5ZBJ
 
 | |
5ZBL
 
 | |
5ZI9
 
 | | Crystal structure of type-II LOG from Streptomyces coelicolor A3 | | Descriptor: | 1,2-ETHANEDIOL, CITRATE ANION, Cytokinin riboside 5'-monophosphate phosphoribohydrolase, ... | | Authors: | Seo, H, Kim, K.-J. | | Deposit date: | 2018-03-14 | | Release date: | 2018-04-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and biochemical characterization of the type-II LOG protein from Streptomyces coelicolor A3.
Biochem. Biophys. Res. Commun., 499, 2018
|
|
7U5W
 
 | |
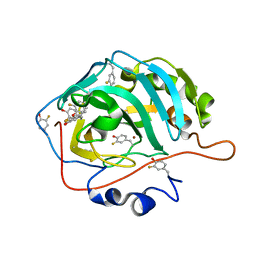
7U5X
 
 | |
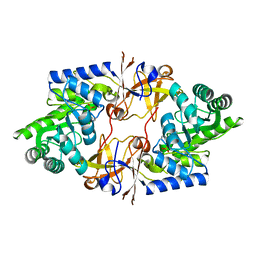
7DIB
 
 | |
5ZBK
 
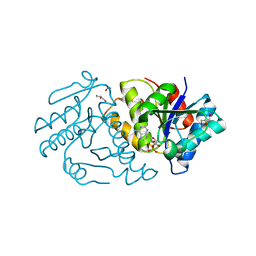 | |
7TUN
 
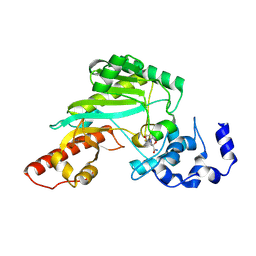 | |
7TUO
 
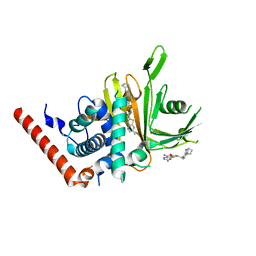 | | Crystal structure analysis of human USP28 complex with a compound | | Descriptor: | 7-amino-N-(2-{4-[(1R,3s,5S)-8-azabicyclo[3.2.1]octan-3-yl]phenyl}ethyl)-3-methylthieno[2,3-b]pyrazine-6-carboxamide, CHLORIDE ION, Ubiquitin carboxyl-terminal hydrolase 28 | | Authors: | Seo, H.-S, Dhe-Paganon, S. | | Deposit date: | 2022-02-03 | | Release date: | 2023-02-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Crystal Structure Analysis of human USP28 complex with a compound
To Be Published
|
|
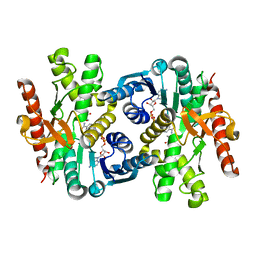
6ITK
 
 | |
6MR4
 
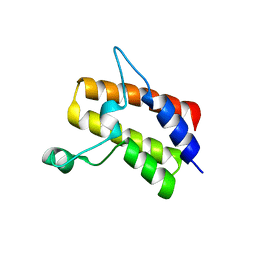 | | Crystal structure of the Sth1 bromodomain from S.cerevisiae | | Descriptor: | Nuclear protein STH1/NPS1 | | Authors: | Seo, H.S, Hashimoto, H, Krolak, A, Debler, E.W, Blus, B.J. | | Deposit date: | 2018-10-11 | | Release date: | 2019-07-24 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.71 Å) | | Cite: | Substrate Affinity and Specificity of the ScSth1p Bromodomain Are Fine-Tuned for Versatile Histone Recognition.
Structure, 27, 2019
|
|
6O34
 
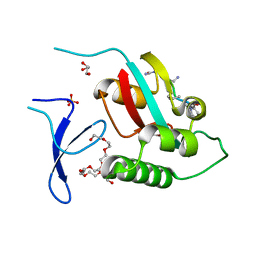 | | Crystal Structure Analysis of PIN1 | | Descriptor: | GLYCEROL, NONAETHYLENE GLYCOL, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1, ... | | Authors: | Seo, H.-S, Dhe-Paganon, S. | | Deposit date: | 2019-02-25 | | Release date: | 2020-02-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Crystal Structure Analysis of PIN1
To be Published
|
|
6O33
 
 | | Crystal Structure Analysis of PIN1 | | Descriptor: | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1, SULFATE ION, peptide | | Authors: | Seo, H.-S, Dhe-Paganon, S. | | Deposit date: | 2019-02-25 | | Release date: | 2020-02-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Crystal Structure Analysis of PIN1
To be Published
|
|
4ZC9
 
 | | Crystal Structure of the BRD4a/DB-2-190 complex | | Descriptor: | 2-[(6S)-4-(4-chlorophenyl)-2,3,9-trimethyl-6H-thieno[3,2-f][1,2,4]triazolo[4,3-a][1,4]diazepin-6-yl]-N-(4-{[({2-[(3S)-2,6-dioxopiperidin-3-yl]-1,3-dioxo-2,3-dihydro-1H-isoindol-4-yl}oxy)acetyl]amino}butyl)acetamide, Bromodomain-containing protein 4 | | Authors: | Seo, H.-S, DeAngelo, S, Bradner, J.E. | | Deposit date: | 2015-04-15 | | Release date: | 2015-11-18 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (0.99 Å) | | Cite: | DRUG DEVELOPMENT. Phthalimide conjugation as a strategy for in vivo target protein degradation.
Science, 348, 2015
|
|
7C11
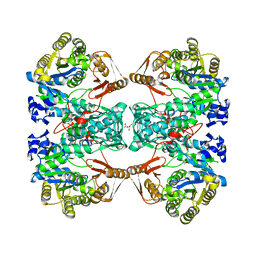 
 | | Formate--tetrahydrofolate ligase from Methylobacterium extorquens CM4 strain | | Descriptor: | ACETATE ION, CITRATE ANION, Formate-tetrahydrofolate ligase, ... | | Authors: | Kim, K.-J, Kim, S, Seo, H, Lee, S. | | Deposit date: | 2020-05-02 | | Release date: | 2020-10-28 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.815 Å) | | Cite: | Biochemical properties and crystal structure of formate-tetrahydrofolate ligase from Methylobacterium extorquens CM4.
Biochem.Biophys.Res.Commun., 528, 2020
|
|
5TX3
 
 | | Structure of Maternal Embryonic Leucine Zipper Kinase | | Descriptor: | 7-[(1S)-4-hydroxy-2,3-dihydro-1H-inden-1-yl]-5,5-dimethyl-2-({3-[(pyrrolidin-1-yl)methyl]phenyl}amino)-5,7-dihydro-6H-pyrrolo[2,3-d]pyrimidin-6-one, Maternal embryonic leucine zipper kinase | | Authors: | Li, Q, Seo, H.-S, Huang, H.-T, Gray, N.S, Dhe-Paganon, S, Eck, M.J. | | Deposit date: | 2016-11-15 | | Release date: | 2017-11-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | MELK is not necessary for the proliferation of basal-like breast cancer cells.
Elife, 6, 2017
|
|
5TWU
 
 | | Structure of Maternal Embryonic Leucine Zipper Kinase | | Descriptor: | Maternal embryonic leucine zipper kinase | | Authors: | Li, Q, Seo, H.-S, Huang, H.-T, Gray, N.S, Dhe-Paganon, S, Eck, M.J. | | Deposit date: | 2016-11-14 | | Release date: | 2017-11-22 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.603 Å) | | Cite: | MELK is not necessary for the proliferation of basal-like breast cancer cells.
Elife, 6, 2017
|
|
6UCH
 
 | | SMARCB1 nucleosome-interacting C-terminal alpha helix | | Descriptor: | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily B member 1 | | Authors: | Valencia, A.M, Sun, Z.Y.J, Seo, H.S, Vangos, H.S, Yeoh, Z.C, Mashtalir, N, Dhe-Paganon, S, Kadoch, C. | | Deposit date: | 2019-09-16 | | Release date: | 2019-11-27 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Recurrent SMARCB1 Mutations Reveal a Nucleosome Acidic Patch Interaction Site That Potentiates mSWI/SNF Complex Chromatin Remodeling.
Cell, 179, 2019
|
|
3ISO
 
 | | Crystal structure of 26 kDa GST of Clonorchis sinensis in P3221 symmetry | | Descriptor: | GLUTATHIONE, Putative glutathione transferase, SULFATE ION, ... | | Authors: | Han, Y.H, Seo, H.A, Kim, G.H, Chung, Y.J. | | Deposit date: | 2009-08-27 | | Release date: | 2010-09-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A histidine substitution confers metal binding affinity to a Schistosoma japonicum Glutathione S-transferase.
Protein Expr.Purif., 73, 2010
|
|
8HK9
 
 | |
4MBO
 
 | | 1.65 Angstrom Crystal Structure of Serine-rich Repeat Adhesion Glycoprotein (Srr1) from Streptococcus agalactiae | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, BETA-MERCAPTOETHANOL, CALCIUM ION, ... | | Authors: | Minasov, G, Shuvalova, L, Dubrovska, I, Winsor, J, Seo, H.S, Seepersaud, R, Doran, K.S, Iverson, T.M, Sullam, P.M, Anderson, W.F, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2013-08-19 | | Release date: | 2013-11-06 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Characterization of Fibrinogen Binding by Glycoproteins Srr1 and Srr2 of Streptococcus agalactiae.
J.Biol.Chem., 288, 2013
|
|
8FAL
 
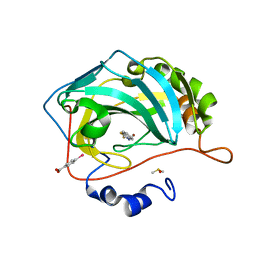 | | Masking thiol reactivity with thioamide-based MBPs- carbonic anhydrase II complexed with benzo[d]thiazole-2(3H)-thione | | Descriptor: | 1,3-benzothiazole-2(3H)-thione, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Kohlbrand, A.J, Seo, H. | | Deposit date: | 2022-11-28 | | Release date: | 2023-03-01 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | Masking thiol reactivity with thioamide, thiourea, and thiocarbamate-based MBPs.
Chem.Commun.(Camb.), 59, 2023
|
|
8FAU
 
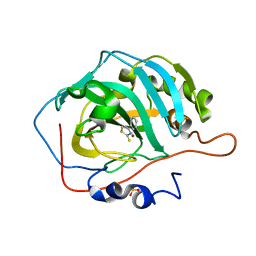 | | Masking thiol reactivity with thioamide-based MBPs- carbonic anhydrase II complexed with 4-phenylthiazole-2(3H)-thione | | Descriptor: | 4-phenyl-1,3-thiazole-2(3H)-thione, Carbonic anhydrase 2, SULFATE ION, ... | | Authors: | Kohlbrand, A.J, Seo, H. | | Deposit date: | 2022-11-28 | | Release date: | 2023-03-01 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.44 Å) | | Cite: | Masking thiol reactivity with thioamide, thiourea, and thiocarbamate-based MBPs.
Chem.Commun.(Camb.), 59, 2023
|
|
8H5J
 
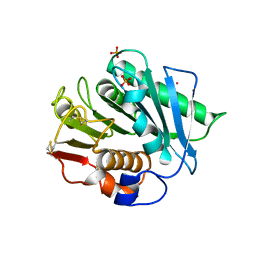 | |
8H5L
 
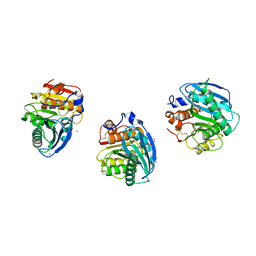 | |
