3OYZ
 
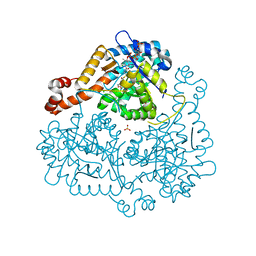 | | Haloferax volcanii Malate Synthase Pyruvate/Acetyl-CoA Ternary Complex | | Descriptor: | ACETYL COENZYME *A, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Howard, B.R, Bracken, C, Neighbor, A, Thomas, G, Lamlenn, K.K, Schubert, H.L, Whitby, F.G. | | Deposit date: | 2010-09-24 | | Release date: | 2011-06-01 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structures of a halophilic archaeal malate synthase from Haloferax volcanii and comparisons with isoforms A and G.
Bmc Struct.Biol., 11, 2011
|
|
3OYX
 
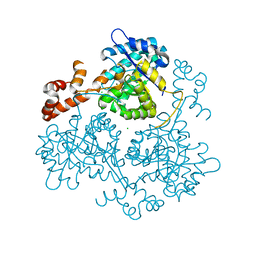 | | Haloferax volcanii Malate Synthase Magnesium/Glyoxylate Complex | | Descriptor: | CHLORIDE ION, GLYOXYLIC ACID, MAGNESIUM ION, ... | | Authors: | Howard, B.R, Bracken, C, Neighbor, A, Thomas, G, Lamlenn, K.K, Schubert, H.L, Whitby, F.G. | | Deposit date: | 2010-09-24 | | Release date: | 2011-06-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Crystal structures of a halophilic archaeal malate synthase from Haloferax volcanii and comparisons with isoforms A and G.
Bmc Struct.Biol., 11, 2011
|
|
3PUG
 
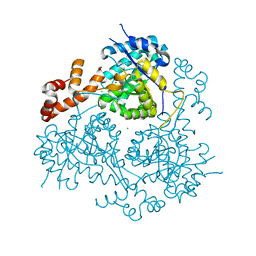 | | Haloferax volcanii Malate Synthase Native at 3mM Glyoxylate | | Descriptor: | CHLORIDE ION, GLYOXYLIC ACID, MAGNESIUM ION, ... | | Authors: | Howard, B.R, Bracken, C.D, Neighbor, A.M, Thomas, G.C, Lamlenn, K.K, Schubert, H.L, Whitby, F.G. | | Deposit date: | 2010-12-04 | | Release date: | 2011-06-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structures of a halophilic archaeal malate synthase from Haloferax volcanii and comparisons with isoforms A and G.
Bmc Struct.Biol., 11, 2011
|
|
1YTW
 
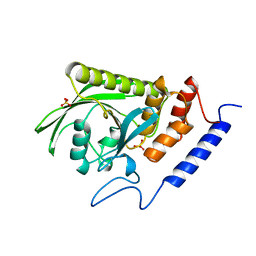 | | YERSINIA PTPASE COMPLEXED WITH TUNGSTATE | | Descriptor: | SULFATE ION, TUNGSTATE(VI)ION, YERSINIA PROTEIN TYROSINE PHOSPHATASE | | Authors: | Fauman, E.B, Schubert, H.L, Saper, M.A. | | Deposit date: | 1996-05-01 | | Release date: | 1996-11-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The X-ray crystal structures of Yersinia tyrosine phosphatase with bound tungstate and nitrate. Mechanistic implications.
J.Biol.Chem., 271, 1996
|
|
7MQS
 
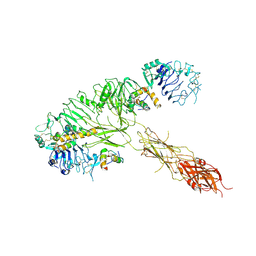 | | The insulin receptor ectodomain in complex with three venom hybrid insulin molecules - asymmetric conformation | | Descriptor: | Insulin A chain, Insulin B chain, Isoform Short of Insulin receptor | | Authors: | Blakely, A.D, Xiong, X, Kim, J.H, Menting, J, Schafer, I.B, Schubert, H.L, Agrawal, R, Gutmann, T, Delaine, C, Zhang, Y, Artik, G.O, Merriman, A, Eckert, D, Lawrence, M.C, Coskun, U, Fisher, S.J, Forbes, B.E, Safavi-Hemami, H, Hill, C.P, Chou, D.H.C. | | Deposit date: | 2021-05-06 | | Release date: | 2022-03-16 | | Last modified: | 2024-10-23 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Symmetric and asymmetric receptor conformation continuum induced by a new insulin.
Nat.Chem.Biol., 18, 2022
|
|
7MQR
 
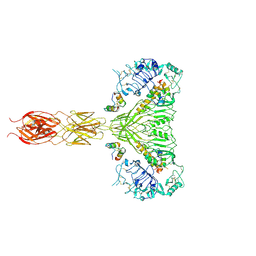 | | The insulin receptor ectodomain in complex with four venom hybrid insulins - symmetric conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Insulin A chain, Insulin B chain, ... | | Authors: | Blakely, A.D, Xiong, X, Kim, J.H, Menting, J, Schafer, I.B, Schubert, H.L, Agrawal, R, Gutmann, T, Delaine, C, Zhang, Y, Artik, G.O, Merriman, A, Eckert, D, Lawrence, M.C, Coskun, U, Fisher, S.J, Forbes, B.E, Safavi-Hemami, H, Hill, C.P, Chou, D.H.C. | | Deposit date: | 2021-05-06 | | Release date: | 2022-03-16 | | Last modified: | 2023-03-29 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Symmetric and asymmetric receptor conformation continuum induced by a new insulin.
Nat.Chem.Biol., 18, 2022
|
|
7MQO
 
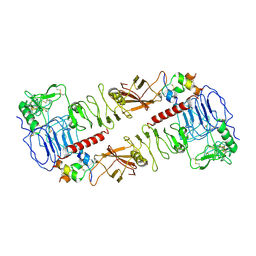 | | The insulin receptor ectodomain in complex with a venom hybrid insulin analog - "head" region | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Insulin A chain, ... | | Authors: | Blakely, A.D, Xiong, X, Kim, J.H, Menting, J, Schafer, I.B, Schubert, H.L, Agrawal, R, Gutmann, T, Delaine, C, Zhang, Y, Artik, G.O, Merriman, A, Eckert, D, Lawrence, M.C, Coskun, U, Fisher, S.J, Forbes, B.E, Safavi-Hemami, H, Hill, C.P, Chou, D.H.C. | | Deposit date: | 2021-05-06 | | Release date: | 2022-03-16 | | Last modified: | 2023-03-29 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Symmetric and asymmetric receptor conformation continuum induced by a new insulin.
Nat.Chem.Biol., 18, 2022
|
|
1XXP
 
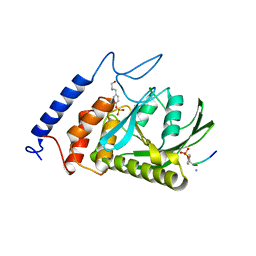 | | Yersinia YopH (residues 163-468) C403S binds phosphotyrosyl peptide at two sites | | Descriptor: | Hexapeptide ASP-ALA-ASP-GLU-PTR-CLE, Protein-tyrosine phosphatase yopH | | Authors: | Ivanov, M.I, Stuckey, J.A, Schubert, H.L, Saper, M.A, Bliska, J.B. | | Deposit date: | 2004-11-07 | | Release date: | 2005-03-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Two substrate-targeting sites in the Yersinia protein tyrosine phosphatase co-operate to promote bacterial virulence
Mol.Microbiol., 55, 2005
|
|
1XXV
 
 | | Yersinia YopH (residues 163-468) binds phosphonodifluoromethyl-Phe containing hexapeptide at two sites | | Descriptor: | Epidermal growth factor receptor derived peptide, Protein-tyrosine phosphatase yopH | | Authors: | Ivanov, M.I, Stuckey, J.A, Schubert, H.L, Saper, M.A, Bliska, J.B. | | Deposit date: | 2004-11-08 | | Release date: | 2005-03-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Two substrate-targeting sites in the Yersinia protein tyrosine phosphatase co-operate to promote bacterial virulence
Mol.Microbiol., 55, 2005
|
|
8U7T
 
 | | Substrate-bound Cdc48, Class 1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Lin, H.J.L, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-15 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
1JR2
 
 | | Structure of Uroporphyrinogen III Synthase | | Descriptor: | UROPORPHYRINOGEN-III SYNTHASE | | Authors: | Mathews, M.A, Schubert, H.L, Whitby, F.G, Alexander, K.J, Schadick, K, Bergonia, H.A, Phillips, J.D, Hill, C.P. | | Deposit date: | 2001-08-10 | | Release date: | 2001-11-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal structure of human uroporphyrinogen III synthase.
EMBO J., 20, 2001
|
|
8U9Q
 
 | | Cdc48-Shp1 unfolding native substrate, Class 6 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-19 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8U9P
 
 | | Cdc48-Shp1 unfolding native substrate, Class 2 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-19 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
1S4D
 
 | | Crystal Structure Analysis of the S-adenosyl-L-methionine dependent uroporphyrinogen-III C-methyltransferase SUMT | | Descriptor: | GLYCEROL, S-ADENOSYL-L-HOMOCYSTEINE, Uroporphyrin-III C-methyltransferase | | Authors: | Vevodova, J, Graham, R.M, Raux, E, Schubert, H.L, Roper, D.I, Brindley, A.A, Scott, A.I, Roessner, C.A, Stamford, N.P.J, Stroupe, M.E, Getzoff, E.D, Warren, M.J, Wilson, K.S. | | Deposit date: | 2004-01-16 | | Release date: | 2004-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure/Function Studies on a S-Adenosyl-l-methionine-dependent Uroporphyrinogen III C Methyltransferase (SUMT), a Key Regulatory Enzyme of Tetrapyrrole Biosynthesis
J.Mol.Biol., 344, 2004
|
|
8U8I
 | | Cdc48-Shp1 unfolding native substrate, Class 4 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-18 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8U9C
 
 | | Cdc48-Shp1 unfolding native substrate, Class 5 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-19 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.7 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8U9Z
 
 | | Cdc48-Shp1 unfolding native substrate, Class 7 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-20 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8UA1
 
 | | Cdc48-Shp1 unfolding native substrate, Class 9 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-20 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8UA0
 
 | | Cdc48-Shp1 unfolding native substrate, Class 8 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-20 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8UAA
 
 | | Cdc48-Shp1 unfolding native substrate, Class 3 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-20 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
8UB4
 
 | | Cdc48-Shp1 unfolding native substrate, consensus structure | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Cell division control protein 48, MAGNESIUM ION, ... | | Authors: | Cooney, I, Schubert, H.L, Cedeno, K, Carson, R, Fisher, O.N, Price, J.C, Hill, C.P, Shen, P.S. | | Deposit date: | 2023-09-22 | | Release date: | 2024-11-06 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Visualization of the Cdc48 AAA+ ATPase protein unfolding pathway.
Nat Commun, 15, 2024
|
|
1QNR
 
 | | The 3-D structure of a Trichoderma reesei b-mannanase from glycoside hydrolase family 5 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ENDO-1,4-B-D-MANNANASE, GLYCEROL, ... | | Authors: | Sabini, E, Schubert, H, Murshudov, G, Wilson, K.S, Siika-Aho, M, Penttila, M. | | Deposit date: | 1999-10-20 | | Release date: | 2000-10-19 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The Three-Dimensional Structure of a Trichoderma Reesei Beta-Mannanase from Glycoside Hydrolase Family 5.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1QNP
 
 | | The 3-D structure of a Trichoderma reesei b-mannanase from glycoside hydrolase family 5 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ENDO-1,4-B-D-MANNANASE, GLYCEROL, ... | | Authors: | Sabini, E, Schubert, H, Murshudov, G, Wilson, K.S, Siika-Aho, M, Penttila, M. | | Deposit date: | 1999-10-20 | | Release date: | 2000-10-19 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Three-Dimensional Structure of a Trichoderma Reesei Beta-Mannanase from Glycoside Hydrolase Family 5.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1QNQ
 
 | | The 3-D structure of a Trichoderma reesei b-mannanase from glycoside hydrolase family 5 | | Descriptor: | 2,2':6',2''-TERPYRIDINE PLATINUM(II) Chloride, 2-acetamido-2-deoxy-beta-D-glucopyranose, ENDO-1,4-B-D-MANNANASE, ... | | Authors: | Sabini, E, Schubert, H, Murshudov, G, Wilson, K.S, Siika-Aho, M, Penttila, M. | | Deposit date: | 1999-10-20 | | Release date: | 2000-10-19 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The Three-Dimensional Structure of a Trichoderma Reesei Beta-Mannanase from Glycoside Hydrolase Family 5.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1S1Q
 
 | | TSG101(UEV) domain in complex with Ubiquitin | | Descriptor: | ACETIC ACID, COPPER (II) ION, SULFATE ION, ... | | Authors: | Sundquist, W.I, Schubert, H.L, Kelly, B.N, Hill, G.C, Holton, J.M, Hill, C.P. | | Deposit date: | 2004-01-07 | | Release date: | 2004-05-04 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Ubiquitin recognition by the human TSG101 protein
Mol.Cell, 13, 2004
|
|
