4E3D
 
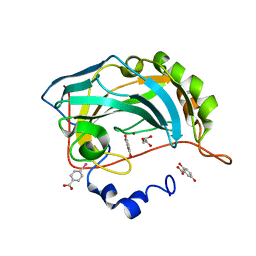 | |
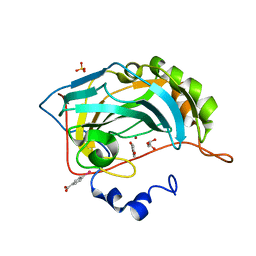
4E3G
 
 | |
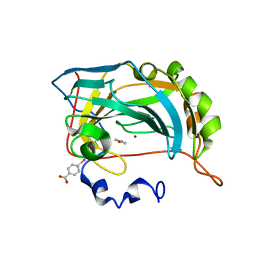
4E3H
 
 | |
1RS4
 
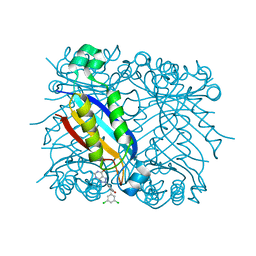 | | DHNA, 7,8-Dihydroneopterin Aldolase complexed with 3-(5-Amino-7-hydroxy-[1,2,3]triazolo[4,5-d]pyrimidin-2-yl)-N-(3,5-dichlorobenzyl)-benzamide | | Descriptor: | 3-(5-AMINO-7-HYDROXY-[1,2,3]TRIAZOLO[4,5-D]PYRIMIDIN-2-YL)-N-(3,5-DICHLOROBENZYL)-BENZAMIDE, Dihydroneopterin aldolase | | Authors: | Sanders, W.J, Nienaber, V.L, Lerner, C.G, McCall, J.O, Merrick, S.M, Swanson, S.J, Harlan, J.E, Stoll, V.S, Stamper, G.F, Betz, S.F, Condroski, K.R, Meadows, R.P, Severin, J.M, Walter, K.A, Magdalinos, P, Jakob, C.G, Wagner, R, Beutel, B.A. | | Deposit date: | 2003-12-09 | | Release date: | 2004-03-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Discovery of Potent Inhibitors of Dihydroneopterin Aldolase Using CrystaLEAD High-Throughput X-ray Crystallographic Screening and Structure-Directed Lead Optimization.
J.Med.Chem., 47, 2004
|
|
1RSI
 
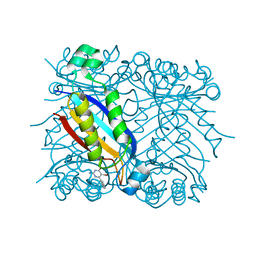 | | DHNA complex with 2-Amino-5-bromo-3-hydroxy-6-phenylpyrimidine | | Descriptor: | 2-AMINO-5-BROMO-6-PHENYLPYRIMIDIN-4-OL, Dihydroneopterin aldolase | | Authors: | Sanders, W.J, Nienaber, V.L, Lerner, C.G, McCall, J.O, Merrick, S.M, Swanson, S.J, Harlan, J.E, Stoll, V.S, Stamper, G.F, Betz, S.F, Condroski, K.R, Meadows, R.P, Severin, J.M, Walter, K.A, Magdalinos, P, Jakob, C.G, Wagner, R, Beutel, B.A. | | Deposit date: | 2003-12-09 | | Release date: | 2004-03-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Discovery of Potent Inhibitors of Dihydroneopterin Aldolase Using CrystaLEAD High-Throughput X-ray Crystallographic Screening and Structure-Directed Lead Optimization.
J.Med.Chem., 47, 2004
|
|
1R2W
 
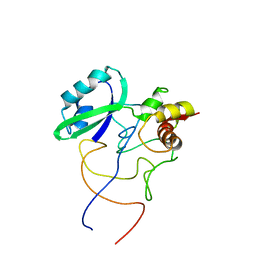 | | Coordinates of L11 with 58nts of 23S rRNA fitted into the cryo-EM map of the 70S ribosome | | Descriptor: | 50S ribosomal protein L11, 58nts of 23S rRNA | | Authors: | Valle, M, Zavialov, A, Li, W, Stagg, S.M, Sengupta, J, Nielsen, R.C, Nissen, P, Harvey, S.C, Ehrenberg, M, Frank, J. | | Deposit date: | 2003-09-30 | | Release date: | 2003-11-04 | | Last modified: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (9 Å) | | Cite: | Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron Microscopy
Nat.Struct.Biol., 10, 2003
|
|
5C9H
 
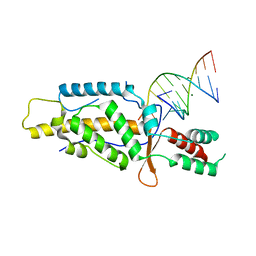 | | Structural Basis of Template Boundary Definition in Tetrahymena Telomerase | | Descriptor: | MAGNESIUM ION, RNA (5'-R(P*AP*GP*AP*AP*CP*UP*GP*UP*CP*A)-3'), RNA (5'-R(P*UP*CP*AP*UP*UP*CP*AP*GP*UP*UP*CP*U)-3'), ... | | Authors: | Jansson, L.I, Akiyama, B.M, Ooms, A, Lu, C, Rubin, S.M, Stone, M.D. | | Deposit date: | 2015-06-26 | | Release date: | 2015-10-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis of template-boundary definition in Tetrahymena telomerase.
Nat.Struct.Mol.Biol., 22, 2015
|
|
4EMK
 
 | | Crystal structure of SpLsm5/6/7 | | Descriptor: | U6 snRNA-associated Sm-like protein LSm5, U6 snRNA-associated Sm-like protein LSm6, U6 snRNA-associated Sm-like protein LSm7 | | Authors: | Jiang, S.M, Wu, D.H, Song, H.W. | | Deposit date: | 2012-04-12 | | Release date: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structures of Lsm3, Lsm4 and Lsm5/6/7 from Schizosaccharomyces pombe.
Plos One, 7, 2012
|
|
1ZKD
 
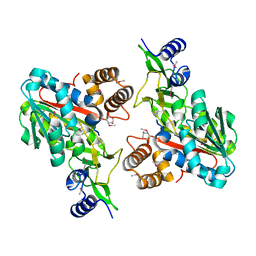 | | X-Ray structure of the putative protein Q6N1P6 from Rhodopseudomonas palustris at the resolution 2.1 A , Northeast Structural Genomics Consortium target RpR58 | | Descriptor: | DUF185 | | Authors: | Kuzin, A.P, Yong, W, Vorobiev, S.M, Acton, T, Ma, L, Xiao, R, Montelione, G, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-05-02 | | Release date: | 2005-05-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | X-Ray structure of the putative protein Q6N1P6 from Rhodopseudomonas palustris at the resolution 2.1 A , Northeast Structural Genomics Consortium target RpR58
To be Published
|
|
1RHQ
 
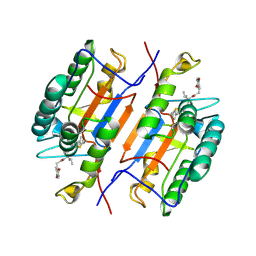 | |
1Z7W
 
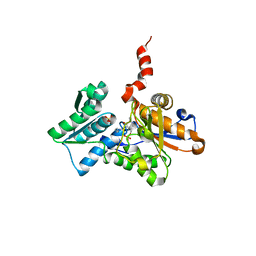 | | Crystal Structure of O-Acetylserine Sulfhydrylase from Arabidopsis thaliana | | Descriptor: | Cysteine synthase, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION | | Authors: | Bonner, E.R, Cahoon, R.E, Knapke, S.M, Jez, J.M. | | Deposit date: | 2005-03-28 | | Release date: | 2005-09-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular Basis of Cysteine Biosynthesis in Plants: STRUCTURAL AND FUNCTIONAL ANALYSIS OF O-ACETYLSERINE SULFHYDRYLASE FROM ARABIDOPSIS THALIANA.
J.Biol.Chem., 280, 2005
|
|
1ZBS
 
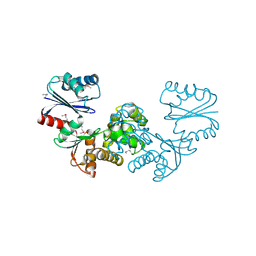 | | Crystal Structure of the Putative N-acetylglucosamine Kinase (PG1100) from Porphyromonas gingivalis, Northeast Structural Genomics Target PgR18 | | Descriptor: | hypothetical protein PG1100 | | Authors: | Forouhar, F, Abashidze, M, Kuzin, A, Vorobiev, S.M, Conover, K, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-04-08 | | Release date: | 2005-11-15 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of the Putative N-acetylglucosamine Kinase (PG1100) from Porphyromonas gingivalis, Northeast Structural Genomics Target PgR18
To be Published
|
|
4EMH
 
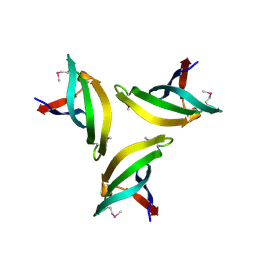 | | Crystal structure of SpLsm4 | | Descriptor: | Probable U6 snRNA-associated Sm-like protein LSm4 | | Authors: | Jiang, S.M, Wu, D.H, Song, H.W. | | Deposit date: | 2012-04-12 | | Release date: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structures of Lsm3, Lsm4 and Lsm5/6/7 from Schizosaccharomyces pombe.
Plos One, 7, 2012
|
|
1RXY
 
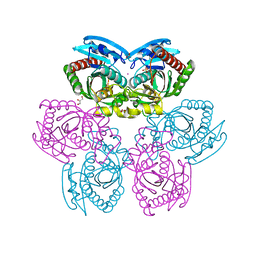 | | E. coli uridine phosphorylase: type-B native | | Descriptor: | ACETATE ION, GLYCEROL, POTASSIUM ION, ... | | Authors: | Caradoc-Davies, T.T, Cutfield, S.M, Lamont, I.L, Cutfield, J.F. | | Deposit date: | 2003-12-18 | | Release date: | 2004-04-13 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structures of escherichia coli uridine phosphorylase in two native and three complexed forms reveal basis of substrate specificity, induced conformational changes and influence of potassium
J.Mol.Biol., 337, 2004
|
|
1ZEE
 
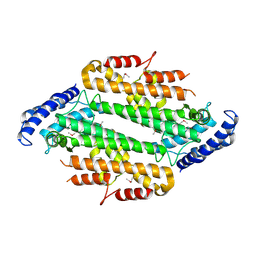 | | X-Ray Crystal Structure of Protein SO4414 from Shewanella oneidensis. Northeast Structural Genomics Consortium Target SoR52. | | Descriptor: | hypothetical protein SO4414 | | Authors: | Forouhar, F, Abashidze, M, Vorobiev, S.M, Conover, K, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2005-04-18 | | Release date: | 2005-05-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Crystal Structure of the Hypothetical Protein SO4414 from Shewanella oneidensis, NESG Target SoR52
To be Published
|
|
5CD4
 
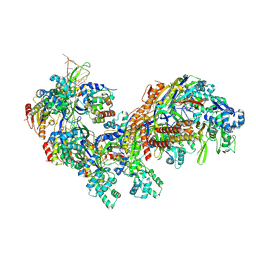 | | The Type IE CRISPR Cascade complex from E. coli, with two assemblies in the asymmetric unit arranged back-to-back | | Descriptor: | CRISPR system Cascade subunit CasA, CRISPR system Cascade subunit CasB, CRISPR system Cascade subunit CasC, ... | | Authors: | Jackson, R.N, Golden, S.M, Carter, J, Wiedenheft, B. | | Deposit date: | 2015-07-03 | | Release date: | 2015-08-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Mechanism of CRISPR-RNA guided recognition of DNA targets in Escherichia coli.
Nucleic Acids Res., 43, 2015
|
|
1S1P
 
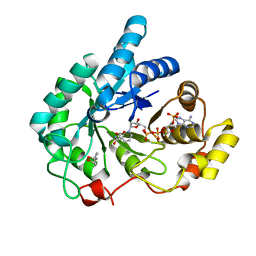 | | Crystal structures of prostaglandin D2 11-ketoreductase (AKR1C3) in complex with the non-steroidal anti-inflammatory drugs flufenamic acid and indomethacin | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ACETATE ION, Aldo-keto reductase family 1 member C3, ... | | Authors: | Lovering, A.L, Ride, J.P, Bunce, C.M, Desmond, J.C, Cummings, S.M, White, S.A. | | Deposit date: | 2004-01-07 | | Release date: | 2004-03-23 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal structures of prostaglandin D(2) 11-ketoreductase (AKR1C3) in complex with the nonsteroidal anti-inflammatory drugs flufenamic acid and indomethacin.
Cancer Res., 64, 2004
|
|
4EMG
 
 | | Crystal structure of SpLsm3 | | Descriptor: | Probable U6 snRNA-associated Sm-like protein LSm3 | | Authors: | Jiang, S.M, Wu, D.H, Song, H.W. | | Deposit date: | 2012-04-12 | | Release date: | 2012-06-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structures of Lsm3, Lsm4 and Lsm5/6/7 from Schizosaccharomyces pombe.
Plos One, 7, 2012
|
|
2C61
 
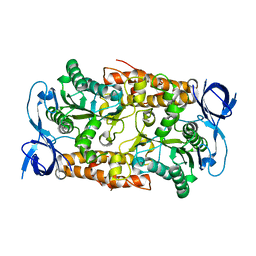 | | Crystal structure of the non-catalytic B subunit of A-type ATPase from M. mazei Go1 | | Descriptor: | A-TYPE ATP SYNTHASE NON-CATALYTIC SUBUNIT B | | Authors: | Schaefer, I, Bailer, S.M, Dueser, M, Boersch, M, Bernal, R.A, Grueber, G, Stock, D. | | Deposit date: | 2005-11-06 | | Release date: | 2006-12-11 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of the Archaeal A1Ao ATP Synthase Subunit B from Methanosarcina Mazei Go1: Implications of Nucleotide-Binding Differences in the Major A1Ao Subunits a and B. Subunits a and B
J.Mol.Biol., 358, 2006
|
|
4E49
 
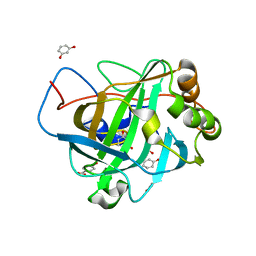 | |
2BY6
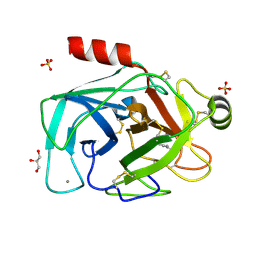 
 | | Is radiation damage dependent on the dose-rate used during macromolecular crystallography data collection | | Descriptor: | BENZAMIDINE, CALCIUM ION, CATIONIC TRYPSIN, ... | | Authors: | Leiros, H.-K.S, Timmins, J, Ravelli, R.B.G, McSweeney, S.M. | | Deposit date: | 2005-07-29 | | Release date: | 2006-02-06 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Is Radiation Damage Dependent on the Dose-Rate Used During Macromolecular Crystallography Data Collection?
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2BY7
 
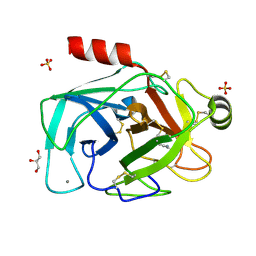 | | Is radiation damage dependent on the dose-rate used during macromolecular crystallography data collection | | Descriptor: | BENZAMIDINE, CALCIUM ION, CATIONIC TRYPSIN, ... | | Authors: | Leiros, H.-K.S, Timmins, J, Ravelli, R.B.G, McSweeney, S.M. | | Deposit date: | 2005-07-29 | | Release date: | 2006-02-06 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Is Radiation Damage Dependent on the Dose-Rate Used During Macromolecular Crystallography Data Collection?
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2BY9
 
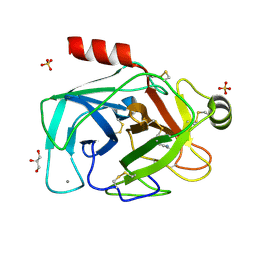 | | Is radiation damage dependent on the dose-rate used during macromolecular crystallography data collection | | Descriptor: | BENZAMIDINE, CALCIUM ION, CATIONIC TRYPSIN, ... | | Authors: | Leiros, H.-K.S, Timmins, J, Ravelli, R.B.G, McSweeney, S.M. | | Deposit date: | 2005-07-29 | | Release date: | 2006-02-06 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Is Radiation Damage Dependent on the Dose-Rate Used During Macromolecular Crystallography Data Collection?
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2BYA
 
 | | Is radiation damage dependent on the dose-rate used during macromolecular crystallography data collection | | Descriptor: | BENZAMIDINE, CALCIUM ION, CATIONIC TRYPSIN, ... | | Authors: | Leiros, H.-K.S, Timmins, J, Ravelli, R.B.G, McSweeney, S.M. | | Deposit date: | 2005-07-29 | | Release date: | 2006-02-06 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Is Radiation Damage Dependent on the Dose-Rate Used During Macromolecular Crystallography Data Collection?
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2BY5
 
 | | Is radiation damage dependent on the dose-rate used during macromolecular crystallography data collection | | Descriptor: | BENZAMIDINE, CALCIUM ION, CATIONIC TRYPSIN, ... | | Authors: | Leiros, H.-K.S, Timmins, J, Ravelli, R.B.G, McSweeney, S.M. | | Deposit date: | 2005-07-29 | | Release date: | 2006-02-06 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Is Radiation Damage Dependent on the Dose-Rate Used During Macromolecular Crystallography Data Collection?
Acta Crystallogr.,Sect.D, 62, 2006
|
|
