3M6Z
 
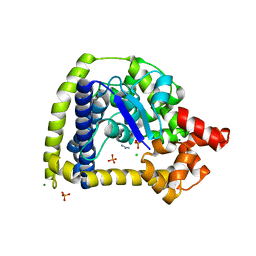 | | Crystal structure of an N-terminal 44 kDa fragment of topoisomerase V in the presence of guanidium hydrochloride | | Descriptor: | CHLORIDE ION, GUANIDINE, MAGNESIUM ION, ... | | Authors: | Rajan, R, Taneja, B, Mondragon, A. | | Deposit date: | 2010-03-16 | | Release date: | 2010-08-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structures of minimal catalytic fragments of topoisomerase v reveals conformational changes relevant for DNA binding.
Structure, 18, 2010
|
|
5HM5
 
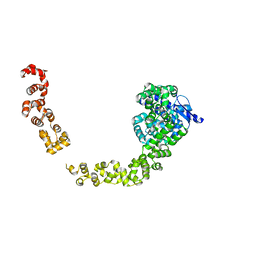 | |
4GFJ
 
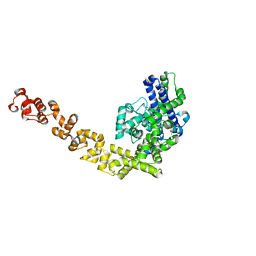 | | Crystal structure of Topo-78, an N-terminal 78kDa fragment of topoisomerase V | | Descriptor: | GLYCEROL, Topoisomerase V, ZINC ION | | Authors: | Rajan, R, Prasad, R, Taneja, B, Wilson, S.H, Mondragon, A. | | Deposit date: | 2012-08-03 | | Release date: | 2012-12-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Identification of one of the apurinic/apyrimidinic lyase active sites of topoisomerase V by structural and functional studies.
Nucleic Acids Res., 41, 2013
|
|
1YCL
 
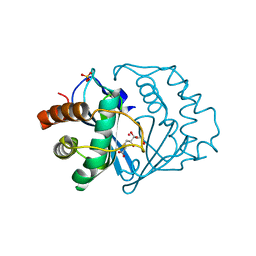 | | Crystal Structure of B. subtilis LuxS in Complex with a Catalytic 2-Ketone Intermediate | | Descriptor: | (S)-2-AMINO-4-[(2S,3R)-2,3,5-TRIHYDROXY-4-OXO-PENTYL]MERCAPTO-BUTYRIC ACID, COBALT (II) ION, S-ribosylhomocysteinase, ... | | Authors: | Rajan, R, Zhu, J, Hu, X, Pei, D, Bell, C.E. | | Deposit date: | 2004-12-22 | | Release date: | 2005-03-15 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of S-Ribosylhomocysteinase (LuxS) in Complex with a Catalytic 2-Ketone Intermediate.
Biochemistry, 44, 2005
|
|
3M7D
 
 | |
3M7G
 
 | |
3M6K
 
 | |
1RBF
 
 | |
1RBI
 
 | |
1RBC
 
 | |
1RBE
 
 | |
1RBG
 
 | |
1RBD
 
 | |
1RBH
 
 | |
1XP8
 
 | | Deinococcus radiodurans RecA in complex with ATP-gamma-S | | Descriptor: | PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, RecA protein | | Authors: | Bell, C.E, Rajan, R. | | Deposit date: | 2004-10-08 | | Release date: | 2004-12-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of RecA from Deinococcus radiodurans: insights into the structural basis of extreme radioresistance.
J.Mol.Biol., 344, 2004
|
|
2FQO
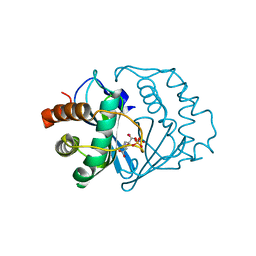 
 | | Crystal structure of B. subtilis LuxS in complex with (2S)-2-Amino-4-[(2R,3R)-2,3-dihydroxy-3-N- hydroxycarbamoyl-propylmercapto]butyric acid | | Descriptor: | (2S)-2-AMINO-4-[(2R,3R)-2,3-DIHYDROXY-3-N-HYDROXYCARBAMOYL-PROPYLMERCAPTO]BUTYRIC ACID, COBALT (II) ION, S-ribosylhomocysteine lyase, ... | | Authors: | Shen, G, Rajan, R, Zhu, J, Bell, C.E, Pei, D. | | Deposit date: | 2006-01-18 | | Release date: | 2006-05-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Design and Synthesis of Substrate and Intermediate Analogue Inhibitors of S-Ribosylhomocysteinase
J.Med.Chem., 49, 2006
|
|
2FQT
 
 | | Crystal structure of B.subtilis LuxS in complex with (2S)-2-Amino-4-[(2R,3S)-2,3-dihydroxy-3-N-hydroxycarbamoyl-propylmercapto]butyric acid | | Descriptor: | (2S)-2-AMINO-4-[(2R,3S)-2,3-DIHYDROXY-3-N-HYDROXYCARBAMOYL-PROPYLMERCAPTO]BUTYRIC ACID, COBALT (II) ION, S-ribosylhomocysteine lyase, ... | | Authors: | Shen, G, Rajan, R, Zhu, J, Bell, C.E, Pei, D. | | Deposit date: | 2006-01-18 | | Release date: | 2006-05-30 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Design and Synthesis of Substrate Analogue Inhibitors of S-Ribosylhomocysteinase (LuxS)
J.Med.Chem., 49, 2006
|
|
2RLN
 
 | |
2RNS
 
 | | REFINEMENT OF THE CRYSTAL STRUCTURE OF RIBONUCLEASE S. COMPARISON WITH AND BETWEEN THE VARIOUS RIBONUCLEASE A STRUCTURES | | Descriptor: | RIBONUCLEASE S, SULFATE ION | | Authors: | Kim, E.E, Varadarajan, R, Wyckoff, H.W, Richards, F.M. | | Deposit date: | 1992-02-19 | | Release date: | 1994-01-31 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Refinement of the crystal structure of ribonuclease S. Comparison with and between the various ribonuclease A structures.
Biochemistry, 31, 1992
|
|
7X7N
 
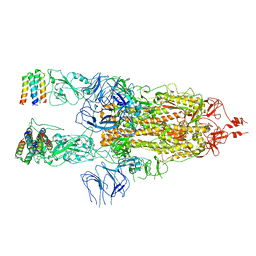 | | 3D model of the 3-RBD up single trimeric spike protein of SARS-CoV2 in the presence of synthetic peptide SIH-5. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, Synthetic peptide SIH-5 | | Authors: | Khatri, B, Pramanick, I, Malladi, S.K, Rajmani, R.S, Kumar, S, Ghosh, P, Sengupta, N, Rahisuddin, R, Kumaran, S, Ringe, R.P, Varadarajan, R, Dutta, S, Chatterjee, J. | | Deposit date: | 2022-03-10 | | Release date: | 2022-04-27 | | Last modified: | 2022-11-16 | | Method: | ELECTRON MICROSCOPY (4.47 Å) | | Cite: | A dimeric proteomimetic prevents SARS-CoV-2 infection by dimerizing the spike protein.
Nat.Chem.Biol., 18, 2022
|
|
1FEV
 
 | |
1KEB
 
 | | Crystal Structure of Double Mutant M37L,P40S E.coli Thioredoxin | | Descriptor: | COPPER (II) ION, Thioredoxin 1 | | Authors: | Rudresh, Jain, R, Dani, V, Mitra, A, Srivastava, S, Sarma, S.P, Varadarajan, R, Ramakumar, S. | | Deposit date: | 2001-11-15 | | Release date: | 2002-11-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Consequences of Replacement of an alpha-helical Pro Residue in E.coli Thioredoxin
PROTEIN ENG., 15, 2002
|
|
1SGU
 
 | | Comparing the Accumulation of Active Site and Non-active Site Mutations in the HIV-1 Protease | | Descriptor: | N-[2(R)-HYDROXY-1(S)-INDANYL]-5-[(2(S)-TERTIARY BUTYLAMINOCARBONYL)-4(3-PYRIDYLMETHYL)PIPERAZINO]-4(S)-HYDROXY-2(R)-PHENYLMETHYLPENTANAMIDE, POL polyprotein | | Authors: | Clemente, J.C, Moose, R.E, Hemrajani, R, Govindasamy, L, Reutzel, R, Mckenna, R, Abandje-McKenna, M, Goodenow, M.M, Dunn, B.M. | | Deposit date: | 2004-02-24 | | Release date: | 2004-10-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Comparing the Accumulation of Active- and Nonactive-Site Mutations in the HIV-1 Protease.
Biochemistry, 43, 2004
|
|
1SH9
 
 | | Comparing the Accumulation of Active Site and Non-active Site Mutations in the HIV-1 Protease | | Descriptor: | POL polyprotein, RITONAVIR | | Authors: | Clemente, J.C, Moose, R.E, Hemrajani, R, Govindasamy, L, Reutzel, R, McKenna, R, Abanje-McKenna, M, Goodenow, M.M, Dunn, B.M. | | Deposit date: | 2004-02-25 | | Release date: | 2004-10-05 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Comparing the Accumulation of Active- and Nonactive-Site Mutations in the HIV-1 Protease.
Biochemistry, 43, 2004
|
|
1D5D
 
 | |
