8W4C
 
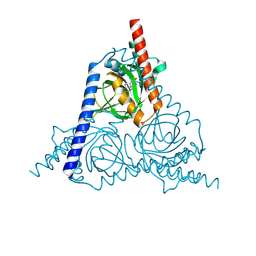 | | The sigma-1 receptor from Xenopus laevis in complex with progesterone by soaking | | Descriptor: | PROGESTERONE, Sigma non-opioid intracellular receptor 1 | | Authors: | Xiao, Y, Fu, C, Sun, Z, Zhou, X. | | Deposit date: | 2023-08-23 | | Release date: | 2024-07-17 | | Method: | X-RAY DIFFRACTION (2.676 Å) | | Cite: | Insight into binding of endogenous neurosteroid ligands to the sigma-1 receptor.
Nat Commun, 15, 2024
|
|
8W4B
 
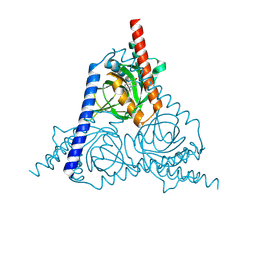 | | The sigma-1 receptor from Xenopus laevis in complex with progesterone by co-crystallization | | Descriptor: | PROGESTERONE, Sigma non-opioid intracellular receptor 1 | | Authors: | Xiao, Y, Fu, C, Sun, Z, Zhou, X. | | Deposit date: | 2023-08-23 | | Release date: | 2024-07-17 | | Method: | X-RAY DIFFRACTION (2.152 Å) | | Cite: | Insight into binding of endogenous neurosteroid ligands to the sigma-1 receptor.
Nat Commun, 15, 2024
|
|
8W4E
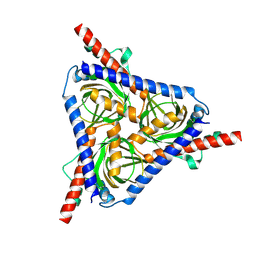 
 | |
8YBB
 
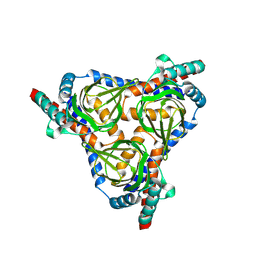 | |
8W4D
 
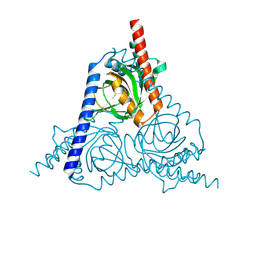 | |
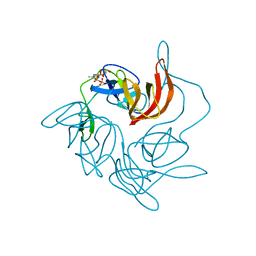
3GQK
 
 | |
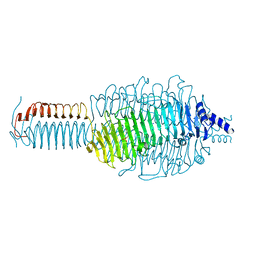
3GQ9
 
 | |
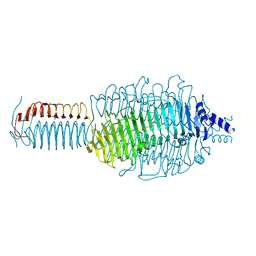
3GQ8
 
 | |
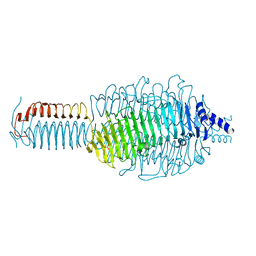
3GQA
 
 | |
3GQ7
 
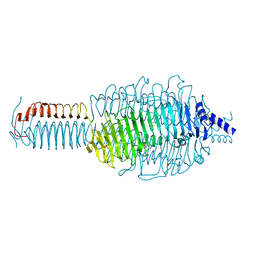 | |
3GQH
 
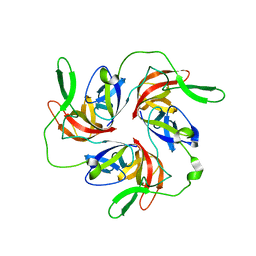 | |
7WNV
 
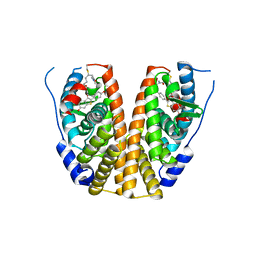 | | Crystal structure of mutant estrogen receptor alpha Y537S in complex with CO9 | | Descriptor: | (~{Z})-4-[2-[4-[[2-(4-hydroxyphenyl)-6-oxidanyl-1-benzothiophen-3-yl]oxy]phenoxy]ethylamino]-~{N},~{N}-dimethyl-but-2-enamide, Estrogen receptor | | Authors: | Xiao, Y, Lv, Y. | | Deposit date: | 2022-01-19 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | X-ray crystallography study and optimization of novel benzothiophene analogs as potent selective estrogen receptor covalent antagonists (SERCAs) with improved potency and safety profiles.
Bioorg.Chem., 141, 2023
|
|
3CSZ
 
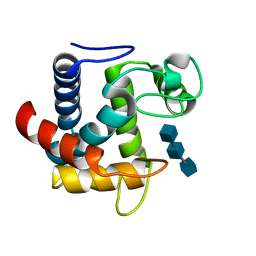 | |
3CT5
 
 | |
3CT0
 
 | |
3CSQ
 
 | |
3CSR
 
 | |
3CT1
 
 | |
3QC7
 
 | |
3R02
 
 | | The discovery of novel benzofuran-2-carboxylic acids as potent Pim-1 inhibitors | | Descriptor: | 7-[(cis-4-aminocyclohexyl)amino]-5-bromo-1-benzofuran-2-carboxylic acid, IMIDAZOLE, Proto-oncogene serine/threonine-protein kinase pim-1 | | Authors: | Xiang, Y, Hirth, B, Asmussen, G, Biemann, H.-P, Good, A, Fitzgerald, M, Gladysheva, T, Jancsics, K, Liu, J, Metz, M, Papoulis, A, Skerlj, R, Stepp, D.J, Wei, R.R. | | Deposit date: | 2011-03-07 | | Release date: | 2011-05-11 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The discovery of novel benzofuran-2-carboxylic acids as potent Pim-1 inhibitors.
Bioorg.Med.Chem.Lett., 21, 2011
|
|
3R04
 
 | | The discovery of novel benzofuran-2-carboxylic acids as potent Pim-1 inhibitors | | Descriptor: | 5-{6-[(trans-4-aminocyclohexyl)amino]pyrazin-2-yl}-1-benzofuran-2-carboxylic acid, IMIDAZOLE, Proto-oncogene serine/threonine-protein kinase pim-1 | | Authors: | Xiang, Y, Hirth, B, Asmussen, G, Biemann, H.-P, Good, A, Fitzgerald, M, Gladysheva, T, Jancsics, K, Liu, J, Metz, M, Papoulis, A, Skerlj, R, Stepp, D.J, Wei, R.R. | | Deposit date: | 2011-03-07 | | Release date: | 2011-05-11 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The discovery of novel benzofuran-2-carboxylic acids as potent Pim-1 inhibitors.
Bioorg.Med.Chem.Lett., 21, 2011
|
|
7SWF
 
 | |
7SWQ
 
 | |
7SVA
 
 | | Cryo-EM structure of Arabidopsis Ago10-guide RNA complex | | Descriptor: | MAGNESIUM ION, Protein argonaute 10, RNA (5'-R(P*UP*GP*GP*AP*GP*UP*GP*UP*GP*AP*CP*AP*AP*UP*GP*GP*UP*GP*UP*UP*U)-3') | | Authors: | Xiao, Y, MacRae, I.J. | | Deposit date: | 2021-11-18 | | Release date: | 2022-11-23 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Structural basis for RNA slicing by a plant Argonaute.
Nat.Struct.Mol.Biol., 30, 2023
|
|
8IXJ
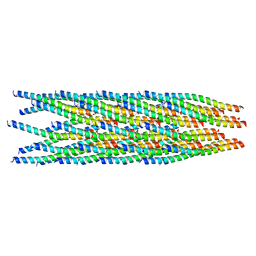 
 | |
