3RBV
 
 | | Crystal Structure of KijD10, a 3-ketoreductase from Actinomadura kijaniata incomplex with NADP | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Holden, H.M, Kubiak, R.L. | | Deposit date: | 2011-03-30 | | Release date: | 2011-06-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Combined Structural and Functional Investigation of a C-3''-Ketoreductase Involved in the Biosynthesis of dTDP-l-Digitoxose.
Biochemistry, 50, 2011
|
|
3R9M
 
 | | Crystal structure of the Brox Bro1 domain | | Descriptor: | 1,2-ETHANEDIOL, BRO1 domain-containing protein BROX, FORMIC ACID | | Authors: | Mu, R.L, Jiang, J.S, Snyder, G, Smith, P, Xiao, T. | | Deposit date: | 2011-03-25 | | Release date: | 2011-09-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Phe105 Loop of Alix Bro1 Domain Plays a Key Role in HIV-1 Release.
Structure, 19, 2011
|
|
3RA3
 
 | | Crystal structure of a section of a de novo design gigaDalton protein fibre | | Descriptor: | SODIUM ION, p1c, p2f | | Authors: | Zaccai, N.R, Sharp, T.H, Bruning, M, Woolfson, D.N, Brady, R.L. | | Deposit date: | 2011-03-27 | | Release date: | 2012-08-08 | | Last modified: | 2013-06-19 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Cryo-transmission electron microscopy structure of a gigadalton peptide fiber of de novo design
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1OES
 
 | | Oxidation state of protein tyrosine phosphatase 1B | | Descriptor: | MAGNESIUM ION, PROTEIN-TYROSINE PHOSPHATASE, NON-RECEPTOR TYPE 1 | | Authors: | van Montfort, R.L.M, Congreve, M, Tisi, D, Carr, R, Jhoti, H. | | Deposit date: | 2003-03-31 | | Release date: | 2003-06-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Oxidation state of the active-site cysteine in protein tyrosine phosphatase 1B.
Nature, 423, 2003
|
|
1OET
 
 | | Oxidation state of protein tyrosine phosphatase 1B | | Descriptor: | -TYROSINE PHOSPHATASE, NON-RECEPTOR TYPE 1, MAGNESIUM ION | | Authors: | van Montfort, R.L.M, Congreve, M, Tisi, D, Carr, R, Jhoti, H. | | Deposit date: | 2003-03-31 | | Release date: | 2003-06-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Oxidation state of the active-site cysteine in protein tyrosine phosphatase 1B.
Nature, 423, 2003
|
|
1Z12
 
 | | Crystal Structure of Bovine Low Molecular Weight PTPase Complexed with Vanadate | | Descriptor: | Low molecular weight phosphotyrosine protein phosphatase, VANADATE ION | | Authors: | Zhang, M, Zhou, M, Van Etten, R.L, Stauffacher, C.V. | | Deposit date: | 2005-03-03 | | Release date: | 2005-04-05 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of Bovine Low Molecular Weight Phosphotyrosyl Phosphatase Complexed with the Transition State Analog Vanadate
Biochemistry, 36, 1997
|
|
1YWG
 
 | | The structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, glyceraldehyde-3-phosphate dehydrogenase | | Authors: | Satchell, J.F, Klonis, N, Malby, R.L, Adisa, A, Alpyurek, A.E, Colman, P.M, Tilley, L. | | Deposit date: | 2005-02-17 | | Release date: | 2005-11-29 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
3RC2
 
 | | Crystal Structure of KijD10, a 3-ketoreductase from Actinomadura kijaniata in complex with TDP-benzene and NADP; open conformation | | Descriptor: | 1,2-ETHANEDIOL, 5'-O-[(S)-hydroxy{[(S)-hydroxy(phenoxy)phosphoryl]oxy}phosphoryl]thymidine, CHLORIDE ION, ... | | Authors: | Holden, H.M, Kubiak, R.L. | | Deposit date: | 2011-03-30 | | Release date: | 2011-06-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Combined Structural and Functional Investigation of a C-3''-Ketoreductase Involved in the Biosynthesis of dTDP-l-Digitoxose.
Biochemistry, 50, 2011
|
|
3RCB
 
 | |
3R8Q
 
 | |
3R9B
 
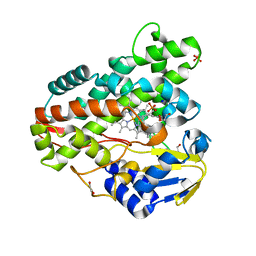 | | Crystal structure of Mycobacterium smegmatis CYP164A2 in ligand free state | | Descriptor: | 1,2-ETHANEDIOL, CYTOCHROME P450 164A2, DODECANE, ... | | Authors: | Agnew, C.R.J, Warrilow, A.G.S, Kelly, S.L, Brady, R.L. | | Deposit date: | 2011-03-25 | | Release date: | 2012-04-04 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | An enlarged, adaptable active site in CYP164 family P450 enzymes, the sole P450 in Mycobacterium leprae.
Antimicrob.Agents Chemother., 56, 2012
|
|
2BMA
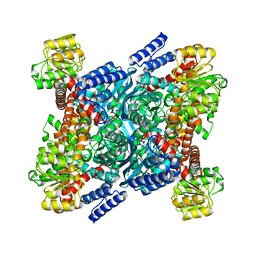 
 | | The crystal structure of Plasmodium falciparum glutamate dehydrogenase, a putative target for novel antimalarial drugs | | Descriptor: | GLUTAMATE DEHYDROGENASE (NADP+) | | Authors: | Werner, C, Stubbs, M.T, Krauth-Siege, R.L, Klebe, G. | | Deposit date: | 2005-03-10 | | Release date: | 2005-05-19 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The Crystal Structure of Plasmodium Falciparum Glutamate Dehydrogenase, a Putative Target for Novel Antimalarial Drugs
J.Mol.Biol., 349, 2005
|
|
295D
 
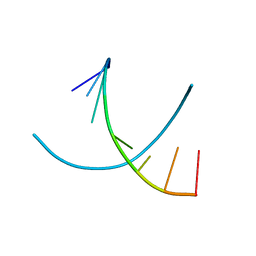 | | CRYSTAL AND SOLUTION STRUCTURES OF THE OLIGONUCLEOTIDE D(ATGCGCAT)2: A COMBINED X-RAY AND NMR STUDY | | Descriptor: | DNA (5'-D(*AP*TP*GP*CP*GP*CP*AP*T)-3') | | Authors: | Clark, G.R, Brown, D.G, Sanderson, M.R, Chwalinski, T, Neidle, S, Veal, J.M, Jones, R.L, Wilson, W.D, Zon, G, Garman, E, Stuart, D.I. | | Deposit date: | 1991-05-28 | | Release date: | 1996-12-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal and solution structures of the oligonucleotide d(ATGCGCAT)2: a combined X-ray and NMR study.
Nucleic Acids Res., 18, 1990
|
|
2A5C
 
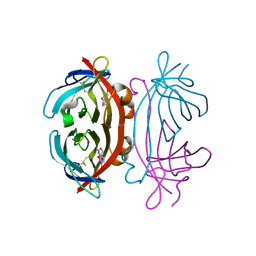 | | Structure of Avidin in complex with the ligand 8-oxodeoxyadenosine | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 8-OXODEOXYADENOSINE, Avidin | | Authors: | Conners, R, Hooley, E, Thomas, S, Brady, R.L. | | Deposit date: | 2005-06-30 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Recognition of oxidatively modified bases within the biotin-binding site of avidin.
J.Mol.Biol., 357, 2006
|
|
3R47
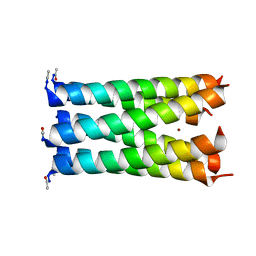 
 | | Crystal structure of a 6-helix coiled coil CC-hex-H24 | | Descriptor: | BROMIDE ION, coiled coil helix L24H | | Authors: | Zaccai, N.R, Chi, B.H.C, Woolfson, D.N, Brady, R.L. | | Deposit date: | 2011-03-17 | | Release date: | 2011-11-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.5002 Å) | | Cite: | A de novo peptide hexamer with a mutable channel.
Nat.Chem.Biol., 7, 2011
|
|
3R9C
 
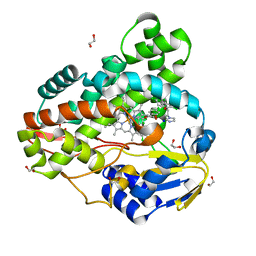 | | Crystal structure of Mycobacterium smegmatis CYP164A2 with Econazole bound | | Descriptor: | 1,2-ETHANEDIOL, 1-[(2R)-2-[(4-chlorobenzyl)oxy]-2-(2,4-dichlorophenyl)ethyl]-1H-imidazole, Cytochrome P450 164A2, ... | | Authors: | Agnew, C.R.J, Kelly, S.L, Brady, R.L. | | Deposit date: | 2011-03-25 | | Release date: | 2012-03-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | An enlarged, adaptable active site in CYP164 family P450 enzymes, the sole P450 in Mycobacterium leprae.
Antimicrob.Agents Chemother., 56, 2012
|
|
2A9X
 
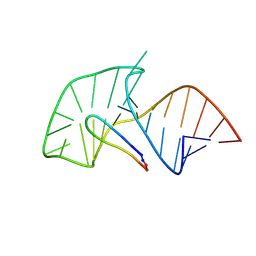 | | TAR RNA recognition by a cyclic peptidomimetic of Tat protein | | Descriptor: | BIV TAR RNA, BIV-2 cyclic peptide | | Authors: | Leeper, T.C, Athanassiou, Z, Dias, R.L, Robinson, J.A, Varani, G. | | Deposit date: | 2005-07-12 | | Release date: | 2005-11-01 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | TAR RNA recognition by a cyclic peptidomimetic of Tat protein.
Biochemistry, 44, 2005
|
|
2C8W
 
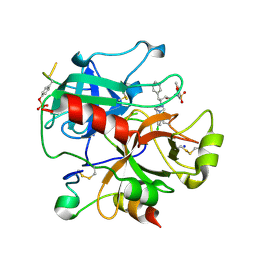 | | thrombin inhibitors | | Descriptor: | (2S,3R)-N-[5-CHLORO-2-(2,3-DIHYDRO-1H-TETRAZOL-1-YL)BENZYL]-3-HYDROXY-4-{[(4-METHOXYPHENYL)SULFONYL]AMINO}-1-PHENYLBUTA N-2-AMINIUM, HIRUDIN VARIANT-2, SODIUM ION, ... | | Authors: | Howard, N, Abell, C, Blakemore, W, Carr, R, Chessari, G, Congreve, M, Howard, S, Jhoti, H, Murray, C.W, Seavers, L.C.A, van Montfort, R.L.M. | | Deposit date: | 2005-12-08 | | Release date: | 2006-07-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Application of Fragment Screening and Fragment Linking to the Discovery of Novel Thrombin Inhibitors
J.Med.Chem., 49, 2006
|
|
1IIB
 
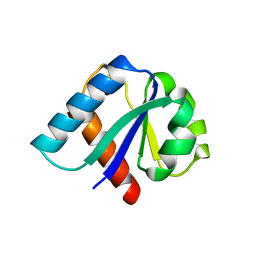 | | CRYSTAL STRUCTURE OF IIBCELLOBIOSE FROM ESCHERICHIA COLI | | Descriptor: | ENZYME IIB OF THE CELLOBIOSE-SPECIFIC PHOSPHOTRANSFERASE SYSTEM | | Authors: | Van Montfort, R.L.M, Pijning, T, Kalk, K.H, Reizer, J, Saier, M.H, Thunnissen, M.M.G.M, Robillard, G.T, Dijkstra, B.W. | | Deposit date: | 1996-12-23 | | Release date: | 1997-12-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structure of an energy-coupling protein from bacteria, IIBcellobiose, reveals similarity to eukaryotic protein tyrosine phosphatases.
Structure, 5, 1997
|
|
2AYE
 
 | |
2C8X
 
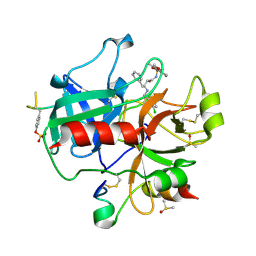 | | thrombin inhibitors | | Descriptor: | DIMETHYL SULFOXIDE, HIRUDIN VARIANT-2, N-{(2R,3S)-3-[(3-CHLOROBENZYL)AMINO]-2-HYDROXY-4-PHENYLBUTYL}-4-METHOXY-2,3,6-TRIMETHYLBENZENESULFONAMIDE, ... | | Authors: | Howard, N, Abell, C, Blakemore, W, Carr, R, Chessari, G, Congreve, M, Howard, S, Jhoti, H, Murray, C.W, Seavers, L.C.A, van Montfort, R.L.M. | | Deposit date: | 2005-12-08 | | Release date: | 2006-07-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Application of Fragment Screening and Fragment Linking to the Discovery of Novel Thrombin Inhibitors
J.Med.Chem., 49, 2006
|
|
2D1G
 
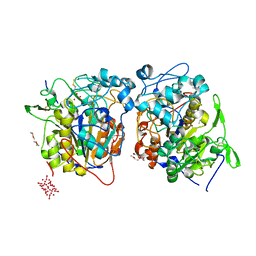 | | Structure of Francisella tularensis Acid Phosphatase A (AcpA) bound to orthovanadate | | Descriptor: | 2-ETHOXYETHANOL, 2-{2-[2-2-(METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, DECAVANADATE, ... | | Authors: | Felts, R.L, Reilly, T.J, Tanner, J.J. | | Deposit date: | 2005-08-20 | | Release date: | 2006-08-15 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of Francisella tularensis AcpA: prototype of a unique superfamily of acid phosphatases and phospholipases C
J.Biol.Chem., 281, 2006
|
|
4ZDO
 
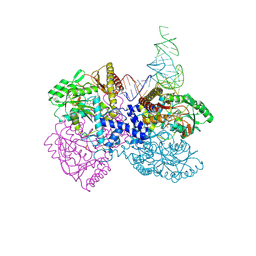 | |
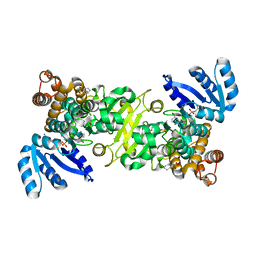
4ZQG
 
 | |
4ZDL
 
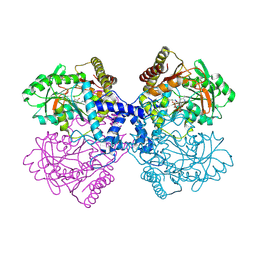 | | The crystal structure of the T325S mutant of the human holo SepSecS | | Descriptor: | (5-HYDROXY-4,6-DIMETHYLPYRIDIN-3-YL)METHYL DIHYDROGEN PHOSPHATE, CITRATE ANION, O-phosphoseryl-tRNA(Sec) selenium transferase | | Authors: | French, R.L, Simonovic, M. | | Deposit date: | 2015-04-17 | | Release date: | 2016-04-20 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Structural basis for early-onset neurological disorders caused by mutations in human selenocysteine synthase.
Sci Rep, 6, 2016
|
|
