1YG2
 
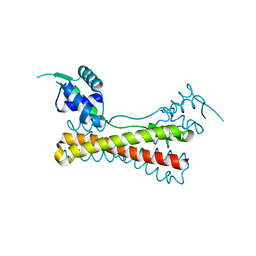 | | Structure of the Vibrio cholerae virulence activator AphA | | Descriptor: | gene activator AphA | | Authors: | De Silva, R.S, Kovacikova, G, Lin, W, Taylor, R.T, Skorupski, K, Kull, F.J. | | Deposit date: | 2005-01-04 | | Release date: | 2005-01-11 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the virulence gene activator AphA from Vibrio cholerae reveals it is a novel member of the winged helix transcription factor superfamily
J.Biol.Chem., 280, 2005
|
|
3T1B
 
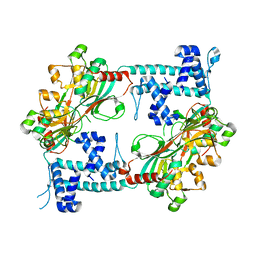 | | Crystal structure of the full-length AphB N100E variant | | Descriptor: | Transcriptional regulator, LysR family | | Authors: | Taylor, J.L, De Silva, R.S, Kovacikova, G, Lin, W, Taylor, R.K, Skorupski, K, Kull, F.J. | | Deposit date: | 2011-07-21 | | Release date: | 2011-11-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The crystal structure of AphB, a virulence gene activator from Vibrio cholerae, reveals residues that influence its response to oxygen and pH.
Mol.Microbiol., 83, 2012
|
|
3SZP
 
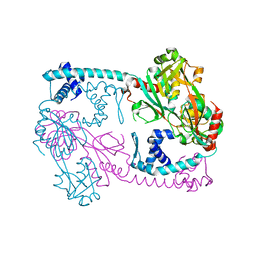 | | Full-length structure of the Vibrio cholerae virulence activator, AphB, a member of the LTTR protein family | | Descriptor: | Transcriptional regulator, LysR family | | Authors: | Taylor, J.L, De Silva, R.S, Kovacikova, G, Lin, W, Taylor, R.K, Skorupski, K, Kull, F.J. | | Deposit date: | 2011-07-19 | | Release date: | 2011-11-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.202 Å) | | Cite: | The crystal structure of AphB, a virulence gene activator from Vibrio cholerae, reveals residues that influence its response to oxygen and pH.
Mol.Microbiol., 83, 2012
|
|
6QGM
 
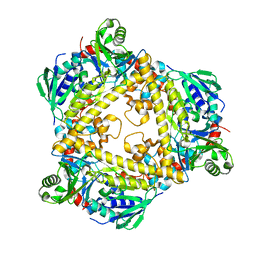 | | VirX1 apo structure | | Descriptor: | VirX1 | | Authors: | Gkotsi, D.S, Ludewig, H, Sharma, S.V, Unsworth, W.P, Taylor, R.J.K, McLachlan, M.M.W, Shanahan, S, Naismith, J.H, Goss, R.J.M. | | Deposit date: | 2019-01-11 | | Release date: | 2019-10-30 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | A marine viral halogenase that iodinates diverse substrates.
Nat.Chem., 11, 2019
|
|
2PBX
 
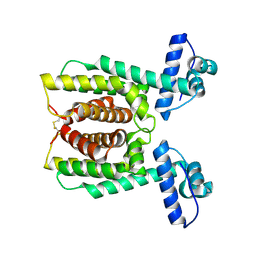 | | Vibrio cholerae HapR | | Descriptor: | Hemagglutinin/protease regulatory protein | | Authors: | Kull, F.J, DeSilva, R.S, Kovacikova, G, Lin, W, Taylor, R.K, Skorupski, K. | | Deposit date: | 2007-03-29 | | Release date: | 2007-10-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the Vibrio cholerae Quorum-Sensing
Regulatory Protein HapR
J.BACTERIOL., 189, 2007
|
|
3J1A
 
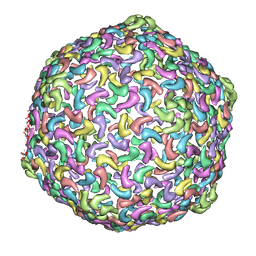 | | HK97-like fold fitted into 3D reconstruction of bacteriophage CW02 | | Descriptor: | capsid protein | | Authors: | Shen, P.S, Domek, M.J, Sanz-Garcia, E, Makaju, A, Taylor, R, Culumber, M, Breakwell, D.P, Prince, J.T, Belnap, D.M. | | Deposit date: | 2012-01-31 | | Release date: | 2012-05-30 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (16 Å) | | Cite: | Sequence and Structural Characterization of Great Salt Lake Bacteriophage CW02, a Member of the T7-Like Supergroup.
J.Virol., 86, 2012
|
|
5F72
 
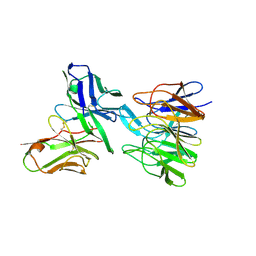 | | De novo design and crystallographic validation of antibodies targeting a pre-selected epitope | | Descriptor: | Kelch-like ECH-associated protein 1, Single chain Fv from a Fab | | Authors: | Liu, X, Taylor, R.D, Griffin, L, Coker, S, Adams, R, Ceska, T, Shi, J, Lawson, A.D.G, Baker, T. | | Deposit date: | 2015-12-07 | | Release date: | 2016-12-14 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | De novo design and crystallographic validation of antibodies targeting a pre-selected epitope
To Be Published
|
|
2KH2
 
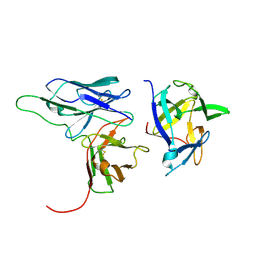 | | Solution structure of a scFv-IL-1B complex | | Descriptor: | Interleukin-1 beta, scFv | | Authors: | Wilkinson, I.C, Hall, C.J, Veverka, V, Muskett, F.W, Stephens, P.E, Taylor, R.J, Henry, A.J, Carr, M.D. | | Deposit date: | 2009-03-23 | | Release date: | 2009-09-08 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | High resolution NMR-based model for the structure of a scFv-IL-1beta complex: potential for NMR as a key tool in therapeutic antibody design and development.
J.Biol.Chem., 284, 2009
|
|
2K8P
 | | Characterisation of the structural features and interactions of sclerostin: molecular insight into a key regulator of Wnt-mediated bone formation | | Descriptor: | Sclerostin | | Authors: | Veverka, V, Henry, A.J, Slocombe, P.M, Ventom, A, Mulloy, B, Muskett, F.W, Muzylak, M, Greenslade, K, Moore, A, Zhang, L, Gong, J, Qian, X, Paszty, C, Taylor, R.J, Robinson, M.K, Carr, M.D. | | Deposit date: | 2008-09-18 | | Release date: | 2009-02-17 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Characterization of the Structural Features and Interactions of Sclerostin: Molecular insight into a key regulator of Wnt-mediated bone formation
J.Biol.Chem., 284, 2009
|
|
2L3Y
 
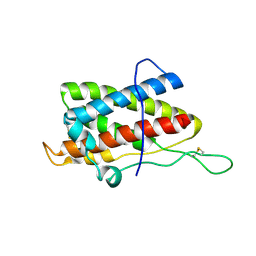 | | Solution structure of mouse IL-6 | | Descriptor: | Interleukin-6 | | Authors: | Veverka, V, Redpath, N.T, Carrington, B, Muskett, F.W, Taylor, R.J, Henry, A.J, Carr, M.D. | | Deposit date: | 2010-09-25 | | Release date: | 2011-09-28 | | Last modified: | 2013-01-02 | | Method: | SOLUTION NMR | | Cite: | Conservation of functional sites on interleukin-6 and implications for evolution of signaling complex assembly and therapeutic intervention.
J.Biol.Chem., 287, 2012
|
|
2NPU
 
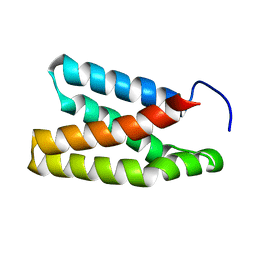 | | The solution structure of the rapamycin-binding domain of mTOR (FRB) | | Descriptor: | FKBP12-rapamycin complex-associated protein | | Authors: | Veverka, V, Crabbe, T, Bird, I, Lennie, G, Muskett, F.W, Taylor, R.J, Carr, M.D. | | Deposit date: | 2006-10-30 | | Release date: | 2007-09-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of the interaction of mTOR with phosphatidic acid and a novel class of inhibitor: compelling evidence for a central role of the FRB domain in small molecule-mediated regulation of mTOR.
Oncogene, 27, 2008
|
|
1RD4
 
 | | An allosteric inhibitor of LFA-1 bound to its I-domain | | Descriptor: | 1-ACETYL-4-(4-{4-[(2-ETHOXYPHENYL)THIO]-3-NITROPHENYL}PYRIDIN-2-YL)PIPERAZINE, Integrin alpha-L | | Authors: | Crump, M.P, Ceska, T.A, Spyracopoulos, L, Henry, A, Archibald, S.C, Alexander, R, Taylor, R.J, Findlow, S.C, O'Connell, J, Robinson, M.K, Shock, A. | | Deposit date: | 2003-11-05 | | Release date: | 2004-03-30 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of an allosteric inhibitor of LFA-1 bound to the I-domain studied by crystallography, NMR, and calorimetry
Biochemistry, 43, 2004
|
|
1JQA
 
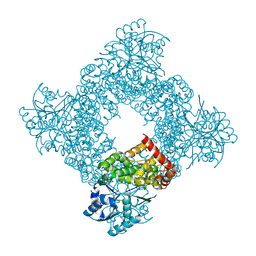 | | Bacillus stearothermophilus glycerol dehydrogenase complex with glycerol | | Descriptor: | GLYCEROL, Glycerol Dehydrogenase, ZINC ION | | Authors: | Ruzheinikov, S.N, Burke, J, Sedelnikova, S, Baker, P.J, Taylor, R, Bullough, P.A, Muir, N.M, Gore, M.G, Rice, D.W. | | Deposit date: | 2001-08-04 | | Release date: | 2001-10-03 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Glycerol dehydrogenase. structure, specificity, and mechanism of a family III polyol dehydrogenase.
Structure, 9, 2001
|
|
1JPU
 
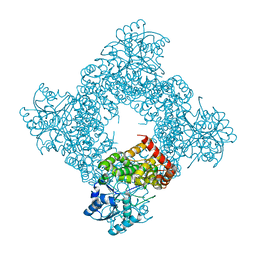 | | Crystal Structure of Bacillus Stearothermophilus Glycerol Dehydrogenase | | Descriptor: | ZINC ION, glycerol dehydrogenase | | Authors: | Ruzheinikov, S.N, Burke, J, Sedelnikova, S, Baker, P.J, Taylor, R, Bullough, P.A, Muir, N.M, Gore, M.G, Rice, D.W. | | Deposit date: | 2001-08-03 | | Release date: | 2001-10-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Glycerol dehydrogenase. structure, specificity, and mechanism of a family III polyol dehydrogenase.
Structure, 9, 2001
|
|
1JQ5
 
 | | Bacillus Stearothermophilus Glycerol dehydrogenase complex with NAD+ | | Descriptor: | Glycerol dehydrogenase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ZINC ION | | Authors: | Ruzheinikov, S.N, Burke, J, Sedelnikova, S, Baker, P.J, Taylor, R, Bullough, P.A, Muir, N.M, Gore, M.G, Rice, D.W. | | Deposit date: | 2001-08-03 | | Release date: | 2001-10-01 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Glycerol dehydrogenase. structure, specificity, and mechanism of a family III polyol dehydrogenase.
Structure, 9, 2001
|
|
