2W1O
 
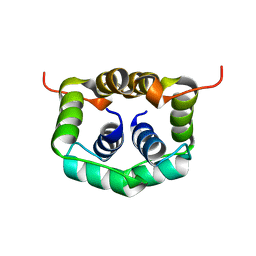 | | NMR structure of dimerization domain of human ribosomal protein P2 | | Descriptor: | 60S ACIDIC RIBOSOMAL PROTEIN P2 | | Authors: | Lee, K.M, Chan, D.S, Sze, K.H, Zhu, G, Shaw, P.C, Wong, K.B. | | Deposit date: | 2008-10-20 | | Release date: | 2009-11-17 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Dimerization Domain of Ribosomal Protein P2 Provides Insights for the Structural Organization of Eukaryotic Stalk.
Nucleic Acids Res., 38, 2010
|
|
2W4P
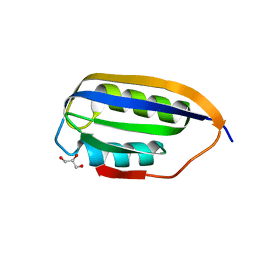 
 | | Human common-type acylphosphatase variant, A99G | | Descriptor: | ACYLPHOSPHATASE-1, GLYCEROL | | Authors: | Lam, S.Y, Sze, K.H, Wong, K.B. | | Deposit date: | 2008-11-29 | | Release date: | 2009-12-29 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A Rigidifying Salt-Bridge Favors the Activity of Thermophilic Enzyme at High Temperatures at the Expense of Low-Temperature Activity.
Plos Biol., 9, 2011
|
|
1UC6
 
 | | Solution Structure of the Carboxyl Terminal Domain of the Ciliary Neurotrophic Factor Receptor | | Descriptor: | Ciliary Neurotrophic Factor Receptor alpha | | Authors: | Man, D, He, W, Sze, K.H, Ke, G, Smith, D.K, Ip, N.Y, Zhu, G. | | Deposit date: | 2003-04-08 | | Release date: | 2004-08-10 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the C-terminal domain of the ciliary neurotrophic factor (CNTF) receptor and ligand free associations among components of the CNTF receptor complex
J.Biol.Chem., 278, 2003
|
|
5ID5
 
 | |
1XS3
 
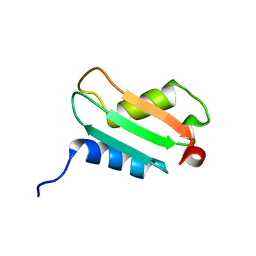 | | Solution Structure Analysis of the XC975 protein | | Descriptor: | hypothetical protein XC975 | | Authors: | Chin, K.-H, Lin, F.-Y, Hu, Y.-C, Sze, K.-H, Lyu, P.-C, Chou, S.-H. | | Deposit date: | 2004-10-18 | | Release date: | 2005-03-29 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Letter to the Editor: NMR structure note - Solution structure of a bacterial BolA-like protein XC975 from a plant pathogen Xanthomonas campestris pv. campestris
J.Biomol.Nmr, 31, 2005
|
|
6M56
 
 | |
2F5H
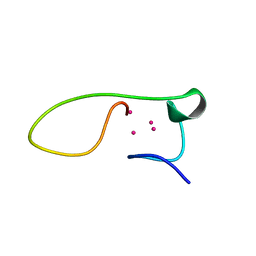 
 | | Solution structure of the alpha-domain of human Metallothionein-3 | | Descriptor: | CADMIUM ION, Metallothionein-3 | | Authors: | Wang, H, Zhang, Q, Cai, B, Li, H.Y, Sze, K.H, Huang, Z.X, Wu, H.M, Sun, H.Z. | | Deposit date: | 2005-11-25 | | Release date: | 2006-05-30 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of human metallothionein-3 (MT-3)
Febs Lett., 580, 2006
|
|
2PQG
 
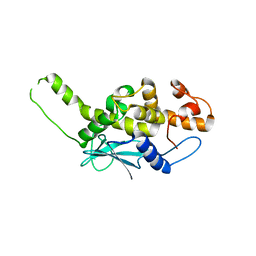 | | Crystal structure of inactive ribosome inactivating protein from maize (b-32) | | Descriptor: | Ribosome-inactivating protein 3 | | Authors: | Mak, A.N.S, Wong, Y.T, Young, J.A, Cha, S.S, Sze, K.H, Au, S.W.N, Wong, K.B, Shaw, P.C. | | Deposit date: | 2007-05-02 | | Release date: | 2008-02-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Structure-function study of maize ribosome-inactivating protein: implications for the internal inactivation region and the sole glutamate in the active site.
Nucleic Acids Res., 35, 2007
|
|
2PQI
 
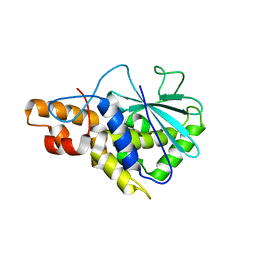 | | Crystal structure of active ribosome inactivating protein from maize (b-32) | | Descriptor: | Ribosome-inactivating protein 3 | | Authors: | Mak, A.N.S, Wong, Y.T, Young, J.A, Cha, S.S, Sze, K.H, Au, S.W.N, Wong, K.B, Shaw, P.C. | | Deposit date: | 2007-05-02 | | Release date: | 2008-02-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure-function study of maize ribosome-inactivating protein: implications for the internal inactivation region and the sole glutamate in the active site.
Nucleic Acids Res., 35, 2007
|
|
2LBF
 
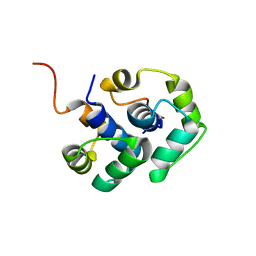 | | Solution structure of the dimerization domain of human ribosomal protein P1/P2 heterodimer | | Descriptor: | 60S acidic ribosomal protein P1, 60S acidic ribosomal protein P2 | | Authors: | Lee, K.-M, Yu, C.W.-H, Chiu, T.Y.-H, Sze, K.-H, Shaw, P.-C, Wong, K.-B. | | Deposit date: | 2011-03-30 | | Release date: | 2011-12-14 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the dimerization domain of the eukaryotic stalk P1/P2 complex reveals the structural organization of eukaryotic stalk complex
Nucleic Acids Res., 2011
|
|
2KNA
 
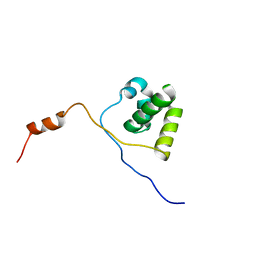 | | Solution structure of UBA domain of XIAP | | Descriptor: | Baculoviral IAP repeat-containing protein 4 | | Authors: | Hui, S.K, Tse, M.K, Sze, K.H. | | Deposit date: | 2009-08-20 | | Release date: | 2010-09-01 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Backbone and side-chain 1H, 13C and 15N assignments of the ubiquitin-associated domain of human X-linked inhibitor of apoptosis protein
Biomol.Nmr Assign., 4, 2010
|
|
2JW2
 
 | |
2L3D
 
 | |
2KSN
 
 | | Solution Structure of the N-terminal Domain of DC-UbP/UBTD2 | | Descriptor: | Ubiquitin domain-containing protein 2 | | Authors: | Song, A, Zhou, C, Guan, X, Sze, K, Hu, H. | | Deposit date: | 2010-01-08 | | Release date: | 2010-05-19 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the N-terminal domain of DC-UbP/UBTD2 and its interaction with ubiquitin
Protein Sci., 19, 2010
|
|
2L37
 
 | | 3D solution structure of arginine/glutamate-rich polypeptide Luffin P1 from the seeds of sponge gourd (Luffa cylindrical) | | Descriptor: | Ribosome-inactivating protein luffin P1 | | Authors: | Ng, Y.M, Yang, Y, Sze, K.H, Zhang, X, Zheng, Y.T, Shaw, P.C. | | Deposit date: | 2010-09-08 | | Release date: | 2011-01-19 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Structural characterization and anti-HIV-1 activities of arginine/glutamate-rich polypeptide Luffin P1 from the seeds of sponge gourd (Luffa cylindrical).
J.Struct.Biol., 2010
|
|
2KDX
 
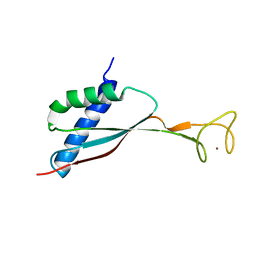 | | Solution structure of HypA protein | | Descriptor: | Hydrogenase/urease nickel incorporation protein hypA, ZINC ION | | Authors: | Xia, W, Li, H, Sze, K.-H. | | Deposit date: | 2009-01-20 | | Release date: | 2009-10-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure of a nickel chaperone, HypA, from Helicobacter pylori reveals two distinct metal binding sites
J.Am.Chem.Soc., 131, 2009
|
|
2KR7
 
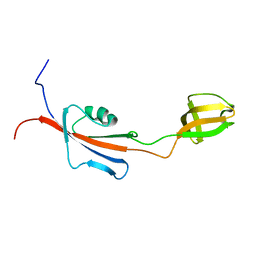 | |
2LXW
 
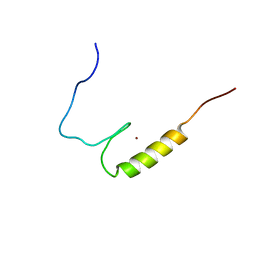 | |
2K6H
 
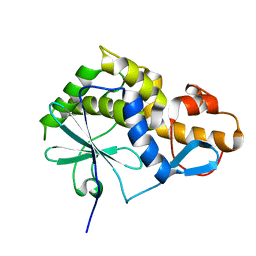 | |
4IDP
 
 | | human atlastin-1 1-446, N440T, GppNHp | | Descriptor: | Atlastin-1, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER | | Authors: | Byrnes, L.J, Singh, A, Szeto, K, Benvin, N.M, O'Donnell, J.P, Zipfel, W.R, Sondermann, H. | | Deposit date: | 2012-12-12 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.587 Å) | | Cite: | Structural basis for conformational switching and GTP loading of the large G protein atlastin.
Embo J., 32, 2013
|
|
4IDQ
 
 | | human atlastin-1 1-446, N440T, GDPAlF4- | | Descriptor: | Atlastin-1, GUANOSINE-5'-DIPHOSPHATE, HEXAETHYLENE GLYCOL, ... | | Authors: | Byrnes, L.J, Singh, A, Szeto, K, Benvin, N.M, O'Donnell, J.P, Zipfel, W.R, Sondermann, H. | | Deposit date: | 2012-12-12 | | Release date: | 2013-01-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.295 Å) | | Cite: | Structural basis for conformational switching and GTP loading of the large G protein atlastin.
Embo J., 32, 2013
|
|
4IDN
 
 | | Human atlastin-1 1-446, C-his6, GppNHp | | Descriptor: | Atlastin-1, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER | | Authors: | Byrnes, L.J, Singh, A, Szeto, K, Benvin, N.M, O'Donnell, J.P, Zipfel, W.R, Sondermann, H. | | Deposit date: | 2012-12-12 | | Release date: | 2013-01-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.252 Å) | | Cite: | Structural basis for conformational switching and GTP loading of the large G protein atlastin.
Embo J., 32, 2013
|
|
4IDO
 
 | | human atlastin-1 1-446, C-his6, GDPAlF4- | | Descriptor: | Atlastin-1, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Byrnes, L.J, Singh, A, Szeto, K, Benvin, N.M, O'Donnell, J.P, Zipfel, W.R, Sondermann, H. | | Deposit date: | 2012-12-12 | | Release date: | 2013-01-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.091 Å) | | Cite: | Structural basis for conformational switching and GTP loading of the large G protein atlastin.
Embo J., 32, 2013
|
|
3MMR
 
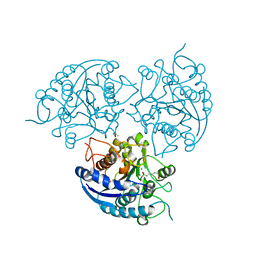 | | Structure of Plasmodium falciparum Arginase in complex with ABH | | Descriptor: | 2(S)-AMINO-6-BORONOHEXANOIC ACID, Arginase, BETA-MERCAPTOETHANOL, ... | | Authors: | Dowling, D.P, Ilies, M, Olszewski, K.L, Portugal, S, Mota, M.M, Llinas, M, Christianson, D.W. | | Deposit date: | 2010-04-20 | | Release date: | 2010-06-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Crystal structure of arginase from Plasmodium falciparum and implications for L-arginine depletion in malarial infection .
Biochemistry, 49, 2010
|
|
2MCY
 
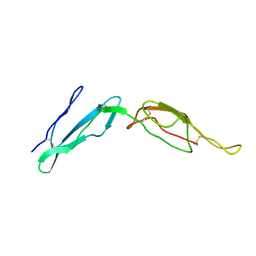 | | CR1 Sushi domains 2 and 3 | | Descriptor: | Complement receptor type 1 | | Authors: | Park, H.J, Guariento, M.J, Maciejewski, M, Hauart, R, Tham, W, Cowman, A.F, Schmidt, C.Q, Martens, H, Liszewski, K.M, Hourcade, D, Barlow, P.N, Atkinson, J.P. | | Deposit date: | 2013-08-27 | | Release date: | 2013-11-13 | | Last modified: | 2014-01-22 | | Method: | SOLUTION NMR | | Cite: | Using Mutagenesis and Structural Biology to Map the Binding Site for the Plasmodium falciparum Merozoite Protein PfRh4 on the Human Immune Adherence Receptor.
J.Biol.Chem., 289, 2014
|
|
