3I6P
 
 | |
6SYM
 
 | |
6UMT
 
 | | High-affinity human PD-1 PD-L2 complex | | Descriptor: | MAGNESIUM ION, Programmed cell death 1 ligand 2, Programmed cell death protein 1 | | Authors: | Tang, S, Kim, P.S. | | Deposit date: | 2019-10-10 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.986 Å) | | Cite: | A high-affinity human PD-1/PD-L2 complex informs avenues for small-molecule immune checkpoint drug discovery.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
3IA0
 
 | |
3I96
 
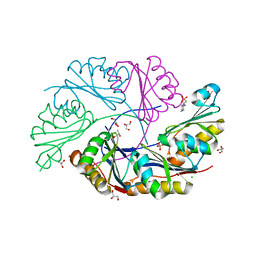 | | Ethanolamine Utilization Microcompartment Shell Subunit, EutS | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, CHLORIDE ION, Ethanolamine utilization protein eutS, ... | | Authors: | Tanaka, S, Sawaya, M.R, Yeates, T.O. | | Deposit date: | 2009-07-10 | | Release date: | 2010-01-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure and Mechanisms of a Protein-Based Organelle in Escherichia coli.
Science, 327, 2010
|
|
3I87
 
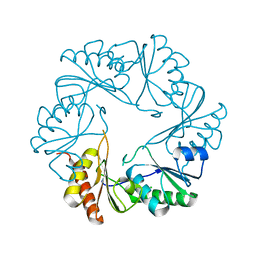 | |
6UMV
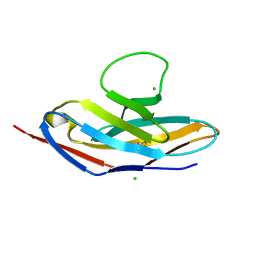 
 | | Human apo PD-1 double mutant | | Descriptor: | CHLORIDE ION, Programmed cell death protein 1 | | Authors: | Tang, S, Kim, P.S. | | Deposit date: | 2019-10-10 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.424 Å) | | Cite: | A high-affinity human PD-1/PD-L2 complex informs avenues for small-molecule immune checkpoint drug discovery.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
5GU7
 
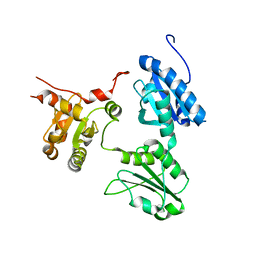 | | Crystal Structure of human ERp44 form II | | Descriptor: | Endoplasmic reticulum resident protein 44 | | Authors: | Watanabe, S, Inaba, K. | | Deposit date: | 2016-08-26 | | Release date: | 2017-04-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural basis of pH-dependent client binding by ERp44, a key regulator of protein secretion at the ER-Golgi interface
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
3I71
 
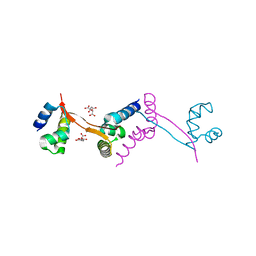 | |
3I82
 
 | |
3AHP
 
 | | Crystal structure of stable protein, CutA1, from a psychrotrophic bacterium Shewanella sp. SIB1 | | Descriptor: | CutA1 | | Authors: | Tanaka, S, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-04-26 | | Release date: | 2011-01-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of stable protein CutA1 from psychrotrophic bacterium Shewanella sp. SIB1
J.SYNCHROTRON RADIAT., 18, 2011
|
|
5FD7
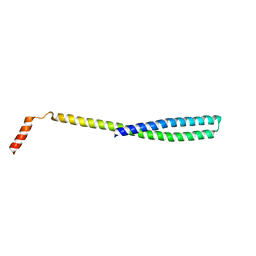 
 | | X-ray Crystal Structure of ESCRT-III Snf7 core domain (conformation A) | | Descriptor: | Vacuolar-sorting protein SNF7 | | Authors: | Tang, S, Henne, W.M, Borbat, P.P, Buchkovich, N.J, Freed, J.H, Mao, Y, Fromme, J.C, Emr, S.D. | | Deposit date: | 2015-12-15 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for activation, assembly and membrane binding of ESCRT-III Snf7 filaments.
Elife, 4, 2015
|
|
5XWM
 
 | | human ERp44 zinc-bound form | | Descriptor: | CHLORIDE ION, Endoplasmic reticulum resident protein 44, ZINC ION | | Authors: | Watanabe, S, Harayama, M, Inaba, K. | | Deposit date: | 2017-06-30 | | Release date: | 2019-01-02 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Zinc regulates ERp44-dependent protein quality control in the early secretory pathway.
Nat Commun, 10, 2019
|
|
2Z30
 
 | | Crystal structure of complex form between mat-Tk-subtilisin and Tk-propeptide | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-05-29 | | Release date: | 2007-12-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Four new crystal structures of Tk-subtilisin in unautoprocessed, autoprocessed and mature forms: insight into structural changes during maturation
J.Mol.Biol., 372, 2007
|
|
2Z2X
 
 | | Crystal structure of mature form of Tk-subtilisin | | Descriptor: | CALCIUM ION, Tk-subtilisin | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-05-29 | | Release date: | 2007-12-04 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Four new crystal structures of Tk-subtilisin in unautoprocessed, autoprocessed and mature forms: insight into structural changes during maturation
J.Mol.Biol., 372, 2007
|
|
4YNC
 
 | | OYE1 W116A COMPLEXED WITH (Z)-METHYL-3-CYANO-3-PHENYLACRYLATE IN A NON PRODUCTIVE BINDING MODE | | Descriptor: | CHLORIDE ION, FLAVIN MONONUCLEOTIDE, NADPH dehydrogenase 1, ... | | Authors: | Santangelo, S, Brenna, E, Stewart, J.D, Powell III, R.W. | | Deposit date: | 2015-03-09 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.498 Å) | | Cite: | Opposite Enantioselectivity in the Bioreduction of (Z)-beta-Aryl-beta-cyanoacrylates Mediated by the Tryptophan 116 Mutants of Old Yellow Enzyme 1: Synthetic Approach to (R)- and (S)-beta-Aryl-gamma-lactams
Adv.Synth.Catal., 357, 2015
|
|
4YIL
 
 | | OYE1 W116A COMPLEXED WITH (Z)-METHYL 3-CYANO-3-(4-FLUOROPHENYL)ACRYLATE IN A NON PRODUCTIVE BINDING MODE | | Descriptor: | FLAVIN MONONUCLEOTIDE, MAGNESIUM ION, NADPH dehydrogenase 1, ... | | Authors: | Santangelo, S, Brenna, E, Stewart, J.D, Powell III, R.W. | | Deposit date: | 2015-03-02 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.461 Å) | | Cite: | Opposite Enantioselectivity in the Bioreduction of (Z)-beta-Aryl-beta-cyanoacrylates Mediated by the Tryptophan 116 Mutants of Old Yellow Enzyme 1: Synthetic Approach to (R)- and (S)-beta-Aryl-gamma-lactams
Adv.Synth.Catal., 357, 2015
|
|
2ZRQ
 
 | | Crystal structure of S324A-subtilisin | | Descriptor: | CALCIUM ION, Tk-subtilisin | | Authors: | Tanaka, S, Takeuchi, Y, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2008-08-28 | | Release date: | 2009-03-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Crystal structure of Tk-subtilisin folded without propeptide: requirement of propeptide for acceleration of folding
Febs Lett., 582, 2008
|
|
5FD9
 
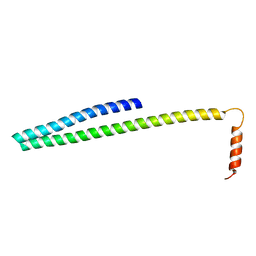 | | X-ray Crystal Structure of ESCRT-III Snf7 core domain (conformation B) | | Descriptor: | Vacuolar-sorting protein SNF7 | | Authors: | Tang, S, Henne, W.M, Borbat, P.P, Buchkovich, N.J, Freed, J.H, Mao, Y, Fromme, J.C, Emr, S.D. | | Deposit date: | 2015-12-15 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis for activation, assembly and membrane binding of ESCRT-III Snf7 filaments.
Elife, 4, 2015
|
|
3A3O
 
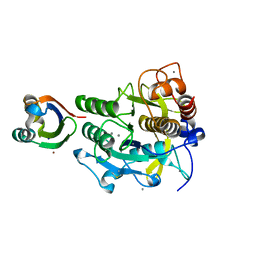 | | Crystal structure of complex between SA-subtilisin and Tk-propeptide with deletion of the five C-terminal residues | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-06-15 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of the interactions critical for propeptide-catalyzed folding of Tk-subtilisin
J.Mol.Biol., 394, 2009
|
|
3A3N
 
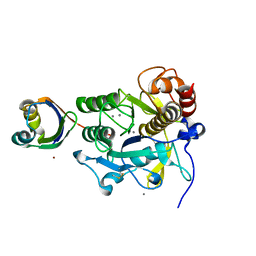 | | Crystal structure of complex between SA-subtilisin and Tk-propeptide with deletion of the two C-terminal residues | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-06-15 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Identification of the interactions critical for propeptide-catalyzed folding of Tk-subtilisin
J.Mol.Biol., 394, 2009
|
|
2Z2Z
 
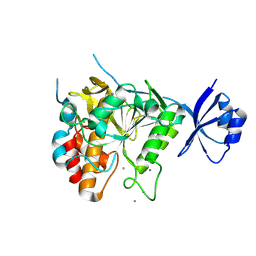 | | Crystal structure of unautoprocessed form of Tk-subtilisin soaked by 10mM CaCl2 | | Descriptor: | CALCIUM ION, Tk-subtilisin precursor | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-05-29 | | Release date: | 2007-12-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Four new crystal structures of Tk-subtilisin in unautoprocessed, autoprocessed and mature forms: insight into structural changes during maturation
J.Mol.Biol., 372, 2007
|
|
2Z2Y
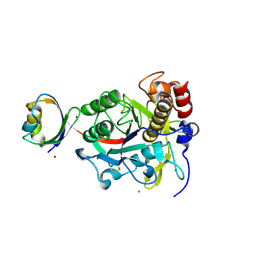 
 | | Crystal structure of autoprocessed form of Tk-subtilisin | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-05-29 | | Release date: | 2007-12-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Four new crystal structures of Tk-subtilisin in unautoprocessed, autoprocessed and mature forms: insight into structural changes during maturation
J.Mol.Biol., 372, 2007
|
|
3A3P
 
 | | Crystal structure of complex between E201A/SA-subtilisin and Tk-propeptide | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-06-15 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of the interactions critical for propeptide-catalyzed folding of Tk-subtilisin
J.Mol.Biol., 394, 2009
|
|
7JST
 
 | | Crystal structure of SARS-CoV-2 3CL in apo form | | Descriptor: | 3C-like proteinase, PHOSPHATE ION | | Authors: | Iketani, S, Forouhar, F, Liu, H, Hong, S.J, Lin, F.-Y, Nair, M.S, Zask, A, Huang, Y, Xing, L, Stockwell, B.R, Chavez, A, Ho, D.D. | | Deposit date: | 2020-08-16 | | Release date: | 2021-03-03 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Lead compounds for the development of SARS-CoV-2 3CL protease inhibitors.
Nat Commun, 12, 2021
|
|
