2INT
 
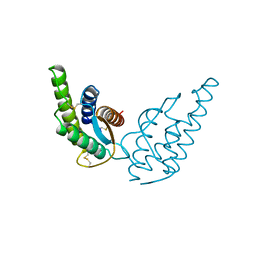 | | CRYSTAL STRUCTURE OF RECOMBINANT HUMAN INTERLEUKIN-4 | | Descriptor: | INTERLEUKIN-4 | | Authors: | Walter, M.R, Cook, W.J, Zhao, B.G, Cameron Junior, R, Ealick, S.E, Walter Junior, R.L, Reichert, P, Nagabhushan, T.L, Trotta, P.P, Bugg, C.E. | | Deposit date: | 1993-07-22 | | Release date: | 1994-01-31 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of recombinant human interleukin-4.
J.Biol.Chem., 267, 1992
|
|
1INR
 
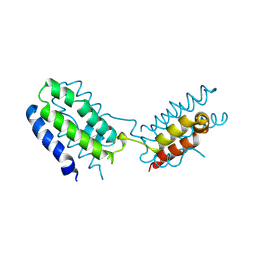 | | CYTOKINE SYNTHESIS | | Descriptor: | INTERLEUKIN-10 | | Authors: | Walter, M.R. | | Deposit date: | 1995-07-31 | | Release date: | 1996-10-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of interleukin 10 reveals an interferon gamma-like fold.
Biochemistry, 34, 1995
|
|
1RH2
 
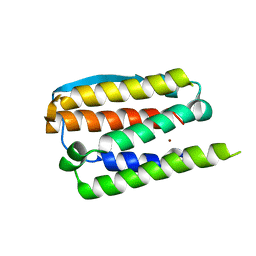 | | RECOMBINANT HUMAN INTERFERON-ALPHA 2B | | Descriptor: | INTERFERON-ALPHA 2B, ZINC ION | | Authors: | Walter, M.R. | | Deposit date: | 1996-11-07 | | Release date: | 1997-11-12 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Zinc mediated dimer of human interferon-alpha 2b revealed by X-ray crystallography.
Structure, 4, 1996
|
|
1TPT
 
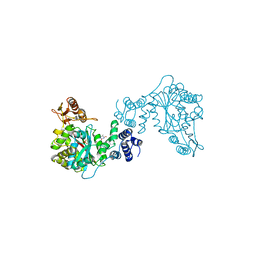 | | THREE-DIMENSIONAL STRUCTURE OF THYMIDINE PHOSPHORYLASE FROM ESCHERICHIA COLI AT 2.8 ANGSTROMS RESOLUTION | | Descriptor: | SULFATE ION, THYMIDINE PHOSPHORYLASE, THYMINE | | Authors: | Walter, M.R, Cook, W.J, Cole, L.B, Short, S.A, Koszalka, G.W, Krenitsky, T.A, Ealick, S.E. | | Deposit date: | 1990-06-14 | | Release date: | 1991-07-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Three-dimensional structure of thymidine phosphorylase from Escherichia coli at 2.8 A resolution.
J.Biol.Chem., 265, 1990
|
|
2V5I
 
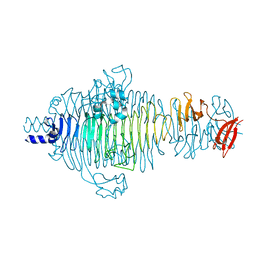 | | Structure of the receptor-binding protein of bacteriophage Det7: a podoviral tailspike in a myovirus | | Descriptor: | SALMONELLA TYPHIMURIUM DB7155 BACTERIOPHAGE DET7 TAILSPIKE, SODIUM ION | | Authors: | Walter, M, Fiedler, C, Grassl, R, Biebl, M, Rachel, R, Hermo-Parrado, X.L, Llamas-Saiz, A.L, Seckler, R, Miller, S, van Raaij, M.J. | | Deposit date: | 2007-07-05 | | Release date: | 2008-02-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of the Receptor-Binding Protein of Bacteriophage Det7: A Podoviral Tail Spike in a Myovirus.
J.Virol., 82, 2008
|
|
1CSG
 
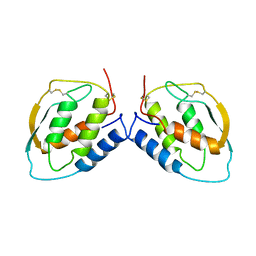 | |
2OGV
 
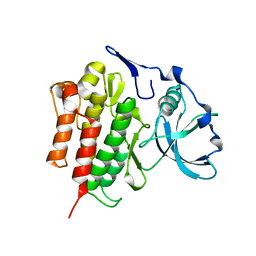 | | Crystal Structure of the Autoinhibited Human c-Fms Kinase Domain | | Descriptor: | Macrophage colony-stimulating factor 1 receptor precursor | | Authors: | Walter, M, Lucet, I.S, Patel, O, Broughton, S.E, Bamert, R, Williams, N.K, Fantino, E, Wilks, A.F, Rossjohn, J. | | Deposit date: | 2007-01-09 | | Release date: | 2007-02-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The 2.7 A crystal structure of the autoinhibited human c-Fms kinase domain.
J.Mol.Biol., 367, 2007
|
|
2TPT
 
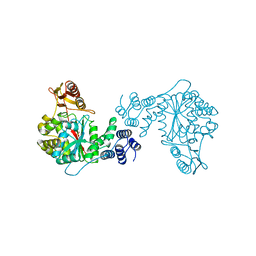 | | STRUCTURAL AND THEORETICAL STUDIES SUGGEST DOMAIN MOVEMENT PRODUCES AN ACTIVE CONFORMATION OF THYMIDINE PHOSPHORYLASE | | Descriptor: | SULFATE ION, THYMIDINE PHOSPHORYLASE | | Authors: | Pugmire, M.J, Cook, W.J, Jasanoff, A, Walter, M.R, Ealick, S.E. | | Deposit date: | 1997-11-24 | | Release date: | 1999-03-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural and theoretical studies suggest domain movement produces an active conformation of thymidine phosphorylase.
J.Mol.Biol., 281, 1998
|
|
3LQM
 
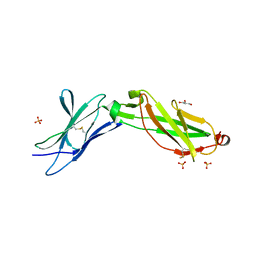 | | Structure of the IL-10R2 Common Chain | | Descriptor: | GLYCEROL, Interleukin-10 receptor subunit beta, SULFATE ION | | Authors: | Yoon, S.I, Walter, M.R. | | Deposit date: | 2010-02-09 | | Release date: | 2010-05-26 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structure and mechanism of receptor sharing by the IL-10R2 common chain.
Structure, 18, 2010
|
|
1LK3
 
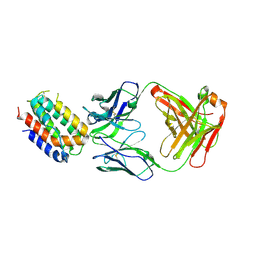 | | ENGINEERED HUMAN INTERLEUKIN-10 MONOMER COMPLEXED TO 9D7 FAB FRAGMENT | | Descriptor: | 9D7 Heavy Chain, 9D7 Light Chain, Interleukin-10 | | Authors: | Josephson, K, Jones, B.C, Walter, L.J, DiGiacomo, R, Indelicato, S.R, Walter, M.R. | | Deposit date: | 2002-04-23 | | Release date: | 2002-07-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Noncompetitive antibody neutralization of IL-10 revealed by protein engineering and x-ray crystallography.
Structure, 10, 2002
|
|
3BES
 
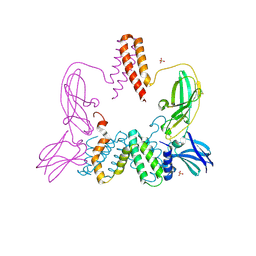 | | Structure of a Poxvirus ifngbp/ifng Complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Interferon gamma, Interferon-gamma binding protein C4R, ... | | Authors: | Nuara, A.A, Walter, M.R. | | Deposit date: | 2007-11-20 | | Release date: | 2008-02-12 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure and mechanism of IFN-gamma antagonism by an orthopoxvirus IFN-gamma-binding protein.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
8FAX
 
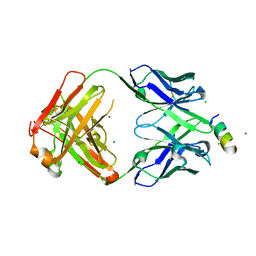 | | Fab 1249A8-MERS Stem Helix Complex | | Descriptor: | 1249A8-HC, 1249A8-LC, CHLORIDE ION, ... | | Authors: | Deshpande, A, Schormann, N, Piepenbrink, M.S, Martinez-Sobrido, L, Kobie, J.J, Walter, M.R. | | Deposit date: | 2022-11-28 | | Release date: | 2023-05-03 | | Last modified: | 2023-07-19 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure and epitope of a neutralizing monoclonal antibody that targets the stem helix of beta coronaviruses.
Febs J., 290, 2023
|
|
1B5L
 
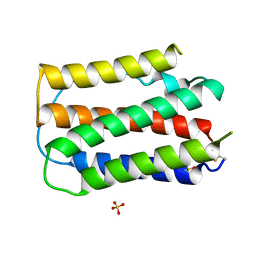 | | OVINE INTERFERON TAU | | Descriptor: | INTERFERON TAU, SULFATE ION | | Authors: | Radhakrishnan, R, Walter, L.J, Subramaniam, P.S, Johnson, H.J, Walter, M.R. | | Deposit date: | 1999-01-07 | | Release date: | 1999-05-18 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of ovine interferon-tau at 2.1 A resolution.
J.Mol.Biol., 286, 1999
|
|
1CTR
 
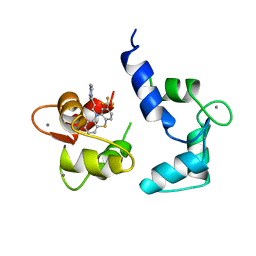 | |
4DOH
 
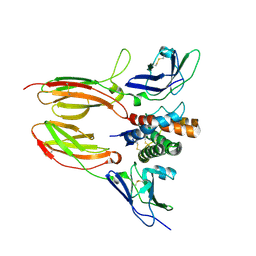 | | IL20/IL201/IL20R2 Ternary Complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Interleukin-20, Interleukin-20 receptor subunit alpha, ... | | Authors: | Logsdon, N.J, Walter, M.R. | | Deposit date: | 2012-02-09 | | Release date: | 2012-07-18 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural basis for receptor sharing and activation by interleukin-20 receptor-2 (IL-20R2) binding cytokines.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1LQS
 
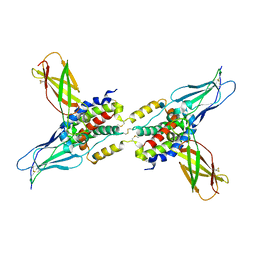 | | CRYSTAL STRUCTURE OF HUMAN CYTOMEGALOVIRUS IL-10 BOUND TO SOLUBLE HUMAN IL-10R1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, INTERLEUKIN-10 RECEPTOR ALPHA CHAIN, INTERLEUKIN-10-LIKE PROTEIN | | Authors: | Jones, B.C, Logsdon, N.J, Josephson, K, Cook, J, Barry, P.A, Walter, M.R. | | Deposit date: | 2002-05-13 | | Release date: | 2002-07-17 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of human cytomegalovirus IL-10 bound to soluble human IL-10R1.
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1J7V
 
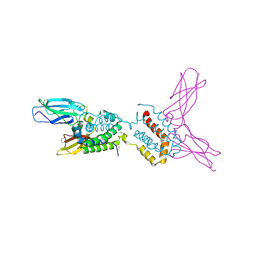 | | HUMAN IL-10 / IL-10R1 COMPLEX | | Descriptor: | INTERLEUKIN-10, INTERLEUKIN-10 RECEPTOR ALPHA CHAIN | | Authors: | Josephson, K, Logsdon, N, Walter, M.R. | | Deposit date: | 2001-05-18 | | Release date: | 2001-09-19 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of the IL-10/IL-10R1 complex reveals a shared receptor binding site.
Immunity, 15, 2001
|
|
3DGC
 
 | | Structure of IL-22/IL-22R1 | | Descriptor: | ACETATE ION, CHLORIDE ION, Interleukin-22, ... | | Authors: | Jones, B.C, Logsdon, N.J, Walter, M.R. | | Deposit date: | 2008-06-13 | | Release date: | 2008-07-15 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of IL-22 Bound to Its High-Affinity IL-22R1 Chain.
Structure, 16, 2008
|
|
1OTP
 
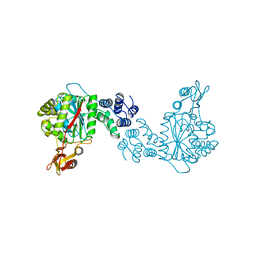 | | STRUCTURAL AND THEORETICAL STUDIES SUGGEST DOMAIN MOVEMENT PRODUCES AN ACTIVE CONFORMATION OF THYMIDINE PHOSPHORYLASE | | Descriptor: | THYMIDINE PHOSPHORYLASE | | Authors: | Pugmire, M.J, Cook, W.J, Jasanoff, A, Walter, M.R, Ealick, S.E. | | Deposit date: | 1997-11-09 | | Release date: | 1998-12-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural and theoretical studies suggest domain movement produces an active conformation of thymidine phosphorylase.
J.Mol.Biol., 281, 1998
|
|
2B7A
 
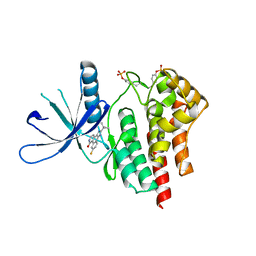 | | The structural basis of Janus Kinase 2 inhibition by a potent and specific pan-Janus kinase inhibitor | | Descriptor: | 2-TERT-BUTYL-9-FLUORO-3,6-DIHYDRO-7H-BENZ[H]-IMIDAZ[4,5-F]ISOQUINOLINE-7-ONE, Tyrosine-protein kinase JAK2 | | Authors: | Lucet, I.S, Fantino, E, Styles, M, Bamert, R, Patel, O, Broughton, S.E, Walter, M, Burns, C.J, Treutlein, H, Wilks, A.F, Rossjohn, J. | | Deposit date: | 2005-10-04 | | Release date: | 2006-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structural basis of Janus kinase 2 inhibition by a potent and specific pan-Janus kinase inhibitor.
Blood, 107, 2006
|
|
1AZY
 
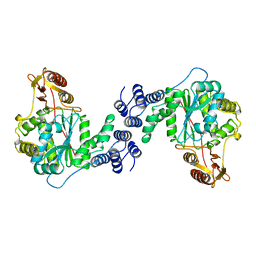 | | STRUCTURAL AND THEORETICAL STUDIES SUGGEST DOMAIN MOVEMENT PRODUCES AN ACTIVE CONFORMATION OF THYMIDINE PHOSPHORYLASE | | Descriptor: | THYMIDINE PHOSPHORYLASE | | Authors: | Pugmire, M.J, Cook, W.J, Jasanoff, A, Walter, M.R, Ealick, S.E. | | Deposit date: | 1997-11-24 | | Release date: | 1999-01-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and theoretical studies suggest domain movement produces an active conformation of thymidine phosphorylase.
J.Mol.Biol., 281, 1998
|
|
1EKU
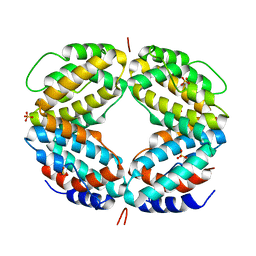 
 | | CRYSTAL STRUCTURE OF A BIOLOGICALLY ACTIVE SINGLE CHAIN MUTANT OF HUMAN IFN-GAMMA | | Descriptor: | Interferon gamma, SULFATE ION | | Authors: | Landar, A, Curry, B, Parker, M.H, DiGiacomo, R, Indelicato, S.R, Walter, M.R. | | Deposit date: | 2000-03-09 | | Release date: | 2000-09-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Design, characterization, and structure of a biologically active single-chain mutant of human IFN-gamma.
J.Mol.Biol., 299, 2000
|
|
1YKB
 
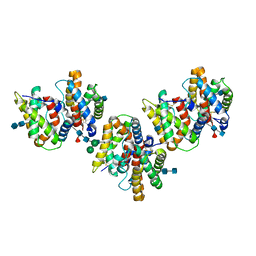 | | Crystal Structure of Insect Cell Expressed IL-22 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Xu, T, Logsdon, N.J, Walter, M.R. | | Deposit date: | 2005-01-17 | | Release date: | 2005-07-12 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of insect-cell-derived IL-22.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1Y6N
 
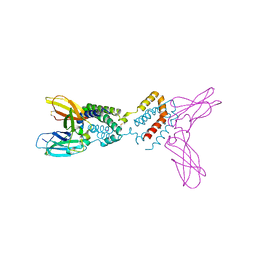 | | Crystal structure of Epstein-Barr virus IL-10 mutant (A87I) complexed with the soluble IL-10R1 chain | | Descriptor: | Interleukin-10 receptor alpha chain, Viral interleukin-10 homolog | | Authors: | Yoon, S.I, Jones, B.C, Logsdon, N.J, Walter, M.R. | | Deposit date: | 2004-12-06 | | Release date: | 2005-05-03 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Same structure, different function crystal structure of the Epstein-Barr virus IL-10 bound to the soluble IL-10R1 chain.
Structure, 13, 2005
|
|
1Y6M
 
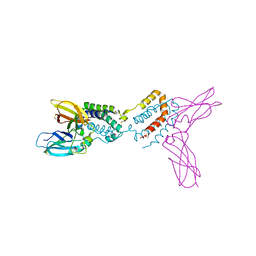 | | Crystal structure of Epstein-Barr virus IL-10 complexed with the soluble IL-10R1 chain | | Descriptor: | Interleukin-10 receptor alpha chain, Viral interleukin-10 homolog | | Authors: | Yoon, S.I, Jones, B.C, Logsdon, N.J, Walter, M.R. | | Deposit date: | 2004-12-06 | | Release date: | 2005-05-03 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Same structure, different function crystal structure of the Epstein-Barr virus IL-10 bound to the soluble IL-10R1 chain.
Structure, 13, 2005
|
|
