2N4S
 
 | |
2N4W
 | |
2N2Z
 
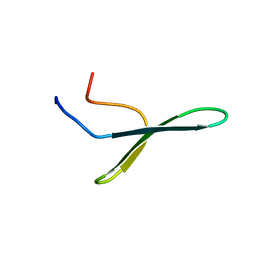
 | | NMR spatial structure of nonspecific lipid transfer protein from the dill Anethum graveolens L. | | Descriptor: | Non-specific lipid-transfer protein | | Authors: | Mineev, K.S, Melnikova, D.N, Finkina, E.I, Arseniev, A.S, Ovchinnikova, T.V. | | Deposit date: | 2015-05-19 | | Release date: | 2016-03-30 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | A novel lipid transfer protein from the dill Anethum graveolens L.: isolation, structure, heterologous expression, and functional characteristics.
J.Pept.Sci., 22, 2016
|
|
2LFW
 
 | | NMR structure of the PhyRSL-NepR complex from Sphingomonas sp. Fr1 | | Descriptor: | NepR anti sigma factor, PhyR sigma-like domain | | Authors: | Campagne, S, Damberger, F.F, Vorholt, J.A, Allain, F.H.-T. | | Deposit date: | 2011-07-18 | | Release date: | 2012-04-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis for sigma factor mimicry in the general stress response of Alphaproteobacteria.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1HZN
 
 | |
2M1F
 
 | | NMR Structure of Antiamoebin I (peptaibol antibiotic) bound to DMPC/DHPC bicelles | | Descriptor: | Antiamoebin I | | Authors: | Shenkarev, Z.O, Paramonov, A.S, Gizatullina, A.K. | | Deposit date: | 2012-11-27 | | Release date: | 2012-12-12 | | Last modified: | 2013-10-02 | | Method: | SOLUTION NMR | | Cite: | Peptaibol antiamoebin I: spatial structure, backbone dynamics, interaction with bicelles and lipid-protein nanodiscs, and pore formation in context of barrel-stave model.
Chem.Biodivers., 10, 2013
|
|
2MFZ
 
 | | NMR structure of C-terminal domain from A. ventricosus minor ampullate spidroin (MiSp) | | Descriptor: | Minor ampullate spidroin | | Authors: | Otikovs, M, Jaudzems, K, Andersson, M, Chen, G, Landreh, M, Nordling, K, Kronqvist, N, Westermark, P, Jornvall, H, Knight, S, Ridderstrale, Y, Holm, L, Meng, Q, Chesler, M, Johansson, J, Rising, A. | | Deposit date: | 2013-10-24 | | Release date: | 2014-08-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Carbonic Anhydrase Generates CO2 and H+ That Drive Spider Silk Formation Via Opposite Effects on the Terminal Domains
Plos Biol., 12, 2014
|
|
1IVT
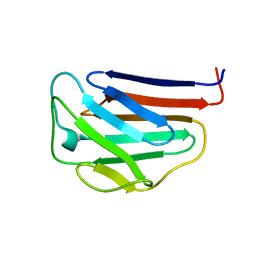 
 | | NMR structures of the C-terminal globular domain of human lamin A/C | | Descriptor: | Lamin A/C | | Authors: | Krimm, I, Ostlund, C, Gilquin, B, Couprie, J, Hossenlopp, P, Mornon, J.P, Bonn, G, Courvalin, J.C, Worman, H.J, Zinn-Justin, S. | | Deposit date: | 2002-03-29 | | Release date: | 2002-08-21 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The Ig-like structure of the C-terminal domain of lamin A/C, mutated in muscular dystrophies, cardiomyopathy, and partial lipodystrophy.
Structure, 10, 2002
|
|
1ILO
 
 | | NMR structure of a thioredoxin, MtH895, from the archeon Methanobacterium thermoautotrophicum strain delta H. | | Descriptor: | conserved hypothetical protein MtH895 | | Authors: | Bhattacharyya, S, Habibi-Nazhad, B, Slupsky, C.M, Sykes, B.D, Wishart, D.S, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2001-05-08 | | Release date: | 2001-11-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Identification of a novel archaebacterial thioredoxin: determination of function through structure.
Biochemistry, 41, 2002
|
|
2JXJ
 
 | |
2MBY
 
 | | NMR Structure of Rrp7 C-terminal Domain | | Descriptor: | Ribosomal RNA-processing protein 7 | | Authors: | Lin, J, Feng, Y, Ye, K. | | Deposit date: | 2013-08-08 | | Release date: | 2013-09-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | An RNA-Binding Complex Involved in Ribosome Biogenesis Contains a Protein with Homology to tRNA CCA-Adding Enzyme.
Plos Biol., 11, 2013
|
|
2LVL
 
 | | NMR Structure the lantibiotic immunity protein SpaI | | Descriptor: | SpaI | | Authors: | Christ, N, Bochmann, S, Gottstein, D, Duchardt-Ferner, E, Hellmich, U.A, Duesterhus, S, Koetter, P, Guentert, P, Entian, K, Woehnert, J. | | Deposit date: | 2012-07-06 | | Release date: | 2012-08-29 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The First Structure of a Lantibiotic Immunity Protein, SpaI from Bacillus subtilis, Reveals a Novel Fold.
J.Biol.Chem., 287, 2012
|
|
2MXO
 
 | |
2HWT
 
 | |
2MQF
 
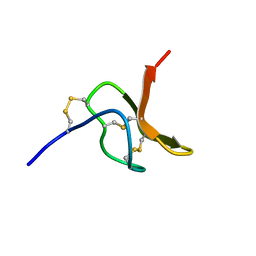 | | NMR structure of spider toxin-TRTX-Hhn2b | | Descriptor: | Mu-theraphotoxin-Hhn2b | | Authors: | Klint, J.K, Chin, Y.K.Y, Mobli, M. | | Deposit date: | 2014-06-19 | | Release date: | 2015-07-15 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Rational Engineering Defines a Molecular Switch That Is Essential for Activity of Spider-Venom Peptides against the Analgesics Target NaV1.7
Mol.Pharmacol., 88, 2015
|
|
2N2R
 
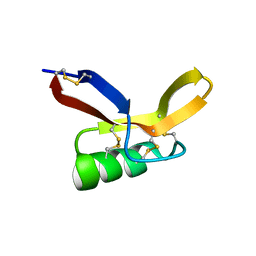 | | NMR solution structure of RsAFP2 | | Descriptor: | Defensin-like protein 2 | | Authors: | Harvey, P.J, Craik, D.J, Vriens, K. | | Deposit date: | 2015-05-12 | | Release date: | 2016-05-25 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | The radish defensins RsAFP1 and RsAFP2 act synergistically with caspofungin against Candida albicans biofilms.
Peptides, 75, 2016
|
|
2N2Q
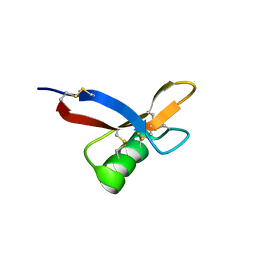 
 | | NMR solution structure of HsAFP1 | | Descriptor: | Defensin-like protein 1 | | Authors: | Harvey, P.J, Craik, D.J, Vriens, K. | | Deposit date: | 2015-05-11 | | Release date: | 2015-07-22 | | Last modified: | 2015-08-19 | | Method: | SOLUTION NMR | | Cite: | Synergistic Activity of the Plant Defensin HsAFP1 and Caspofungin against Candida albicans Biofilms and Planktonic Cultures.
Plos One, 10, 2015
|
|
1KJS
 
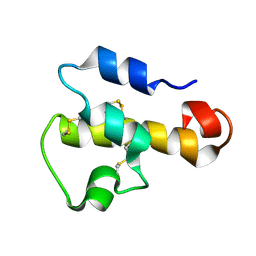 | | NMR SOLUTION STRUCTURE OF C5A AT PH 5.2, 303K, 20 STRUCTURES | | Descriptor: | C5A | | Authors: | Zhang, X, Boyar, W, Toth, M, Wennogle, L, Gonnella, N.C. | | Deposit date: | 1997-01-09 | | Release date: | 1997-05-15 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural definition of the C5a C terminus by two-dimensional nuclear magnetic resonance spectroscopy.
Proteins, 28, 1997
|
|
2KJH
 
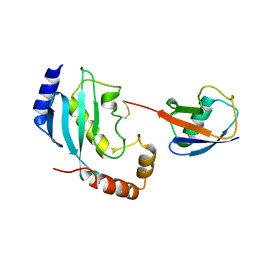 | |
2O00
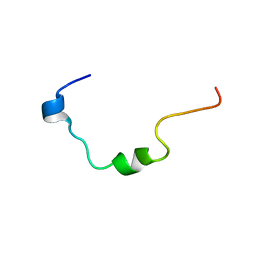 
 | | NMR structure analysis of the Penetratin conjugated Gas (374-394) peptide | | Descriptor: | Penetratin conjugated Gas (374-394) peptide | | Authors: | D'Ursi, A.M, Rovero, P, Albrizio, S, Espsito, C, D'Errico, G, Novellino, E. | | Deposit date: | 2006-11-27 | | Release date: | 2007-06-05 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Driving forces in the delivery of penetratin conjugated G protein fragment.
J.Med.Chem., 50, 2007
|
|
1KRI
 
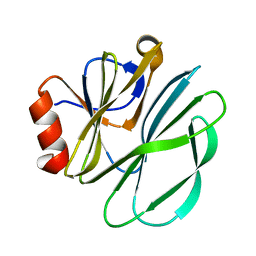 | |
2NZZ
 
 | | NMR structure analysis of the Penetratin conjugated Gas (374-394) peptide | | Descriptor: | Penetratin conjugated Gas (374-394) peptide | | Authors: | D'Ursi, A.M, Rovero, P, Albrizio, S, Espsito, C, D'Errico, G, Novellino, E. | | Deposit date: | 2006-11-27 | | Release date: | 2007-06-05 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Driving forces in the delivery of penetratin conjugated G protein fragment.
J.Med.Chem., 50, 2007
|
|
2AIH
 
 | | 1H-NMR solution structure of a trypsin/chymotrypsin Bowman-Birk inhibitor from Lens culinaris. | | Descriptor: | Bowman-Birk type protease inhibitor, LCTI, CHLORIDE ION | | Authors: | Ragg, E.M, Galbusera, V, Scarafoni, A, Negri, A, Tedeschi, G, Consonni, A, Sessa, F, Duranti, M. | | Deposit date: | 2005-07-29 | | Release date: | 2006-08-01 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Inhibitory properties and solution structure of a potent Bowman-Birk protease inhibitor from lentil (Lens culinaris, L) seeds.
Febs J., 273, 2006
|
|
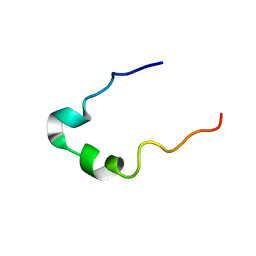
2GS0
 
 | | NMR structure of the complex between the PH domain of the Tfb1 subunit from TFIIH and the activation domain of p53 | | Descriptor: | Cellular tumor antigen p53, RNA polymerase II transcription factor B subunit 1 | | Authors: | Di Lello, P, Jones, T.N, Nguyen, B.D, Legault, P, Omichinski, J.G. | | Deposit date: | 2006-04-25 | | Release date: | 2006-10-31 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structure of the Tfb1/p53 complex: Insights into the interaction between the p62/Tfb1 subunit of TFIIH and the activation domain of p53.
Mol.Cell, 22, 2006
|
|
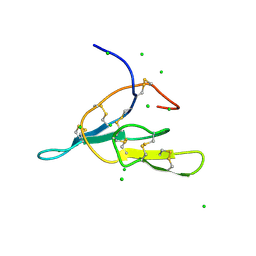
1JRF
 
 | | NMR Solution Structure of the Viral Receptor Domain of Tva | | Descriptor: | CALCIUM ION, SUBGROUP A ROUS SARCOMA VIRUS RECEPTORS PG800 AND PG950 | | Authors: | Wang, Q.-Y, Huang, W, Dolmer, K, Gettins, P.G.W, Rong, L. | | Deposit date: | 2001-08-13 | | Release date: | 2002-03-08 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the viral receptor domain of Tva and its implications in viral entry.
J.Virol., 76, 2002
|
|
