3E2Q
 
 | |
3E2S
 
 | | Crystal Structure Reduced PutA86-630 Mutant Y540S Complexed with L-proline | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, PENTAETHYLENE GLYCOL, PROLINE, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2008-08-06 | | Release date: | 2009-02-03 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A conserved active site tyrosine residue of proline dehydrogenase helps enforce the preference for proline over hydroxyproline as the substrate.
Biochemistry, 48, 2009
|
|
3E2R
 
 | |
3ET4
 
 | | Structure of Recombinant Haemophilus Influenzae E(P4) Acid Phosphatase | | Descriptor: | MAGNESIUM ION, Outer membrane protein P4, NADP phosphatase, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2008-10-06 | | Release date: | 2008-10-14 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of Recombinant Haemophilus Influenzae E (P4) Acid Phosphatase
Reveals a New Member of the Haloacid Dehalogenase Superfamily.
Biochemistry, 46, 2007
|
|
3ET5
 
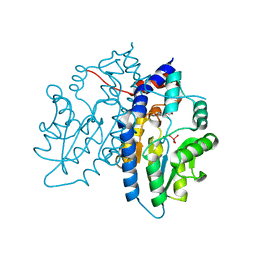 | |
3FST
 
 | |
3FSU
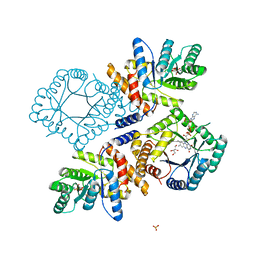
 | | Crystal Structure of Escherichia coli Methylenetetrahydrofolate Reductase Double Mutant Phe223LeuGlu28Gln complexed with methyltetrahydrofolate | | Descriptor: | 5,10-methylenetetrahydrofolate reductase, 5-METHYL-5,6,7,8-TETRAHYDROFOLIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2009-01-12 | | Release date: | 2009-08-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Functional role for the conformationally mobile phenylalanine 223 in the reaction of methylenetetrahydrofolate reductase from Escherichia coli.
Biochemistry, 48, 2009
|
|
3HAZ
 
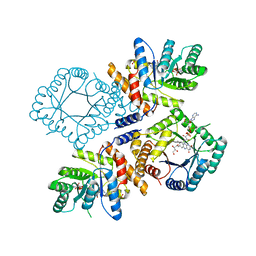
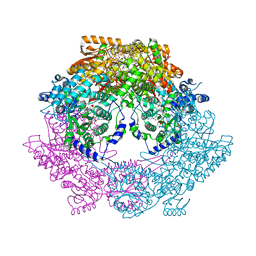 | | Crystal structure of bifunctional proline utilization A (PutA) protein | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2009-05-03 | | Release date: | 2010-02-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of the bifunctional proline utilization A flavoenzyme from Bradyrhizobium japonicum
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
8T8L
 | |
8T8K
 
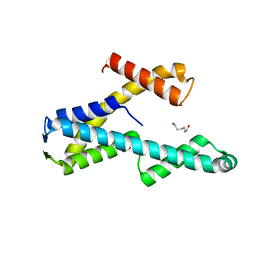 | |
8TCW
 
 | |
8TCX
 
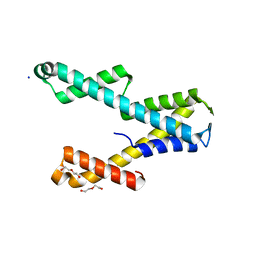
 | | Structure of PYCR1 complexed with 2,4-dioxo-1,2,3,4-tetrahydroquinazoline-6-carboxylic acid | | Descriptor: | 2,4-dioxo-1,2,3,4-tetrahydroquinazoline-6-carboxylic acid, Pyrroline-5-carboxylate reductase 1, mitochondrial, ... | | Authors: | Tanner, J.J, Meeks, K.R. | | Deposit date: | 2023-07-02 | | Release date: | 2024-03-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Novel Fragment Inhibitors of PYCR1 from Docking-Guided X-ray Crystallography.
J.Chem.Inf.Model., 64, 2024
|
|
8TCY
 
 | | Structure of PYCR1 complexed with 7-fluoro-2-oxo-1,2,3,4-tetrahydroquinoline-6-carboxylic acid | | Descriptor: | 7-fluoro-2-oxo-1,2,3,4-tetrahydroquinoline-6-carboxylic acid, DI(HYDROXYETHYL)ETHER, Pyrroline-5-carboxylate reductase 1, ... | | Authors: | Tanner, J.J, Meeks, K.R. | | Deposit date: | 2023-07-02 | | Release date: | 2024-03-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Novel Fragment Inhibitors of PYCR1 from Docking-Guided X-ray Crystallography.
J.Chem.Inf.Model., 64, 2024
|
|
8TD0
 
 | | Structure of PYCR1 complexed with 5-oxo-7a-phenyl-hexahydropyrrolo[2,1-b][1,3]thiazole-3-carboxylic acid | | Descriptor: | (3R,4S,7aR)-5-oxo-7a-phenylhexahydropyrrolo[2,1-b][1,3]thiazole-3-carboxylic acid, Pyrroline-5-carboxylate reductase 1, mitochondrial, ... | | Authors: | Tanner, J.J, Meeks, K.R. | | Deposit date: | 2023-07-02 | | Release date: | 2024-03-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Novel Fragment Inhibitors of PYCR1 from Docking-Guided X-ray Crystallography.
J.Chem.Inf.Model., 64, 2024
|
|
8TD1
 
 | |
8TCZ
 
 | |
8TCU
 
 | |
8TCV
 
 | | Structure of PYCR1 complexed with 4-bromobenzene-1,3-dicarboxylic acid | | Descriptor: | 4-bromobenzene-1,3-dicarboxylic acid, Pyrroline-5-carboxylate reductase 1, mitochondrial, ... | | Authors: | Tanner, J.J, Meeks, K.R. | | Deposit date: | 2023-07-02 | | Release date: | 2024-03-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Novel Fragment Inhibitors of PYCR1 from Docking-Guided X-ray Crystallography.
J.Chem.Inf.Model., 64, 2024
|
|
2FR4
 
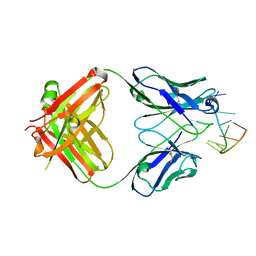 | | Structure of Fab DNA-1 complexed with a stem-loop DNA ligand | | Descriptor: | 5'-D(*CP*TP*GP*CP*CP*TP*TP*CP*AP*G)-3', TETRAETHYLENE GLYCOL, antibody heavy chain FAB, ... | | Authors: | Tanner, J.J, Ou, Z. | | Deposit date: | 2006-01-18 | | Release date: | 2007-01-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Impact of DNA hairpin folding energetics on antibody-ssDNA association.
J.Mol.Biol., 374, 2007
|
|
2FZN
 
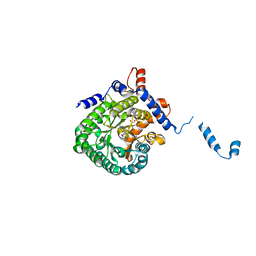 | |
2FZM
 
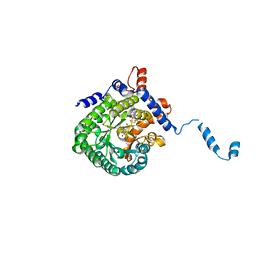 | |
2G37
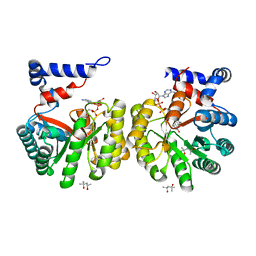 
 | | Structure of Thermus thermophilus L-proline dehydrogenase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, FLAVIN-ADENINE DINUCLEOTIDE, proline dehydrogenase/delta-1-pyrroline-5-carboxylate dehydrogenase | | Authors: | Tanner, J.J, White, T.A. | | Deposit date: | 2006-02-17 | | Release date: | 2007-02-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and Kinetics of Monofunctional Proline Dehydrogenase from Thermus thermophilus.
J.Biol.Chem., 282, 2007
|
|
3SME
 
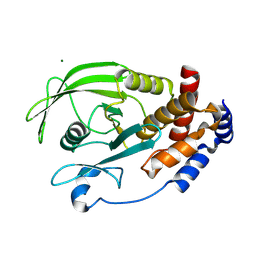 | | Structure of PTP1B inactivated by H2O2/bicarbonate | | Descriptor: | MAGNESIUM ION, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Tanner, J.J, Singh, H. | | Deposit date: | 2011-06-27 | | Release date: | 2011-10-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Biological Buffer Bicarbonate/CO(2) Potentiates H(2)O(2)-Mediated Inactivation of Protein Tyrosine Phosphatases.
J.Am.Chem.Soc., 133, 2011
|
|
1XVJ
 
 | | Crystal Structure Of Rat alpha-Parvalbumin D94S/G98E Mutant | | Descriptor: | CALCIUM ION, Parvalbumin alpha | | Authors: | Tanner, J.J, Agah, S, Lee, Y.H, Henzl, M.T. | | Deposit date: | 2004-10-28 | | Release date: | 2005-09-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of the D94S/G98E Variant of Rat alpha-Parvalbumin. An Explanation for the Reduced Divalent Ion Affinity.
Biochemistry, 44, 2005
|
|
1XKJ
 
 | | BACTERIAL LUCIFERASE BETA2 HOMODIMER | | Descriptor: | BETA2 LUCIFERASE | | Authors: | Tanner, J.J, Krause, K.L. | | Deposit date: | 1996-10-08 | | Release date: | 1997-07-07 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of bacterial luciferase beta 2 homodimer: implications for flavin binding.
Biochemistry, 36, 1997
|
|
