*CP*U)-D(P*(CBR))-R(P*GP*AP*AP*UP*CP*C)-3'), SILVER ION | | Authors: | Kondo, J, Uchida, Y. | | Deposit date: | 2021-03-12 | | Release date: | 2022-03-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | RNA duplex containing C-Ag-A and U-Ag-A base pair
To Be Published
|
|
7EEM
 
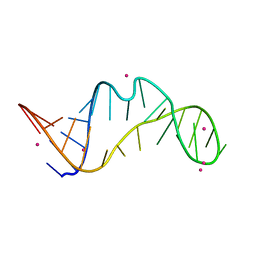 | | RNA bulged-G motif | | Descriptor: | IRIDIUM ION, RNA (27-MER) | | Authors: | Kondo, J, Sekiguchi, S. | | Deposit date: | 2021-03-19 | | Release date: | 2022-03-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.593 Å) | | Cite: | RNA bulged-G motif
To Be Published
|
|
7EI7
 
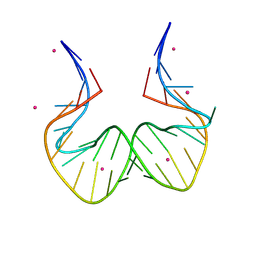 | | RNA kink-turn motif with 2-aminopurine | | Descriptor: | COBALT (II) ION, RNA (5'-R(*GP*GP*CP*GP*A)-D(P*(2PR))-R(P*GP*AP*AP*CP*CP*GP*GP*GP*GP*AP*GP*CP*C)-3') | | Authors: | Kondo, J, Saisu, S. | | Deposit date: | 2021-03-30 | | Release date: | 2022-03-30 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | RNA kink-turn motif with 2-aminopurine
To Be Published
|
|
7EI8
 
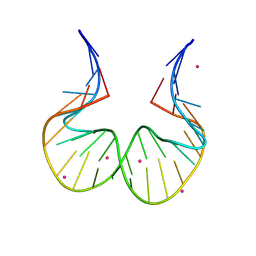 | | RNA kink-turn motif with 2-aminopurine | | Descriptor: | COBALT (II) ION, RNA (5'-R(*GP*GP*C)-D(P*(2PR)P*(2PR)P*(2PR)P*(2PR))-R(P*AP*AP*CP*CP*GP*GP*GP*GP*AP*GP*CP*C)-3') | | Authors: | Kondo, J, Saisu, S. | | Deposit date: | 2021-03-30 | | Release date: | 2022-03-30 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | RNA kink-turn motif with 2-aminopurine
To Be Published
|
|
7EIA
 
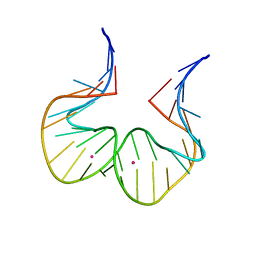 | | RNA kink-turn motif with pyrrolo cytosine | | Descriptor: | POTASSIUM ION, RNA (5'-R(*GP*GP*CP*GP*A)-D(P*(4PC))-R(P*GP*AP*AP*CP*CP*GP*GP*GP*GP*AP*GP*CP*C)-3') | | Authors: | Kondo, J, Miyauchi, T. | | Deposit date: | 2021-03-30 | | Release date: | 2022-03-30 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | RNA kink-turn motif with pyrrolo cytosine
To Be Published
|
|
2O9T
 
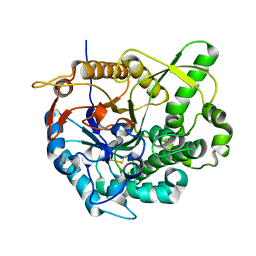 | |
2O9R
 
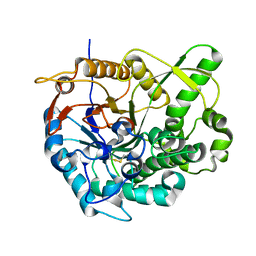 | | beta-glucosidase B complexed with thiocellobiose | | Descriptor: | Beta-glucosidase B, beta-D-glucopyranose-(1-4)-4-thio-beta-D-glucopyranose | | Authors: | Isorna, P, Polaina, J, Sanz-Aparicio, J. | | Deposit date: | 2006-12-14 | | Release date: | 2007-10-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structures of Paenibacillus polymyxa beta-Glucosidase B Complexes Reveal the Molecular Basis of Substrate Specificity and Give New Insights into the Catalytic Machinery of Family I Glycosidases
J.Mol.Biol., 371, 2007
|
|
7F8Z
 
 | | RNA kink-turn motif with 2-aminopurine | | Descriptor: | POTASSIUM ION, RNA (5'-R(*GP*GP*CP*GP*A)-D(P*(2PR))-R(P*GP*AP*AP*CP*CP*GP*GP*GP*GP*AP*GP*CP*C)-3') | | Authors: | Kondo, J, Miyauchi, T. | | Deposit date: | 2021-07-03 | | Release date: | 2022-07-06 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | RNA kink-turn motif with 2-aminopurine
To Be Published
|
|
1KAJ
 
 | |
1WOY
 
 | |
2RN1
 
 | | Liquid crystal solution structure of the kissing complex formed by the apical loop of the HIV TAR RNA and a high affinity RNA aptamer optimized by SELEX | | Descriptor: | RNA (5'-R(P*GP*AP*GP*CP*CP*CP*UP*GP*GP*GP*AP*GP*GP*CP*UP*C)-3'), RNA (5'-R(P*GP*CP*UP*GP*GP*UP*CP*CP*CP*AP*GP*AP*CP*AP*GP*C)-3') | | Authors: | Van Melckebeke, H, Devany, M, Di Primo, C, Beaurain, F, Toulme, J, Bryce, D.L, Boisbouvier, J. | | Deposit date: | 2007-12-05 | | Release date: | 2008-09-23 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Liquid-crystal NMR structure of HIV TAR RNA bound to its SELEX RNA aptamer reveals the origins of the high stability of the complex
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
4WKG
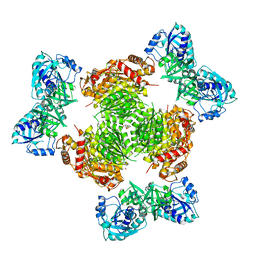 
 | |
2NQZ
 
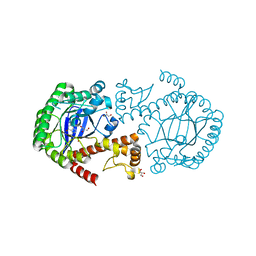 | | Trna-guanine transglycosylase (TGT) mutant in complex with 7-deaza-7-aminomethyl-guanine | | Descriptor: | 7-DEAZA-7-AMINOMETHYL-GUANINE, GLYCEROL, Queuine tRNA-ribosyltransferase, ... | | Authors: | Tidten, N, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2006-11-01 | | Release date: | 2007-11-13 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | Investigation of Specificity Determinants in Bacterial tRNA-Guanine Transglycosylase Reveals Queuine, the Substrate of Its Eucaryotic Counterpart, as Inhibitor
Plos One, 8, 2013
|
|
2NSO
 
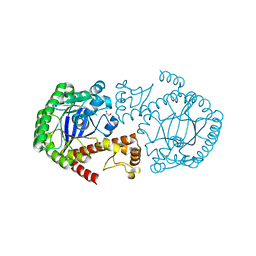 | | Trna-gunanine-transglycosylase (TGT) mutant Y106F, C158V, A232S, V233G- APO-Structure | | Descriptor: | GLYCEROL, Queuine tRNA-ribosyltransferase, ZINC ION | | Authors: | Tidten, N, Heine, A, Reuter, K, Klebe, G. | | Deposit date: | 2006-11-06 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Investigation of Specificity Determinants in Bacterial tRNA-Guanine Transglycosylase Reveals Queuine, the Substrate of Its Eucaryotic Counterpart, as Inhibitor.
Plos One, 8, 2013
|
|
1RNG
 
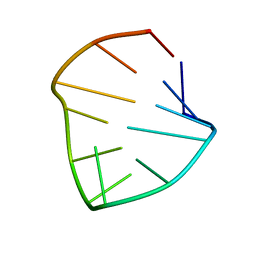 | |
1RHT
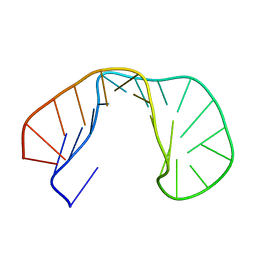 
 | | 24-MER RNA HAIRPIN COAT PROTEIN BINDING SITE FOR BACTERIOPHAGE R17 (NMR, MINIMIZED AVERAGE STRUCTURE) | | Descriptor: | RNA (5'-R(P*GP*GP*GP*AP*CP*UP*GP*AP*CP*GP*AP*UP*CP*AP*CP*GP*CP*AP*GP*UP*CP*UP*AP*U)-3') | | Authors: | Borer, P.N, Lin, Y, Wang, S, Roggenbuck, M.W, Gott, J.M, Uhlenbeck, O.C, Pelczer, I. | | Deposit date: | 1995-03-03 | | Release date: | 1995-06-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Proton NMR and structural features of a 24-nucleotide RNA hairpin.
Biochemistry, 34, 1995
|
|
2M8K
 
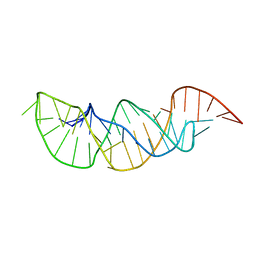 | | A pyrimidine motif triple helix in the Kluyveromyces lactis telomerase RNA pseudoknot is essential for function in vivo | | Descriptor: | RNA (48-MER) | | Authors: | Cash, D.D, Cohen, O, Kim, N, Shefer, K, Brown, Y, Ulyanov, N.B, Tzfati, Y, Feigon, J. | | Deposit date: | 2013-05-22 | | Release date: | 2013-06-19 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Pyrimidine motif triple helix in the Kluyveromyces lactis telomerase RNA pseudoknot is essential for function in vivo.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4YU7
 
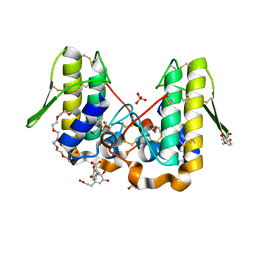 | | Crystal structure of Piratoxin I (PrTX-I) complexed to caffeic acid | | Descriptor: | 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, Basic phospholipase A2 homolog piratoxin-1, CAFFEIC ACID, ... | | Authors: | Fernandes, C.A.H, Fontes, M.R.M. | | Deposit date: | 2015-03-18 | | Release date: | 2015-08-05 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.647 Å) | | Cite: | Structural Basis for the Inhibition of a Phospholipase A2-Like Toxin by Caffeic and Aristolochic Acids.
Plos One, 10, 2015
|
|
1RAU
 
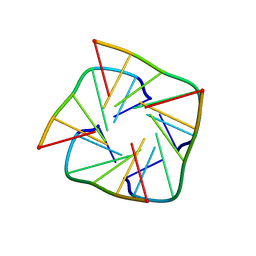 | |
2OOM
 
 | | NMR structure of a kissing complex formed between the TAR RNA element of HIV-1 and a LNA/RNA aptamer | | Descriptor: | RNA 16-mer, TAR RNA element of HIV-1 | | Authors: | Lebars, I, Richard, T, Di Primo, C, Toulme, J.J. | | Deposit date: | 2007-01-26 | | Release date: | 2008-01-29 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR structure of a kissing complex formed between the TAR RNA element of HIV-1 and a LNA-modified aptamer
Nucleic Acids Res., 35, 2007
|
|
