2YFP
 | | STRUCTURE OF YELLOW-EMISSION VARIANT OF GFP | | Descriptor: | PROTEIN (GREEN FLUORESCENT PROTEIN) | | Authors: | Wachter, R.M, Elsliger, M.A, Kallio, K, Hanson, G.T, Remington, S.J. | | Deposit date: | 1998-08-17 | | Release date: | 1999-01-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural basis of spectral shifts in the yellow-emission variants of green fluorescent protein.
Structure, 6, 1998
|
|
1F09
 
 | | CRYSTAL STRUCTURE OF THE GREEN FLUORESCENT PROTEIN (GFP) VARIANT YFP-H148Q WITH TWO BOUND IODIDES | | Descriptor: | GREEN FLUORESCENT PROTEIN, IODIDE ION | | Authors: | Wachter, R.M, Yarbrough, D, Kallio, K, Remington, S.J. | | Deposit date: | 2000-05-15 | | Release date: | 2000-11-17 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Crystallographic and energetic analysis of binding of selected anions to the yellow variants of green fluorescent protein.
J.Mol.Biol., 301, 2000
|
|
1F0B
 
 | |
1BFP
 
 | | BLUE VARIANT OF GREEN FLUORESCENT PROTEIN | | Descriptor: | BLUE FLUORESCENT PROTEIN | | Authors: | Wachter, R.M, Remington, S.J. | | Deposit date: | 1997-04-09 | | Release date: | 1997-07-07 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure and photodynamic behavior of the blue emission variant Y66H/Y145F of green fluorescent protein.
Biochemistry, 36, 1997
|
|
1YFP
 | | STRUCTURE OF YELLOW-EMISSION VARIANT OF GFP | | Descriptor: | YELLOW FLUORESCENT PROTEIN | | Authors: | Wachter, R.M, Elsliger, M.-A, Kallio, K, Hanson, G.T, Remington, S.J. | | Deposit date: | 1998-08-28 | | Release date: | 1998-10-28 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis of spectral shifts in the yellow-emission variants of green fluorescent protein.
Structure, 6, 1998
|
|
1XA9
 
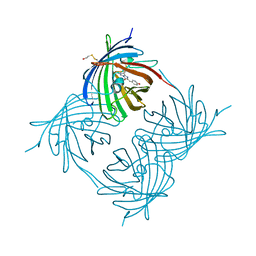 | | Crystal structure of yellow fluorescent protein zFP538 K66M green mutant | | Descriptor: | BETA-MERCAPTOETHANOL, fluorescent protein FP538 | | Authors: | Remington, S.J, Wachter, R.M, Yarbrough, D.K, Branchaud, B, Anderson, D.C, Kallio, K, Lukyanov, K.A. | | Deposit date: | 2004-08-25 | | Release date: | 2005-02-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | zFP538, a yellow-fluorescent protein from Zoanthus, contains a novel three-ring chromophore.
Biochemistry, 44, 2005
|
|
3THG
 
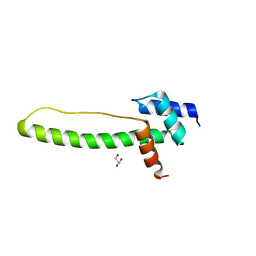 | | Crystal structure of the creosote Rubisco activase C-domain | | Descriptor: | GLYCEROL, Ribulose bisphosphate carboxylase/oxygenase activase 1, chloroplastic | | Authors: | Henderson, J.N, Kuriata, A.M, Fromme, R, Salvucci, M.E, Wachter, R.M. | | Deposit date: | 2011-08-18 | | Release date: | 2011-08-31 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Atomic resolution x-ray structure of the substrate recognition domain of higher plant ribulose-bisphosphate carboxylase/oxygenase (Rubisco) activase.
J.Biol.Chem., 286, 2011
|
|
4GOB
 
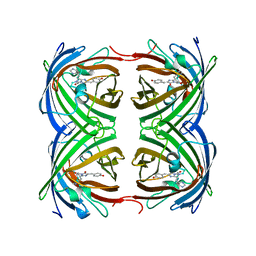 | | Low pH Crystal Structure of a reconstructed Kaede-type Red Fluorescent Protein, Least Evolved Ancestor (LEA) | | Descriptor: | Kaede-type Fluorescent Protein | | Authors: | Kim, H, Grunkemeyer, T.J, Chen, L, Fromme, R, Wachter, R.M. | | Deposit date: | 2012-08-19 | | Release date: | 2013-07-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Acid-base catalysis and crystal structures of a least evolved ancestral GFP-like protein undergoing green-to-red photoconversion.
Biochemistry, 52, 2013
|
|
3P28
 | |
1S6Z
 
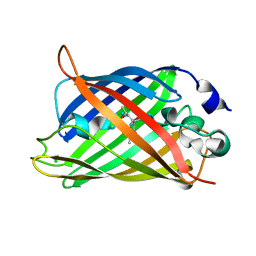 | | Enhanced Green Fluorescent Protein Containing the Y66L Substitution | | Descriptor: | CHLORIDE ION, green fluorescent protein | | Authors: | Rosenow, M.A, Huffman, H.A, Phail, M.E, Wachter, R.M. | | Deposit date: | 2004-01-28 | | Release date: | 2004-05-04 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Crystal Structure of the Y66L Variant of Green Fluorescent Protein Supports a Cyclization-Oxidation-Dehydration Mechanism for Chromophore Maturation(,).
Biochemistry, 43, 2004
|
|
1C4F
 
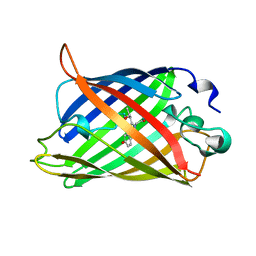 | | GREEN FLUORESCENT PROTEIN S65T AT PH 4.6 | | Descriptor: | GREEN FLUORESCENT PROTEIN | | Authors: | Elsliger, M.A, Wachter, R.M, Kallio, K, Hanson, G.T, Remington, S.J. | | Deposit date: | 1999-08-21 | | Release date: | 1999-08-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and spectral response of green fluorescent protein variants to changes in pH.
Biochemistry, 38, 1999
|
|
1EMG
 
 | | GREEN FLUORESCENT PROTEIN (65-67 REPLACED BY CRO, S65T SUBSTITUTION, Q80R) | | Descriptor: | PROTEIN (GREEN FLUORESCENT PROTEIN) | | Authors: | Elsliger, M.A, Wachter, R.M, Kallio, K, Hanson, G.T, Remington, S.J. | | Deposit date: | 1998-11-12 | | Release date: | 1999-05-12 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and spectral response of green fluorescent protein variants to changes in pH.
Biochemistry, 38, 1999
|
|
1XAE
 
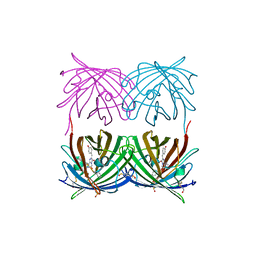 | | Crystal structure of wild type yellow fluorescent protein zFP538 from Zoanthus | | Descriptor: | BETA-MERCAPTOETHANOL, fluorescent protein FP538 | | Authors: | Remington, S.J, Wachter, R.M, Yarbrough, D.K, Branchaud, B, Anderson, D.C, Kallio, K, Lukyanov, K.A. | | Deposit date: | 2004-08-25 | | Release date: | 2005-02-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | zFP538, a yellow-fluorescent protein from Zoanthus, contains a novel three-ring chromophore.
Biochemistry, 44, 2005
|
|
1Z1Q
 | |
1Z1P
 
 | |
1G7K
 
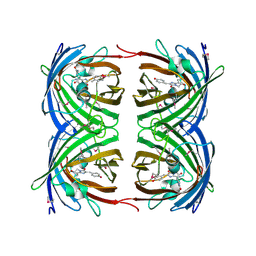 | | CRYSTAL STRUCTURE OF DSRED, A RED FLUORESCENT PROTEIN FROM DISCOSOMA SP. RED | | Descriptor: | FLUORESCENT PROTEIN FP583 | | Authors: | Yarbrough, D, Wachter, R.M, Kallio, K, Matz, M.V, Remington, S.J. | | Deposit date: | 2000-11-10 | | Release date: | 2000-12-06 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Refined crystal structure of DsRed, a red fluorescent protein from coral, at 2.0-A resolution.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
4DXN
 
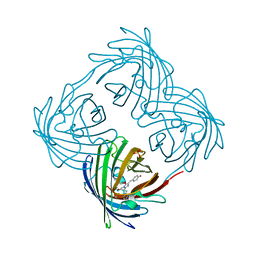 | |
4DXM
 | |
4DXO
 | |
4DXQ
 
 | |
4DXI
 | |
4DXP
 
 | |
