1VGM
 
 | |
2R26
 
 | |
2H12
 
 | |
6Z2H
 
 | |
6ZU0
 
 | |
2C6X
 
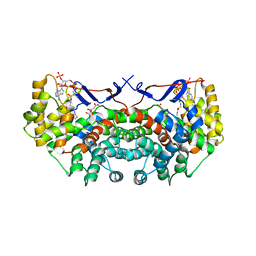 | |
2IFC
 
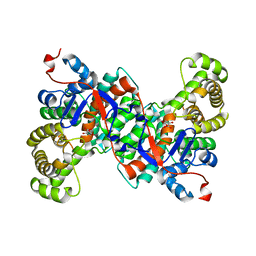 | |
2CSC
 
 | |
2CTS
 
 | |
2IBP
 
 | |
8S97
 
 | |
8S9D
 
 | |
8RJL
 
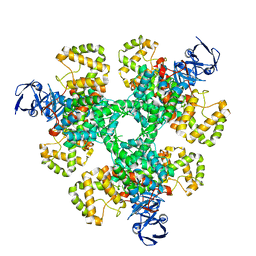
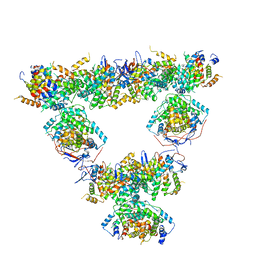 | | Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus | | Descriptor: | Citrate synthase | | Authors: | Lo, Y.K, Bohn, S, Sendker, F.L, Schuller, J.M, Hochberg, G. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (3.34 Å) | | Cite: | Emergence of fractal geometries in the evolution of a metabolic enzyme.
Nature, 628, 2024
|
|
8RJK
 
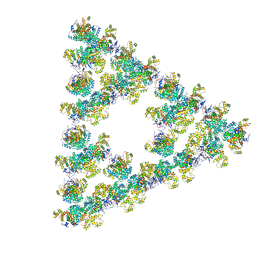 | | Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus | | Descriptor: | Citrate synthase | | Authors: | Lo, Y.K, Bohn, S, Sendker, F.L, Schuller, J.M, Hochberg, G. | | Deposit date: | 2023-12-21 | | Release date: | 2024-02-28 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (5.91 Å) | | Cite: | Emergence of fractal geometries in the evolution of a metabolic enzyme.
Nature, 628, 2024
|
|
6QCL
 
 | |
4JAD
 
 | |
4JAG
 
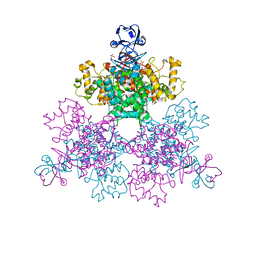 | |
6ABY
 
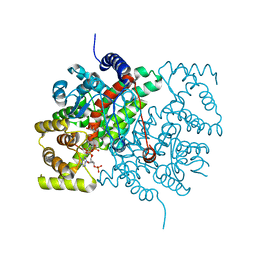
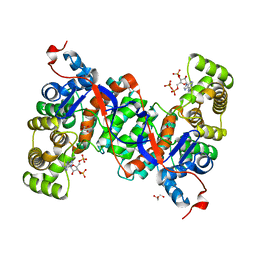 | | Crystal structure of citrate synthase (Msed_1522) from Metallosphaera sedula in complex with oxaloacetate | | Descriptor: | ACETYL COENZYME *A, Citrate synthase, GLYCEROL, ... | | Authors: | Lee, S.-H, Son, H.-F, Kim, K.-J. | | Deposit date: | 2018-07-24 | | Release date: | 2019-03-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights into the inhibition properties of archaeon citrate synthase from Metallosphaera sedula.
PLoS ONE, 14, 2019
|
|
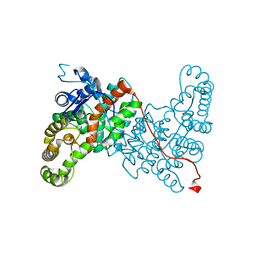
6ABV
 
 | |
4JAE
 
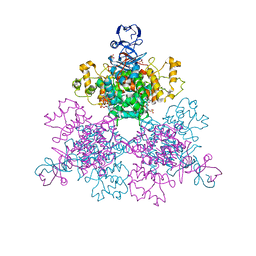 | |
4JAF
 
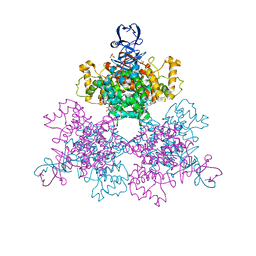 | |
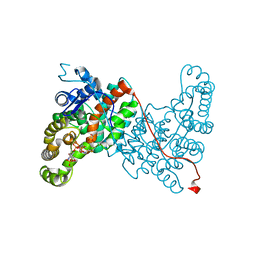
6ABX
 
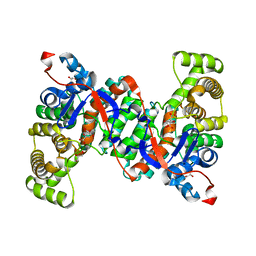 | |
6ABW
 
 | |
1O7X
 
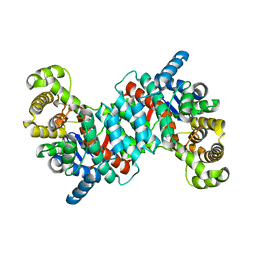 | | Citrate synthase from Sulfolobus solfataricus | | Descriptor: | CITRATE SYNTHASE | | Authors: | Bell, G.S, Russell, R.J.M, Connaris, H, Hough, D.W, Danson, M.J, Taylor, G.L. | | Deposit date: | 2002-11-19 | | Release date: | 2002-12-12 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Stepwise Adaptations of Citrate Synthase to Survival at Life'S Extremes. From Psychrophile to Hyperthermophile.
Eur.J.Biochem., 269, 2002
|
|
1OWC
 
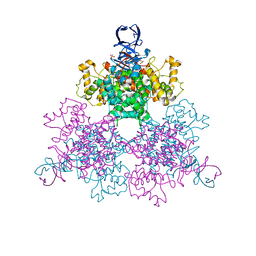 | | Three Dimensional Structure Analysis Of The R109L Variant of the Type II Citrate Synthase From E. Coli | | Descriptor: | Citrate synthase, SULFATE ION | | Authors: | Stokell, D.J, Donald, L.J, Maurus, R, Nguyen, N.T, Sadler, G, Choudhary, K, Hultin, P.G, Brayer, G.D, Duckworth, H.W. | | Deposit date: | 2003-03-28 | | Release date: | 2004-05-18 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Probing the roles of key residues in the unique regulatory NADH binding site of type II citrate synthase of Escherichia coli.
J.Biol.Chem., 278, 2003
|
|
