1D6H
 
 | | CHALONE SYNTHASE (N336A MUTANT COMPLEXED WITH COA) | | Descriptor: | CHALCONE SYNTHASE, COENZYME A, SULFATE ION | | Authors: | Jez, J.M, Ferrer, J.L, Bowman, M.E, Dixon, R.A, Noel, J.P. | | Deposit date: | 1999-10-13 | | Release date: | 2000-02-03 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Dissection of malonyl-coenzyme A decarboxylation from polyketide formation in the reaction mechanism of a plant polyketide synthase.
Biochemistry, 39, 2000
|
|
1D6I
 
 | | CHALCONE SYNTHASE (H303Q MUTANT) | | Descriptor: | CHALCONE SYNTHASE, SULFATE ION | | Authors: | Jez, J.M, Ferrer, J.L, Bowman, M.E, Dixon, R.A, Noel, J.P. | | Deposit date: | 1999-10-13 | | Release date: | 2000-02-03 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Dissection of malonyl-coenzyme A decarboxylation from polyketide formation in the reaction mechanism of a plant polyketide synthase.
Biochemistry, 39, 2000
|
|
1D6J
 
 | |
1D6K
 
 | | NMR SOLUTION STRUCTURE OF THE 5S RRNA E-LOOP/L25 COMPLEX | | Descriptor: | 5S RRNA E-LOOP (5SE), RIBOSOMAL PROTEIN L25 | | Authors: | Stoldt, M, Wohnert, J, Ohlenschlager, O, Gorlach, M, Brown, L.R. | | Deposit date: | 1999-10-14 | | Release date: | 1999-11-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The NMR structure of the 5S rRNA E-domain-protein L25 complex shows preformed and induced recognition.
EMBO J., 18, 1999
|
|
1D6M
 
 | |
1D6N
 
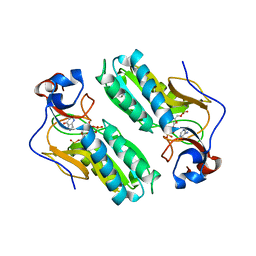 | | TERNARY COMPLEX STRUCTURE OF HUMAN HGPRTASE, PRPP, MG2+, AND THE INHIBITOR HPP REVEALS THE INVOLVEMENT OF THE FLEXIBLE LOOP IN SUBSTRATE BINDING | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, 3H-PYRAZOLO[4,3-D]PYRIMIDIN-7-OL, MAGNESIUM ION, ... | | Authors: | Balendiran, G.K. | | Deposit date: | 1999-10-14 | | Release date: | 1999-12-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Ternary complex structure of human HGPRTase, PRPP, Mg2+, and the inhibitor HPP reveals the involvement of the flexible loop in substrate binding.
Protein Sci., 8, 1999
|
|
1D6O
 
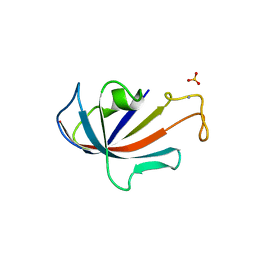 | | NATIVE FKBP | | Descriptor: | AMMONIUM ION, PROTEIN (FK506-BINDING PROTEIN), SULFATE ION | | Authors: | Burkhard, P, Taylor, P, Walkinshaw, M.D. | | Deposit date: | 1999-10-15 | | Release date: | 1999-10-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | X-ray structures of small ligand-FKBP complexes provide an estimate for hydrophobic interaction energies.
J.Mol.Biol., 295, 2000
|
|
1D6P
 
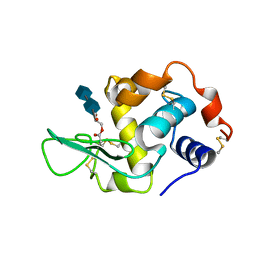 | | HUMAN LYSOZYME L63 MUTANT LABELLED WITH 2',3'-EPOXYPROPYL N,N'-DIACETYLCHITOBIOSE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, LYSOZYME | | Authors: | Muraki, M, Harata, K, Sugita, N, Sato, K. | | Deposit date: | 1999-10-15 | | Release date: | 2000-01-21 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Protein-carbohydrate interactions in human lysozyme probed by combining site-directed mutagenesis and affinity labeling.
Biochemistry, 39, 2000
|
|
1D6Q
 
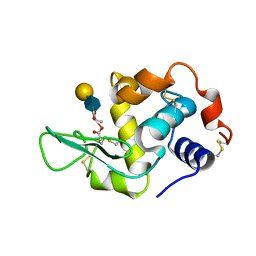 | | HUMAN LYSOZYME E102 MUTANT LABELLED WITH 2',3'-EPOXYPROPYL GLYCOSIDE OF N-ACETYLLACTOSAMINE | | Descriptor: | GLYCEROL, LYSOZYME, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Muraki, M, Harata, K, Sugita, N, Sato, K. | | Deposit date: | 1999-10-15 | | Release date: | 2000-01-21 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Protein-carbohydrate interactions in human lysozyme probed by combining site-directed mutagenesis and affinity labeling.
Biochemistry, 39, 2000
|
|
1D6R
 
 | | CRYSTAL STRUCTURE OF CANCER CHEMOPREVENTIVE BOWMAN-BIRK INHIBITOR IN TERNARY COMPLEX WITH BOVINE TRYPSIN AT 2.3 A RESOLUTION. STRUCTURAL BASIS OF JANUS-FACED SERINE PROTEASE INHIBITOR SPECIFICITY | | Descriptor: | BOWMAN-BIRK PROTEINASE INHIBITOR PRECURSOR, TRYPSINOGEN | | Authors: | Koepke, J, Ermler, U, Wenzl, G, Flecker, P. | | Deposit date: | 1999-10-15 | | Release date: | 2000-05-05 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of cancer chemopreventive Bowman-Birk inhibitor in ternary complex with bovine trypsin at 2.3 A resolution. Structural basis of Janus-faced serine protease inhibitor specificity.
J.Mol.Biol., 298, 2000
|
|
1D6S
 
 | | CRYSTAL STRUCTURE OF THE K41A MUTANT OF O-ACETYLSERINE SULFHYDRYLASE COMPLEXED IN EXTERNAL ALDIMINE LINKAGE WITH METHIONINE | | Descriptor: | METHIONINE, O-ACETYLSERINE SULFHYDRYLASE, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Burkhard, P, Tai, C.H, Ristroph, C.M, Cook, P.F, Jansonius, J.N. | | Deposit date: | 1999-10-15 | | Release date: | 2000-04-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Ligand binding induces a large conformational change in O-acetylserine sulfhydrylase from Salmonella typhimurium.
J.Mol.Biol., 291, 1999
|
|
1D6T
 
 | | RNASE P PROTEIN FROM STAPHYLOCOCCUS AUREUS | | Descriptor: | RIBONUCLEASE P | | Authors: | Spitzfaden, C. | | Deposit date: | 1999-10-15 | | Release date: | 2000-10-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of ribonuclease P protein from Staphylococcus aureus reveals a unique binding site for single-stranded RNA.
J.Mol.Biol., 295, 2000
|
|
1D6U
 
 | | CRYSTAL STRUCTURE OF E. COLI AMINE OXIDASE ANAEROBICALLY REDUCED WITH BETA-PHENYLETHYLAMINE | | Descriptor: | 2-PHENYLETHYLAMINE, CALCIUM ION, COPPER (II) ION, ... | | Authors: | Wilmot, C.M, Hajdu, J, McPherson, M.J, Knowles, P.F, Phillips, S.E.V. | | Deposit date: | 1999-10-15 | | Release date: | 2000-02-02 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Visualization of dioxygen bound to copper during enzyme catalysis.
Science, 286, 1999
|
|
1D6V
 
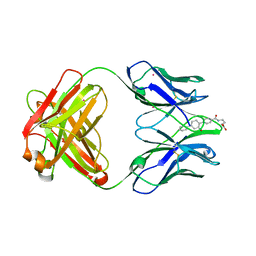 | | CONFORMATION EFFECTS IN BIOLOGICAL CATALYSIS INTRODUCED BY OXY-COPE ANTIBODY MATURATION | | Descriptor: | (1S,2S,5S)2-(4-GLUTARIDYLBENZYL)-5-PHENYL-1-CYCLOHEXANOL, CADMIUM ION, CHIMERIC GERMLINE PRECURSOR OF OXY-COPE CATALYTIC ANTIBODY AZ-28 (HEAVY CHAIN), ... | | Authors: | Mundorff, E.C, Hanson, M.A, Schultz, P.G, Stevens, R.C. | | Deposit date: | 1999-10-15 | | Release date: | 2000-02-09 | | Last modified: | 2017-06-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational effects in biological catalysis: an antibody-catalyzed oxy-cope rearrangement.
Biochemistry, 39, 2000
|
|
1D6W
 
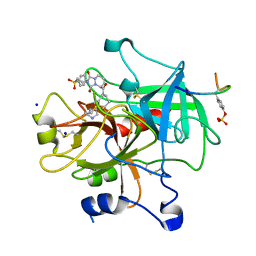 | | STRUCTURE OF THROMBIN COMPLEXED WITH SELECTIVE NON-ELECTROPHILIC INHIBITORS HAVING CYCLOHEXYL MOIETIES AT P1 | | Descriptor: | (5S)-N-[trans-4-(2-amino-1H-imidazol-5-yl)cyclohexyl]-1,3-dioxo-2-[2-(phenylsulfonyl)ethyl]-2,3,5,8-tetrahydro-1H-[1,2,4]triazolo[1,2-a]pyridazine-5-carboxamide, DECAPEPTIDE INHIBITOR, SODIUM ION, ... | | Authors: | Mochalkin, I, Tulinsky, A. | | Deposit date: | 1999-10-15 | | Release date: | 2000-10-25 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of thrombin complexed with selective non-electrophilic inhibitors having cyclohexyl moieties at P1.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1D6X
 
 | |
1D6Y
 
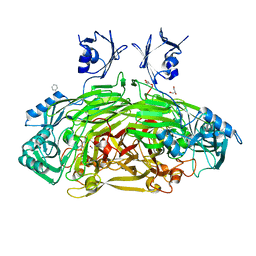 | | CRYSTAL STRUCTURE OF E. COLI COPPER-CONTAINING AMINE OXIDASE ANAEROBICALLY REDUCED WITH BETA-PHENYLETHYLAMINE AND COMPLEXED WITH NITRIC OXIDE. | | Descriptor: | 2-PHENYLETHYLAMINE, CALCIUM ION, COPPER (II) ION, ... | | Authors: | Wilmot, C.M, Hajdu, J, McPherson, M.J, Knowles, P.F, Phillips, S.E.V. | | Deposit date: | 1999-10-16 | | Release date: | 2000-02-02 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Visualization of dioxygen bound to copper during enzyme catalysis.
Science, 286, 1999
|
|
1D6Z
 
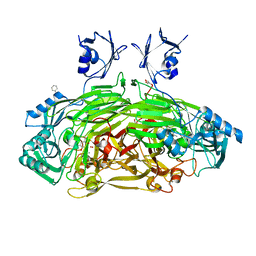 | | CRYSTAL STRUCTURE OF THE AEROBICALLY FREEZE TRAPPED RATE-DETERMINING CATALYTIC INTERMEDIATE OF E. COLI COPPER-CONTAINING AMINE OXIDASE. | | Descriptor: | 2-PHENYLETHYLAMINE, CALCIUM ION, COPPER (II) ION, ... | | Authors: | Wilmot, C.M, Hajdu, J, McPherson, M.J, Knowles, P.F, Phillips, S.E.V. | | Deposit date: | 1999-10-16 | | Release date: | 2000-02-02 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Visualization of dioxygen bound to copper during enzyme catalysis.
Science, 286, 1999
|
|
1D70
 
 | |
1D75
 
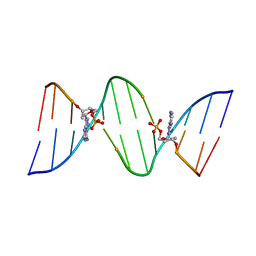 | | CONFORMATION OF THE GUANINE.8-OXOADENINE BASE PAIRS IN THE CRYSTAL STRUCTURE OF D(CGCGAATT(O8A)GCG) | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*(A38)P*GP*CP*G)-3') | | Authors: | Leonard, G.A, Guy, A, Brown, T, Teoule, R, Hunter, W.N. | | Deposit date: | 1992-05-07 | | Release date: | 1993-01-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Conformation of guanine-8-oxoadenine base pairs in the crystal structure of d(CGCGAATT(O8A)GCG).
Biochemistry, 31, 1992
|
|
1D76
 
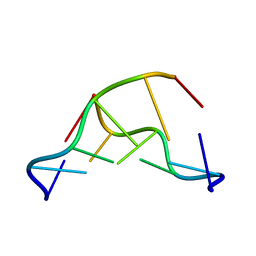 | | CRYSTAL AND MOLECULAR STRUCTURE OF A DNA FRAGMENT CONTAINING A 2-AMINO ADENINE MODIFICATION: THE RELATIONSHIP BETWEEN CONFORMATION, PACKING, AND HYDRATION IN Z-DNA HEXAMERS | | Descriptor: | DNA (5'-D(*CP*GP*UP*(1AP)P*CP*G)-3') | | Authors: | Schneider, B, Ginell, S.L, Jones, R, Gaffney, B, Berman, H.M. | | Deposit date: | 1992-05-19 | | Release date: | 1992-09-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal and molecular structure of a DNA fragment containing a 2-aminoadenine modification: the relationship between conformation, packing, and hydration in Z-DNA hexamers.
Biochemistry, 31, 1992
|
|
1D77
 
 | |
1D78
 
 | | HIGH RESOLUTION REFINEMENT OF THE HEXAGONAL A-DNA OCTAMER D(GTGTACAC) AT 1.4 ANGSTROMS RESOLUTION | | Descriptor: | DNA (5'-D(*GP*TP*GP*TP*AP*CP*AP*C)-3') | | Authors: | Thota, N, Li, X.H, Bingman, C.A, Sundaralingam, M. | | Deposit date: | 1992-06-12 | | Release date: | 1993-04-15 | | Last modified: | 2023-03-22 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | High-resolution refinement of the hexagonal A-DNA octamer d(GTGTACAC) at 1.4 A.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
1D79
 
 | | HIGH RESOLUTION REFINEMENT OF THE HEXAGONAL A-DNA OCTAMER D(GTGTACAC) AT 1.4 ANGSTROMS RESOLUTION | | Descriptor: | DNA (5'-D(*GP*TP*GP*TP*AP*CP*AP*C)-3') | | Authors: | Thota, N, Li, X.H, Bingman, C.A, Sundaralingam, M. | | Deposit date: | 1992-06-12 | | Release date: | 1993-04-15 | | Last modified: | 2023-03-22 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | High-resolution refinement of the hexagonal A-DNA octamer d(GTGTACAC) at 1.4 A.
Acta Crystallogr.,Sect.D, 49, 1993
|
|
1D7A
 | | CRYSTAL STRUCTURE OF E. COLI PURE-MONONUCLEOTIDE COMPLEX. | | Descriptor: | 5-AMINOIMIDAZOLE RIBONUCLEOTIDE, PHOSPHORIBOSYLAMINOIMIDAZOLE CARBOXYLASE | | Authors: | Mathews, I.I, Kappock, T.J, Stubbe, J, Ealick, S.E. | | Deposit date: | 1999-10-16 | | Release date: | 1999-12-03 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of Escherichia coli PurE, an unusual mutase in the purine biosynthetic pathway.
Structure Fold.Des., 7, 1999
|
|
