6Q8Q
 
 | |
6QAR
 
 | |
6RGM
 
 | | urate oxidase under 130 bar of krypton | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 8-AZAXANTHINE, ACETATE ION, ... | | Authors: | Prange, T, Colloc'h, N, Carpentier, P. | | Deposit date: | 2019-04-17 | | Release date: | 2020-05-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Comparative study of the effects of high hydrostatic pressure per se and high argon pressure on urate oxidase ligand stabilization
Acta Cryst. D, 78, 2022
|
|
3J9S
 
 | |
3JW8
 
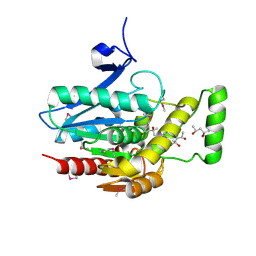 | | Crystal structure of human mono-glyceride lipase | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, MGLL protein | | Authors: | Bertrand, T, Auge, F, Houtmann, J, Rak, A, Vallee, F, Mikol, V, Berne, P.F, Michot, N, Cheuret, D, Hoornaert, C, Mathieu, M. | | Deposit date: | 2009-09-18 | | Release date: | 2010-01-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for human monoglyceride lipase inhibition.
J.Mol.Biol., 396, 2010
|
|
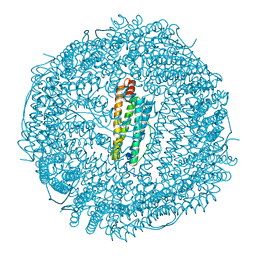
6RA8
 
 | |
5LX5
 
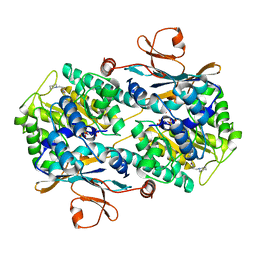 | | CRYSTAL STRUCTURE OF VISFATIN IN COMPLEX WITH SAR154782-RP. | | Descriptor: | DIPHOSPHATE, Nicotinamide phosphoribosyltransferase, [(2~{R},3~{S},4~{R},5~{R})-5-[2-azanyl-5-[[[4-[6-(ethylamino)-5-(2-piperidin-1-ylethylcarbamoyl)pyridin-2-yl]-2-fluoranyl-phenyl]carbamoylamino]methyl]pyridin-1-ium-1-yl]-3,4-bis(oxidanyl)oxolan-2-yl]methyl dihydrogen phosphate | | Authors: | Bertrand, T, Marquette, J.P. | | Deposit date: | 2016-09-20 | | Release date: | 2017-10-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | CRYSTAL STRUCTURE OF VISFATIN IN COMPLEX WITH SAR154782-RP
To Be Published
|
|
3LBG
 
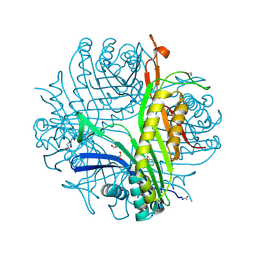 | | Urate oxidase complexed with 8-thio xanthine | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 8-thioxo-3,7,8,9-tetrahydro-1H-purine-2,6-dione, ... | | Authors: | Prange, T, Colloc'h, N, Gabison, L. | | Deposit date: | 2010-01-08 | | Release date: | 2010-06-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Near-atomic resolution structures of urate oxidase complexed with its substrate and analogues: the protonation state of the ligand.
Acta Crystallogr.,Sect.D, 66, 2010
|
|
5LX3
 
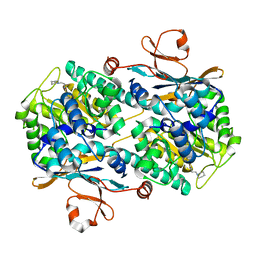 | | CRYSTAL STRUCTURE OF VISFATIN IN COMPLEX WITH SAR154782. | | Descriptor: | 6-[4-[(6-azanylpyridin-3-yl)methylcarbamoylamino]-3-fluoranyl-phenyl]-2-(ethylamino)-~{N}-(2-piperidin-1-ylethyl)pyridine-3-carboxamide, Nicotinamide phosphoribosyltransferase | | Authors: | Bertrand, T, Marquette, J.P. | | Deposit date: | 2016-09-20 | | Release date: | 2017-10-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | CRYSTAL STRUCTURE OF VISFATIN IN COMPLEX WITH SAR154782.
To Be Published
|
|
3KXU
 
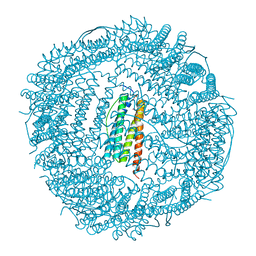 | | Crystal structure of human ferritin FTL498InsTC pathogenic mutant | | Descriptor: | CADMIUM ION, Ferritin, SULFATE ION | | Authors: | Granier, T, Gallois, B, Langlois d'Estaintot, B, Arosio, P. | | Deposit date: | 2009-12-04 | | Release date: | 2010-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Mutant ferritin L-chains that cause neurodegeneration act in a dominant-negative manner to reduce ferritin iron incorporation.
J.Biol.Chem., 285, 2010
|
|
6R1Q
 
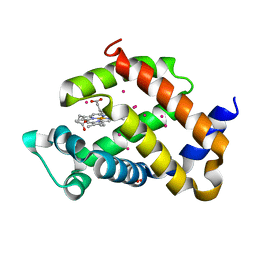 | | murine Neuroglobin under 2 kBar of argon | | Descriptor: | ARGON, CHLORIDE ION, Neuroglobin, ... | | Authors: | Prange, T, Colloc'h, N, Carpentier, P, Vallone, B. | | Deposit date: | 2019-03-14 | | Release date: | 2020-04-01 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Argon-labelling protein crystals by high pressure cooling at 2000 Bar
To Be Published
|
|
2PLB
 
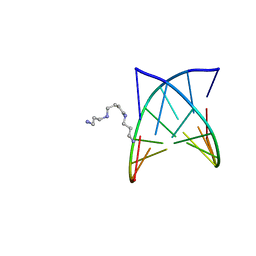 | | D(GTATACC) under hydrostatic pressure of 1.39 GPa | | Descriptor: | 5'-D(*DGP*DGP*DTP*DAP*DTP*DAP*DCP*DC)-3', SPERMINE | | Authors: | Prange, T, Girard, E, Kahn, R, Fourme, R. | | Deposit date: | 2007-04-19 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Adaptation of the base-paired double-helix molecular architecture to extreme pressure.
Nucleic Acids Res., 35, 2007
|
|
2PL4
 
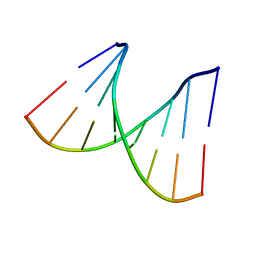 | |
2PL8
 
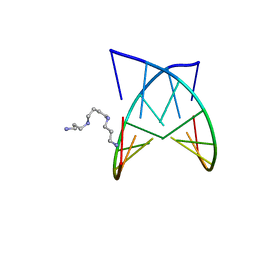 | | D(GTATACC) under hydrostatic pressure of 1.04 GPa | | Descriptor: | 5'-D(*DGP*DGP*DTP*DAP*DTP*DAP*DCP*DC)-3', SPERMINE | | Authors: | Prange, T, Girard, E, Fourme, R, Kahn, R. | | Deposit date: | 2007-04-19 | | Release date: | 2007-07-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Adaptation of the base-paired double-helix molecular architecture to extreme pressure.
Nucleic Acids Res., 35, 2007
|
|
6GDE
 
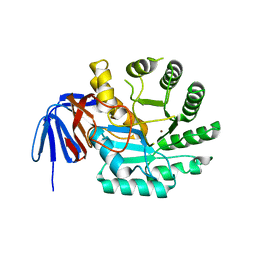 | | DIHYDROOROTASE FROM AQUIFEX AEOLICUS STANDARD (P,T) | | Descriptor: | Dihydroorotase, ZINC ION | | Authors: | Prange, T, Girard, E, Herve, G, Evans, D. | | Deposit date: | 2018-04-23 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Pressure-induced activation of latent dihydroorotase from Aquifex aeolicus as revealed by high pressure protein crystallography.
Febs J., 286, 2019
|
|
6GDD
 
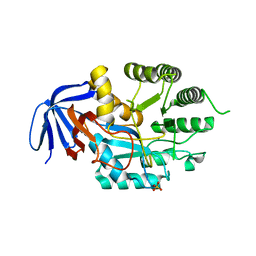 | | DIHYDROOROTASE FROM AQUIFEX AEOLICUS UNDER 1200 BAR OF HYDROSTATIC PRESSURE | | Descriptor: | Dihydroorotase, SULFATE ION, ZINC ION | | Authors: | Prange, T, Girard, E, Herve, G, Evans, D.R. | | Deposit date: | 2018-04-23 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Pressure-induced activation of latent dihydroorotase from Aquifex aeolicus as revealed by high pressure protein crystallography.
Febs J., 286, 2019
|
|
5Y97
 
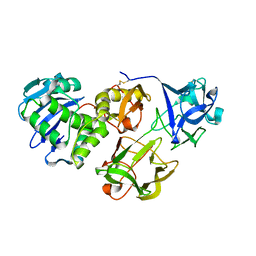 | | Crystal structure of snake gourd seed lectin in complex with lactose | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Seed lectin, beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Chandran, T, Vijayan, M, Sivaji, N, Surolia, A. | | Deposit date: | 2017-08-23 | | Release date: | 2018-08-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Ligand binding and retention in snake gourd seed lectin (SGSL). A crystallographic, thermodynamic and molecular dynamics study
Glycobiology, 28, 2018
|
|
6GDF
 
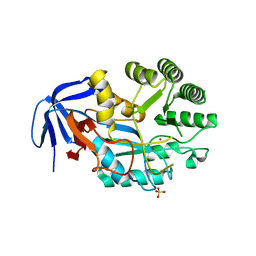 | | DIHYDROOROTASE FROM AQUIFEX AEOLICUS UNDER 600 BAR OF HYDROSTATIC PRESSURE | | Descriptor: | Dihydroorotase, SULFATE ION, ZINC ION | | Authors: | Prange, T, Girard, E, Herve, G, Evans, D.R. | | Deposit date: | 2018-04-23 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Pressure-induced activation of latent dihydroorotase from Aquifex aeolicus as revealed by high pressure protein crystallography.
Febs J., 286, 2019
|
|
2LWB
 
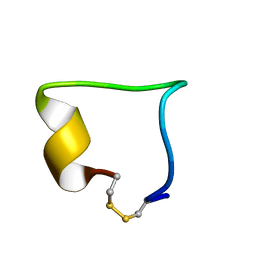 | | Structural model of BAD-1 repeat loop by NMR | | Descriptor: | Adhesin WI-1 | | Authors: | Brandhorst, T, Klein, B, Tonelli, M. | | Deposit date: | 2012-07-26 | | Release date: | 2013-07-31 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structure and function of a fungal adhesin that mimics thrombospondin-1 by binding heparin sulfate glycosaminoglycan and suppressing T cell activation via interaction with CD47
To be Published
|
|
5Y42
 
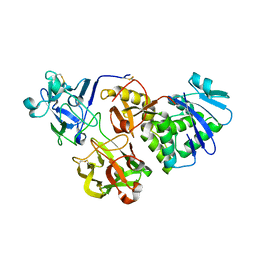 | |
2PLO
 
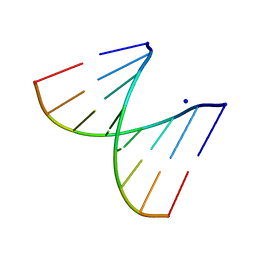 | |
1SJ2
 
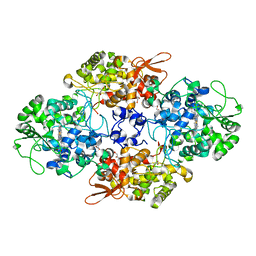 | | Crystal structure of Mycobacterium tuberculosis catalase-peroxidase | | Descriptor: | GLYCEROL, PROTOPORPHYRIN IX CONTAINING FE, Peroxidase/catalase T | | Authors: | Bertrand, T, Eady, N.A.J, Jones, J.N, Bodiguel, J, Jesmin, Nagy, J.M, Raven, E.L, Jamart-Gregoire, B, Brown, K.A. | | Deposit date: | 2004-03-02 | | Release date: | 2004-07-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Crystal Structure of Mycobacterium tuberculosis Catalase-Peroxidase.
J.Biol.Chem., 279, 2004
|
|
4ZBV
 
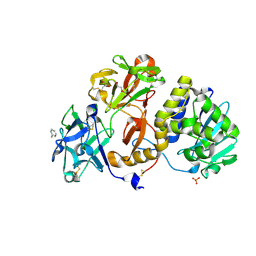 | | Structural studies on a non-toxic homologue of type II RIPs from Momordica charantia (bitter gourd) in complex with benzyl T-antigen | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-alpha-D-galactopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Chandran, T, Sharma, A, Vijayan, M. | | Deposit date: | 2015-04-15 | | Release date: | 2016-03-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural studies on a non-toxic homologue of type II RIPs from bitter gourd: Molecular basis of non-toxicity, conformational selection and glycan structure.
J.Biosci., 40, 2015
|
|
4Z8S
 
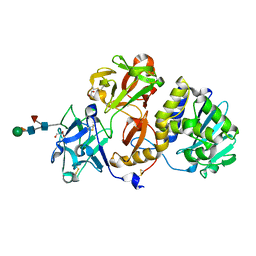 | | Structural studies on a non-toxic homologue of type II RIPs from Momordica charantia (bitter gourd)-Native-1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, ... | | Authors: | Chandran, T, Sharma, A, Vijayan, M. | | Deposit date: | 2015-04-09 | | Release date: | 2016-03-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Structural studies on a non-toxic homologue of type II RIPs from bitter gourd: Molecular basis of non-toxicity, conformational selection and glycan structure.
J.Biosci., 40, 2015
|
|
4ZLB
 
 | | Structural studies on a non-toxic homologue of type II RIPs from Momordica charantia (bitter gourd) in complex with lactose | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, alpha-L-fucopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Chandran, T, Sharma, A, Vijayan, M. | | Deposit date: | 2015-05-01 | | Release date: | 2016-03-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Structural studies on a non-toxic homologue of type II RIPs from bitter gourd: Molecular basis of non-toxicity, conformational selection and glycan structure.
J.Biosci., 40, 2015
|
|
