4JG0
 
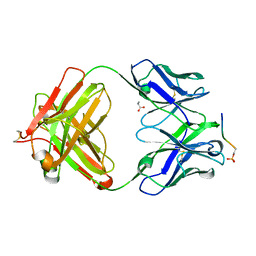 | | Structure of phosphoserine/threonine (pSTAb) scaffold bound to pSer peptide | | Descriptor: | Fab heavy chain, Fab light chain, PROPANOIC ACID, ... | | Authors: | Koerber, J.T, Thomsen, N.D, Hannigan, B.T, Degrado, W.F, Wells, J.A. | | Deposit date: | 2013-02-28 | | Release date: | 2013-08-28 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Nature-inspired design of motif-specific antibody scaffolds.
Nat.Biotechnol., 31, 2013
|
|
4JFZ
 
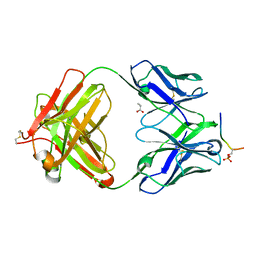 | | Structure of phosphoserine (pSAb) scaffold bound to pSer peptide | | Descriptor: | Fab heavy chain, Fab light chain, PROPANOIC ACID, ... | | Authors: | Koerber, J.T, Thomsen, N.D, Hannigan, B.T, Degrado, W.F, Wells, J.A. | | Deposit date: | 2013-02-28 | | Release date: | 2013-08-28 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Nature-inspired design of motif-specific antibody scaffolds.
Nat.Biotechnol., 31, 2013
|
|
4JQZ
 
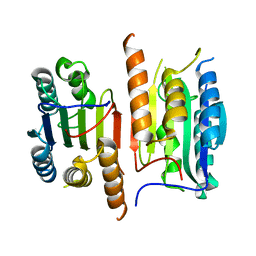 | | Human procaspase-3, crystal form 2 | | Descriptor: | Procaspase-3 | | Authors: | Thomsen, N.D, Wells, J.A. | | Deposit date: | 2013-03-20 | | Release date: | 2013-05-08 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.889 Å) | | Cite: | Structural snapshots reveal distinct mechanisms of procaspase-3 and -7 activation.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4JG1
 
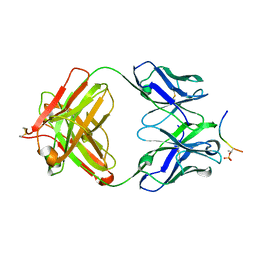 | | Structure of phosphoserine/threonine (pSTAb) scaffold bound to pThr peptide | | Descriptor: | Fab heavy chain, Fab light chain, PROPANOIC ACID, ... | | Authors: | Koerber, J.T, Thomsen, N.D, Hannigan, B.T, Degrado, W.F, Wells, J.A. | | Deposit date: | 2013-02-28 | | Release date: | 2013-08-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Nature-inspired design of motif-specific antibody scaffolds.
Nat.Biotechnol., 31, 2013
|
|
4JQY
 
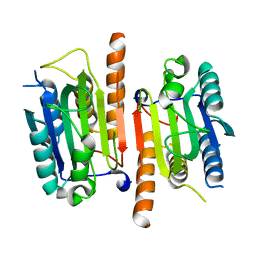 | | Human procaspase-3, crystal form 1 | | Descriptor: | Procaspase-3 | | Authors: | Thomsen, N.D, Wells, J.A. | | Deposit date: | 2013-03-20 | | Release date: | 2013-05-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.495 Å) | | Cite: | Structural snapshots reveal distinct mechanisms of procaspase-3 and -7 activation.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4L8G
 
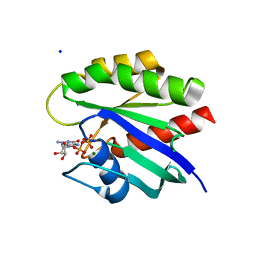 | | Crystal Structure of K-Ras G12C, GDP-bound | | Descriptor: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Ostrem, J.M, Peters, U, Sos, M.L, Wells, J.A, Shokat, K.M. | | Deposit date: | 2013-06-17 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.521 Å) | | Cite: | K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions.
Nature, 503, 2013
|
|
4L9W
 
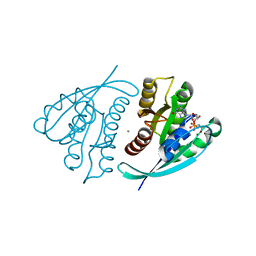 | | Crystal Structure of H-Ras G12C, GMPPNP-bound | | Descriptor: | CALCIUM ION, GTPase HRas, MAGNESIUM ION, ... | | Authors: | Ostrem, J.M, Peters, U, Sos, M.L, Wells, J.A, Shokat, K.M. | | Deposit date: | 2013-06-18 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.952 Å) | | Cite: | K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions.
Nature, 503, 2013
|
|
4L9S
 
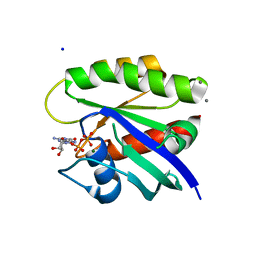 | | Crystal Structure of H-Ras G12C, GDP-bound | | Descriptor: | CALCIUM ION, GTPase HRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Ostrem, J.M, Peters, U, Sos, M.L, Wells, J.A, Shokat, K.M. | | Deposit date: | 2013-06-18 | | Release date: | 2013-11-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.606 Å) | | Cite: | K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions.
Nature, 503, 2013
|
|
4JR0
 | | Human procaspase-3 bound to Ac-DEVD-CMK | | Descriptor: | Ac-DEVD-CMK, CHLORIDE ION, Procaspase-3 | | Authors: | Thomsen, N.D, Wells, J.A. | | Deposit date: | 2013-03-20 | | Release date: | 2013-05-08 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.796 Å) | | Cite: | Structural snapshots reveal distinct mechanisms of procaspase-3 and -7 activation.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4JR2
 
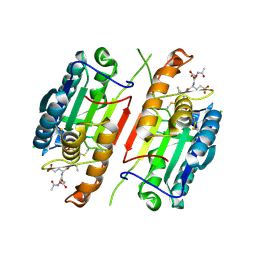 | |
4JFX
 
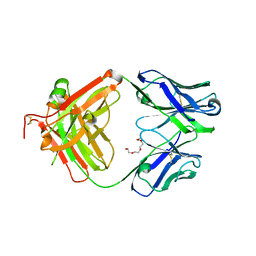 | | Structure of phosphotyrosine (pTyr) scaffold bound to pTyr peptide | | Descriptor: | Fab heavy chain, Fab light chain, Phosphopeptide, ... | | Authors: | Koerber, J.T, Thomsen, N.D, Hannigan, B.T, Degrado, W.F, Wells, J.A. | | Deposit date: | 2013-02-28 | | Release date: | 2013-09-25 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Nature-inspired design of motif-specific antibody scaffolds.
Nat.Biotechnol., 31, 2013
|
|
4JFY
 
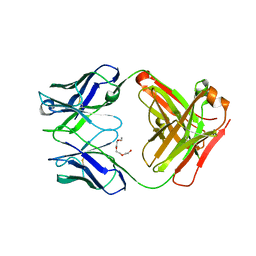 | | Apo structure of phosphotyrosine (pYAb) scaffold | | Descriptor: | Fab heavy chain, Fab light chain, TETRAETHYLENE GLYCOL | | Authors: | Koerber, J.T, Thomsen, N.D, Hannigan, B.T, Degrado, W.F, Wells, J.A. | | Deposit date: | 2013-02-28 | | Release date: | 2013-08-28 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Nature-inspired design of motif-specific antibody scaffolds.
Nat.Biotechnol., 31, 2013
|
|
1QVN
 
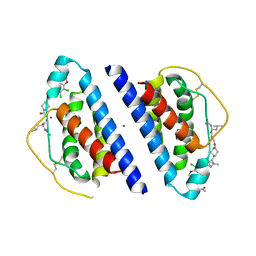 | | Structure of SP4160 Bound to IL-2 V69A | | Descriptor: | 2-GUANIDINO-4-METHYL-PENTANOIC ACID [2-(4-{5-[4-(4-ACETYLAMINO-BENZYLOXY)-2,3-DICHLORO-PHENYL]-2-METHYL-2H-PYRAZOL-3-YL}-PIPERIDIN-1-YL)-2-OXO-ETHYL]-AMIDE, Interleukin-2, ZINC ION | | Authors: | Thanos, C.D, Delano, W.L, Wells, J.A. | | Deposit date: | 2003-08-28 | | Release date: | 2005-04-05 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Hot-spot mimicry of a cytokine receptor by a small molecule.
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
1PW6
 | | Low Micromolar Small Molecule Inhibitor of IL-2 | | Descriptor: | 2-CYCLOHEXYL-N-(2-{4-[5-(2,3-DICHLORO-PHENYL)-2H-PYRAZOL-3-YL]-PIPERIDIN-1-YL}-2-OXO-ETHYL)-2-GUANIDINO-ACETAMIDE, Interleukin-2, SULFATE ION | | Authors: | Thanos, C.D, Randal, M, Wells, J.A. | | Deposit date: | 2003-06-30 | | Release date: | 2004-01-13 | | Last modified: | 2018-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Potent small-molecule binding to a dynamic hot spot on IL-2.
J.Am.Chem.Soc., 125, 2003
|
|
1PY2
 | | Structure of a 60 nM Small Molecule Bound to a Hot Spot on IL-2 | | Descriptor: | 5-[2,3-DICHLORO-4-(5-{1-[2-(2-GUANIDINO-4-METHYL-PENTANOYLAMINO)-ACETYL]-PIPERIDIN-4-YL}-1-METHYL-1H-PYRAZOL-3-YL)-PHENOXYMETHYL]-FURAN-2-CARBOXYLIC ACID, Interleukin-2, ZINC ION | | Authors: | Thanos, C.D, Randal, M, Wells, J.A. | | Deposit date: | 2003-07-07 | | Release date: | 2004-01-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Potent small-molecule binding to a dynamic hot spot on IL-2.
J.Am.Chem.Soc., 125, 2003
|
|
2H4W
 
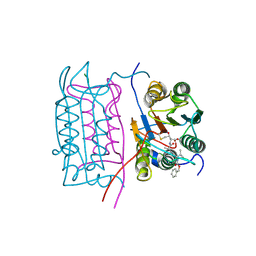 | |
1SHJ
 
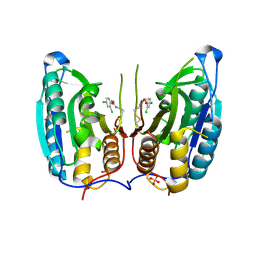 | | Caspase-7 in complex with DICA allosteric inhibitor | | Descriptor: | 2-(2,4-DICHLORO-PHENOXY)-N-(2-MERCAPTO-ETHYL)-ACETAMIDE, Caspase-7, SULFATE ION | | Authors: | Hardy, J.A, Lam, J, Nguyen, J.T, O'Brien, T, Wells, J.A. | | Deposit date: | 2004-02-25 | | Release date: | 2004-08-17 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Discovery of an allosteric site in the caspases
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
2HBQ
 
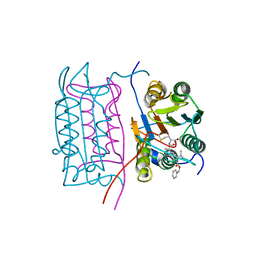 | |
2H48
 
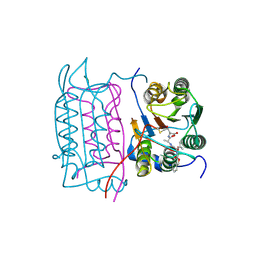 | | Crystal structure of human caspase-1 (Cys362->Ala, Cys364->Ala, Cys397->Ala) in complex with 3-[2-(2-benzyloxycarbonylamino-3-methyl-butyrylamino)-propionylamino]-4-oxo-pentanoic acid (z-VAD-FMK) | | Descriptor: | Caspase 1, isoform gamma, N-[(benzyloxy)carbonyl]-L-valyl-N-[(2S)-1-carboxy-4-fluoro-3-oxobutan-2-yl]-L-alaninamide | | Authors: | Scheer, J.M, Wells, J.A, Romanowski, M.J. | | Deposit date: | 2006-05-23 | | Release date: | 2006-06-06 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A common allosteric site and mechanism in caspases
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
2H54
 
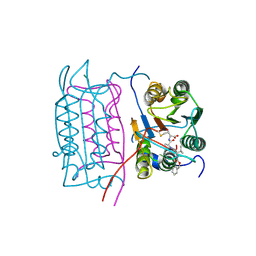 | |
1SHL
 
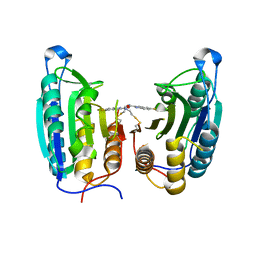 | | CASPASE-7 IN COMPLEX WITH FICA ALLOSTERIC INHIBITOR | | Descriptor: | 5-FLUORO-1H-INDOLE-2-CARBOXYLIC ACID-(2-MERCAPTO-ETHYL)-AMIDE, Caspase-7 | | Authors: | Hardy, J.A, Lam, J, Nguyen, J.T, O'Brien, T, Wells, J.A. | | Deposit date: | 2004-02-25 | | Release date: | 2004-08-17 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Discovery of an allosteric site in the caspases
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
2FQQ
 
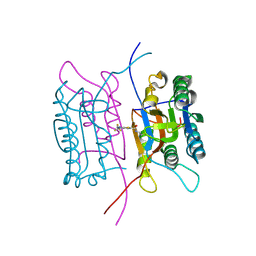 | | Crystal structure of human caspase-1 (Cys285->Ala, Cys362->Ala, Cys364->Ala, Cys397->Ala) in complex with 1-methyl-3-trifluoromethyl-1H-thieno[2,3-c]pyrazole-5-carboxylic acid (2-mercapto-ethyl)-amide | | Descriptor: | 1-METHYL-3-TRIFLUOROMETHYL-1H-THIENO[2,3-C]PYRAZOLE-5-CARBOXYLIC ACID (2-MERCAPTO-ETHYL)-AMIDE, Caspase-1 | | Authors: | Scheer, J.M, Wells, J.A, Romanowski, M.J. | | Deposit date: | 2006-01-18 | | Release date: | 2006-05-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | A common allosteric site and mechanism in caspases
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
2H4Y
 
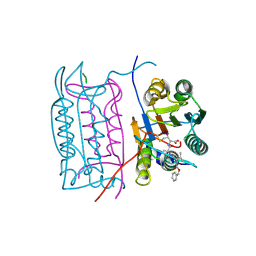 | |
2HBR
 
 | |
2HBZ
 
 | |
