4IDC
 
 | | Structure of the Fragaria x ananassa enone oxidoreductase in complex with NADPH and HDMF | | Descriptor: | (2R)-4-hydroxy-2,5-dimethylfuran-3(2H)-one, 1,2-ETHANEDIOL, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2012-12-12 | | Release date: | 2013-04-17 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural basis for the enzymatic formation of the key strawberry flavor compound 4-hydroxy-2,5-dimethyl-3(2H)-furanone
J.Biol.Chem., 288, 2013
|
|
4IDD
 
 | | Structure of the Fragaria x ananassa enone oxidoreductase in complex with NADPH and EHMF | | Descriptor: | (2R)-2-ethyl-4-hydroxy-5-methylfuran-3(2H)-one, 1,2-ETHANEDIOL, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2012-12-12 | | Release date: | 2013-04-17 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis for the enzymatic formation of the key strawberry flavor compound 4-hydroxy-2,5-dimethyl-3(2H)-furanone
J.Biol.Chem., 288, 2013
|
|
4IDB
 
 | | Structure of the Fragaria x ananassa enone oxidoreductase in complex with NADP+ | | Descriptor: | 1,2-ETHANEDIOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Ripening-induced protein, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2012-12-12 | | Release date: | 2013-04-17 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural basis for the enzymatic formation of the key strawberry flavor compound 4-hydroxy-2,5-dimethyl-3(2H)-furanone
J.Biol.Chem., 288, 2013
|
|
4IDE
 
 | | Structure of the Fragaria x ananassa enone oxidoreductase in complex with NADP+ and EDHMF | | Descriptor: | (2E)-2-ethylidene-4-hydroxy-5-methylfuran-3(2H)-one, 1,2-ETHANEDIOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2012-12-12 | | Release date: | 2013-04-17 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis for the enzymatic formation of the key strawberry flavor compound 4-hydroxy-2,5-dimethyl-3(2H)-furanone
J.Biol.Chem., 288, 2013
|
|
4IDA
 
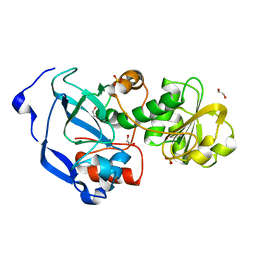 | |
4RUN
 | |
2VO9
 
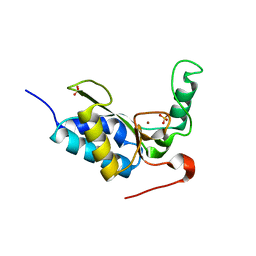 | | CRYSTAL STRUCTURE OF THE ENZYMATICALLY ACTIVE DOMAIN OF THE LISTERIA MONOCYTOGENES BACTERIOPHAGE 500 ENDOLYSIN PLY500 | | Descriptor: | L-ALANYL-D-GLUTAMATE PEPTIDASE, SULFATE ION, ZINC ION | | Authors: | Korndoerfer, I.P, Kanitz, A, Skerra, A. | | Deposit date: | 2008-02-09 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Analysis of the L-Alanoyl-D-Glutamate Endopeptidase Domain of Listeria Bacteriophage Endolysin Ply500 Reveals a New Member of the Las Peptidase Family.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
3T1W
 
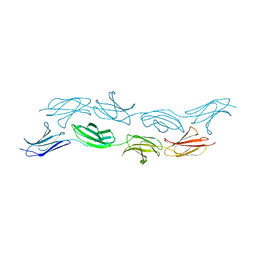 | |
6YJX
 
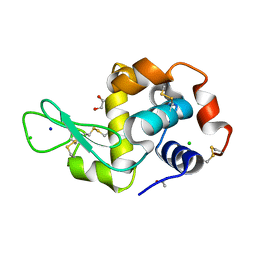 | | Structure of Hen egg-white lysozyme crystallized with PAS polypeptide | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, CHLORIDE ION, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2020-04-05 | | Release date: | 2020-07-22 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Proline/alanine-rich sequence (PAS) polypeptides as an alternative to PEG precipitants for protein crystallization.
Acta Crystallogr.,Sect.F, 76, 2020
|
|
6YJW
 | | Structure of Fragaria ananassa O-methyltransferase crystallized with PAS polypeptide | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, O-methyltransferase, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2020-04-05 | | Release date: | 2020-07-22 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Proline/alanine-rich sequence (PAS) polypeptides as an alternative to PEG precipitants for protein crystallization.
Acta Crystallogr.,Sect.F, 76, 2020
|
|
5E9O
 | | Spirochaeta thermophila X module - CBM64 - mutant G504A | | Descriptor: | 5-amino-2,4,6-triiodobenzene-1,3-dicarboxylic acid, Cellulase, glycosyl hydrolase family 5, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2015-10-15 | | Release date: | 2016-03-02 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for cellulose binding by the type A carbohydrate-binding module 64 of Spirochaeta thermophila.
Proteins, 84, 2016
|
|
5E9P
 
 | | Spirochaeta thermophila X module - CBM64 - wildtype | | Descriptor: | Cellulase, glycosyl hydrolase family 5, TPS linker, ... | | Authors: | Schiefner, A, Skerra, A. | | Deposit date: | 2015-10-15 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structural basis for cellulose binding by the type A carbohydrate-binding module 64 of Spirochaeta thermophila.
Proteins, 84, 2016
|
|
6SUA
 
 | |
3EYC
 | | New crystal structure of human tear lipocalin in complex with 1,4-butanediol in space group P21 | | Descriptor: | 1,4-BUTANEDIOL, Lipocalin-1 | | Authors: | Breustedt, D.A, Keil, L, Skerra, A. | | Deposit date: | 2008-10-20 | | Release date: | 2009-10-06 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A new crystal form of human tear lipocalin reveals high flexibility in the loop region and induced fit in the ligand cavity
Acta Crystallogr.,Sect.D, 65, 2009
|
|
1GPO
 
 | |
5BPK
 
 | | Varying binding modes of inhibitors and structural differences in the binding pockets of different gamma-glutamyltranspeptidases | | Descriptor: | (2S)-amino[(5S)-4,5-dihydro-1,2-oxazol-5-yl]acetic acid, 1,2-ETHANEDIOL, Gamma-glutamyltranspeptidase (Ggt) | | Authors: | Bolz, C, Bach, N.C, Meyer, H, Mueller, G, Dawidowski, M, Popowicz, G, Sieber, S.A, Skerra, A, Gerhard, M. | | Deposit date: | 2015-05-28 | | Release date: | 2016-05-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Varying binding modes of inhibitors and structural differences in the binding pockets of different gamma-glutamyltranspeptidases
To Be Published
|
|
1IVL
 
 | |
6I9Q
 
 | | Structure of the mouse CD98 heavy chain ectodomain | | Descriptor: | 1,2-ETHANEDIOL, 4F2 cell-surface antigen heavy chain, CHLORIDE ION | | Authors: | Schiefner, A, Deuschle, F.-C, Skerra, A. | | Deposit date: | 2018-11-24 | | Release date: | 2019-04-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural differences between the ectodomains of murine and human CD98hc.
Proteins, 87, 2019
|
|
1RST
 
 | |
1RSU
 
 | |
7O31
 
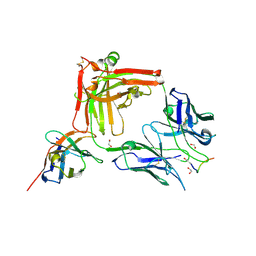 | | Crystal structure of the anti-PAS Fab 1.2 in complex with its epitope peptide and the anti-Kappa VHH domain | | Descriptor: | 1,2-ETHANEDIOL, PAS#1 epitope peptide, anti-Kappa VHH domain, ... | | Authors: | Schilz, J, Schiefner, A, Skerra, A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Molecular recognition of structurally disordered Pro/Ala-rich sequences (PAS) by antibodies involves an Ala residue at the hot spot of the epitope.
J.Mol.Biol., 433, 2021
|
|
7O30
 
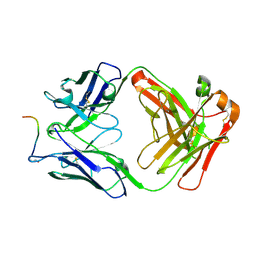 | |
7O2Z
 
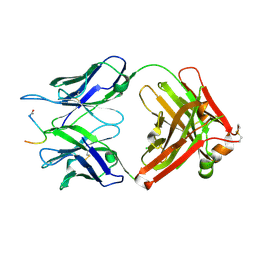 | | Crystal structure of the anti-PAS Fab 2.2 in complex with its epitope peptide | | Descriptor: | CHLORIDE ION, P/A#1 epitope peptide, anti-PAS Fab 2.2 chimeric heavy chain, ... | | Authors: | Schilz, J, Schiefner, A, Skerra, A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Molecular recognition of structurally disordered Pro/Ala-rich sequences (PAS) by antibodies involves an Ala residue at the hot spot of the epitope.
J.Mol.Biol., 433, 2021
|
|
7O33
 
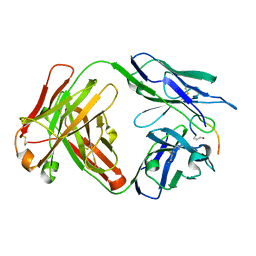 | | Crystal structure of the anti-PAS Fab 3.1 in complex with its epitope peptide | | Descriptor: | APSA epitope peptide, anti-PAS Fab 3.1 chimeric heavy chain, anti-PAS Fab 3.1 chimeric light chain | | Authors: | Schilz, J, Skerra, A. | | Deposit date: | 2021-04-01 | | Release date: | 2021-07-07 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Molecular recognition of structurally disordered Pro/Ala-rich sequences (PAS) by antibodies involves an Ala residue at the hot spot of the epitope.
J.Mol.Biol., 433, 2021
|
|
1SG2
 
 | |
