1M61
 
 | |
1L8G
 
 | | Crystal structure of PTP1B complexed with 7-(1,1-Dioxo-1H-benzo[d]isothiazol-3-yloxymethyl)-2-(oxalyl-amino)-4,7-dihydro-5H-thieno[2,3-c]pyran-3-carboxylic acid | | Descriptor: | 7-(1,1-DIOXO-1H-BENZO[D]ISOTHIAZOL-3-YLOXYMETHYL)-2-(OXALYL-AMINO)-4,7-DIHYDRO-5H-THIENO[2,3-C]PYRAN-3-CARBOXYLIC ACID, PROTEIN-TYROSINE PHOSPHATASE, NON-RECEPTOR TYPE 1 | | Authors: | Iversen, L.F, Andersen, H.S, Moller, K.B, Olsen, O.H, Peters, G.H, Branner, S, Mortensen, S.B, Hansen, T.K, Lau, J, Ge, Y, Holsworth, D.D, Newman, M.J, Moller, N.P.H. | | Deposit date: | 2002-03-20 | | Release date: | 2002-05-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Steric hindrance as a basis for structure-based design of selective inhibitors of protein-tyrosine phosphatases.
Biochemistry, 40, 2001
|
|
1N1M
 
 | | Human Dipeptidyl Peptidase IV/CD26 in complex with an inhibitor | | Descriptor: | 2-AMINO-3-METHYL-1-PYRROLIDIN-1-YL-BUTAN-1-ONE, 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Rasmussen, H.B, Branner, S, Wiberg, F.C, Wagtmann, N.R. | | Deposit date: | 2002-10-18 | | Release date: | 2002-12-27 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of human dipeptidyl peptidase IV/CD26 in complex with a substrate analogue
Nat.Struct.Biol., 10, 2003
|
|
2HZM
 
 | | Structure of the Mediator head subcomplex Med18/20 | | Descriptor: | PHOSPHATE ION, RNA polymerase II mediator complex subunit 18, RNA polymerase II mediator complex subunit 20 | | Authors: | Lariviere, L, Geiger, S, Hoeppner, S, Rother, S, Straesser, K, Cramer, P. | | Deposit date: | 2006-08-09 | | Release date: | 2006-09-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure and TBP binding of the Mediator head subcomplex Med8-Med18-Med20.
Nat.Struct.Mol.Biol., 13, 2006
|
|
1GMC
 
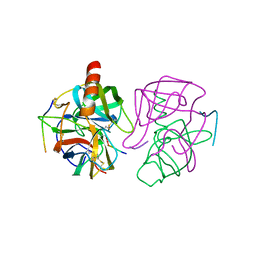 | | THE X-RAY CRYSTAL STRUCTURE OF THE TETRAHEDRAL INTERMEDIATE OF GAMMA-CHYMOTRYPSIN IN HEXANE | | Descriptor: | GAMMA-CHYMOTRYPSIN A, PRO GLY ALA TYR PEPTIDE | | Authors: | Yennawar, N.H, Yennawar, H.P, Banerjee, S, Farber, G.K. | | Deposit date: | 1993-08-20 | | Release date: | 1993-10-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray crystal structure of gamma-chymotrypsin in hexane.
Biochemistry, 33, 1994
|
|
1GMD
 
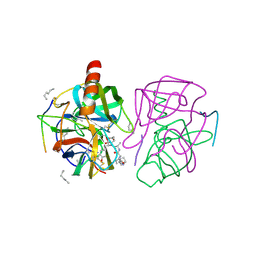 | | X-ray crystal structure of gamma-chymotrypsin in hexane | | Descriptor: | GAMMA-CHYMOTRYPSIN A, HEXANE, PRO GLY ALA TYR ASP PEPTIDE | | Authors: | Yennawar, N.H, Yennawar, H.P, Banerjee, S, Farber, G.K. | | Deposit date: | 1993-08-20 | | Release date: | 1993-10-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray crystal structure of gamma-chymotrypsin in hexane.
Biochemistry, 33, 1994
|
|
1ZZP
 
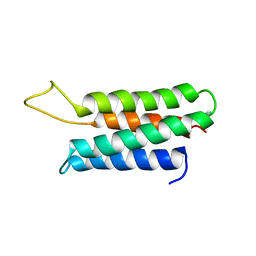 | | Solution structure of the F-actin binding domain of Bcr-Abl/c-Abl | | Descriptor: | Proto-oncogene tyrosine-protein kinase ABL1 | | Authors: | Hantschel, O, Wiesner, S, Guttler, T, Mackereth, C.D, Rix, L.L.R, Mikes, Z, Dehne, J, Gorlich, D, Sattler, M, Superti-Furga, G. | | Deposit date: | 2005-06-14 | | Release date: | 2005-08-30 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for the Cytoskeletal Association of Bcr-Abl/c-Abl.
Mol.Cell, 19, 2005
|
|
2CD2
 
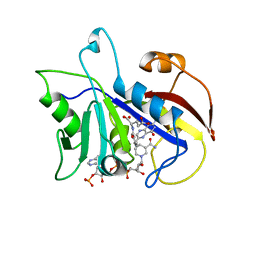 | | LIGAND INDUCED CONFORMATIONAL CHANGES IN THE CRYSTAL STRUCTURES OF PNEUMOCYSTIS CARINII DIHYDROFOLATE REDUCTASE COMPLEXES WITH FOLATE AND NADP+ | | Descriptor: | DIHYDROFOLATE REDUCTASE, FOLIC ACID, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Cody, V, Galitsky, N, Rak, D, Luft, J, Pangborn, W, Queener, S. | | Deposit date: | 1999-03-15 | | Release date: | 2000-03-29 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Ligand-induced conformational changes in the crystal structures of Pneumocystis carinii dihydrofolate reductase complexes with folate and NADP+.
Biochemistry, 38, 1999
|
|
7KJE
 
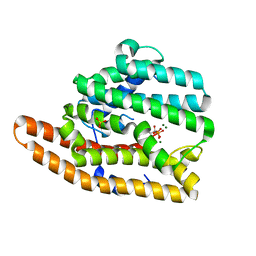 | | F96S epi-isozizaene synthase: complex with 3 Mg2+ and neridronate | | Descriptor: | (6-azanyl-1-oxidanyl-1-phosphono-hexyl)phosphonic acid, MAGNESIUM ION, SULFATE ION, ... | | Authors: | Ronnebaum, T.A, Gardner, S, Christianson, D.W. | | Deposit date: | 2020-10-26 | | Release date: | 2020-12-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | An Aromatic Cluster in the Active Site of epi -Isozizaene Synthase Is an Electrostatic Toggle for Divergent Terpene Cyclization Pathways.
Biochemistry, 59, 2020
|
|
2HZS
 
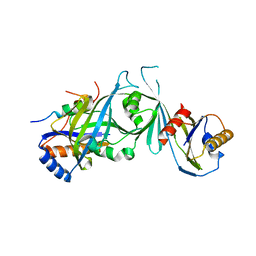 | | Structure of the Mediator head submodule Med8C/18/20 | | Descriptor: | RNA polymerase II mediator complex subunit 18, RNA polymerase II mediator complex subunit 20, RNA polymerase II mediator complex subunit 8 | | Authors: | Lariviere, L, Geiger, S, Hoeppner, S, Rother, S, Straesser, K, Cramer, P. | | Deposit date: | 2006-08-09 | | Release date: | 2006-09-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure and TBP binding of the Mediator head subcomplex Med8-Med18-Med20.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2B7E
 
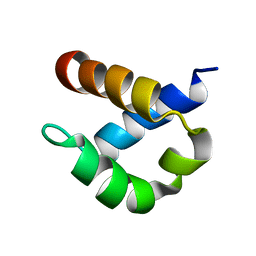 | | First FF domain of Prp40 Yeast Protein | | Descriptor: | Pre-mRNA processing protein PRP40 | | Authors: | Gasch, A, Wiesner, S, Martin-Malpartida, P, Ramirez-Espain, X, Ruiz, L, Macias, M.J. | | Deposit date: | 2005-10-04 | | Release date: | 2005-11-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of Prp40 FF1 domain and its interaction with the crn-TPR1 motif of Clf1 gives a new insight into the binding mode of FF domains.
J.Biol.Chem., 281, 2006
|
|
2MMZ
 
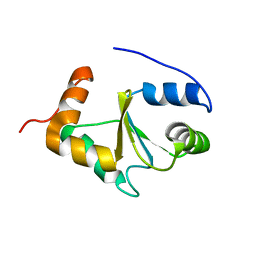 | | Solution structure of the apo form of human glutaredoxin 5 | | Descriptor: | Glutaredoxin-related protein 5, mitochondrial | | Authors: | Banci, L, Brancaccio, D, Ciofi-Baffoni, S, Del Conte, R, Gadepalli, R, Mikolajczyk, M, Neri, S, Piccioli, M, Winkelmann, J. | | Deposit date: | 2014-03-25 | | Release date: | 2014-04-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | [2Fe-2S] cluster transfer in iron-sulfur protein biogenesis.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
2MDR
 
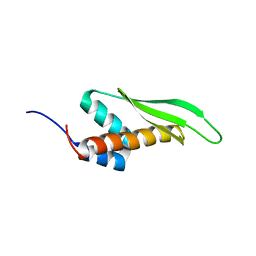 | | Solution structure of the third double-stranded RNA-binding domain (dsRBD3) of human adenosine-deaminase ADAR1 | | Descriptor: | Double-stranded RNA-specific adenosine deaminase | | Authors: | Barraud, P, Banerjee, S, Mohamed, W.I, Jantsch, M.F, Allain, F.H. | | Deposit date: | 2013-09-17 | | Release date: | 2014-04-30 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A bimodular nuclear localization signal assembled via an extended double-stranded RNA-binding domain acts as an RNA-sensing signal for transportin 1.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
5NPO
 
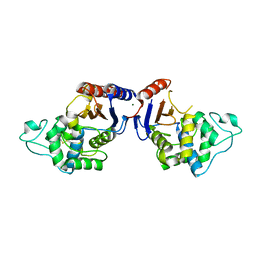 | | Promiscuous Protein Self-Assembly as a Function of Protein Stability | | Descriptor: | Beta-lactamase, Beta-lactamase TEM, MAGNESIUM ION | | Authors: | Cohen-Khait, R, Dym, O, Hamer-Rogotner, S, Schreiber, G. | | Deposit date: | 2017-04-18 | | Release date: | 2017-12-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Promiscuous Protein Binding as a Function of Protein Stability.
Structure, 25, 2017
|
|
5OAK
 
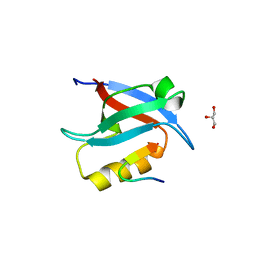 | |
2KIS
 
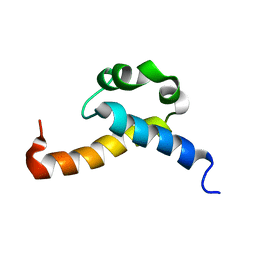 | | Solution structure of CA150 FF1 domain and FF1-FF2 interdomain linker | | Descriptor: | Transcription elongation regulator 1 | | Authors: | Murphy, J.M, Hansen, D, Wiesner, S, Muhandiram, D, Borg, M, Smith, M.J, Sicheri, F, Kay, L.E, Forman-Kay, J.D, Pawson, T. | | Deposit date: | 2009-05-08 | | Release date: | 2009-09-08 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structural studies of FF domains of the transcription factor CA150 provide insights into the organization of FF domain tandem arrays.
J.Mol.Biol., 393, 2009
|
|
2KZT
 
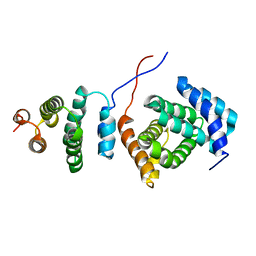 | | Structure of the Tandem MA-3 Region of Pdcd4 | | Descriptor: | Programmed cell death protein 4 | | Authors: | Waters, L.C, Strong, S.L, Oka, O, Muskett, F.W, Veverka, V, Banerjee, S, Schmedt, T, Henry, A.J, Klempnauer, K.H, Carr, M.D. | | Deposit date: | 2010-06-24 | | Release date: | 2011-03-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of the tandem MA-3 region of Pdcd4 protein and characterization of its interactions with eIF4A and eIF4G: molecular mechanisms of a tumor suppressor
J.Biol.Chem., 286, 2011
|
|
2KRB
 
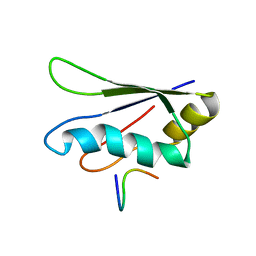 | | Solution structure of EIF3B-RRM bound to EIF3J peptide | | Descriptor: | Eukaryotic translation initiation factor 3 subunit B, Eukaryotic translation initiation factor 3 subunit J | | Authors: | Elantak, L, Wagner, S, Herrmannova, A, Janoskova, M, Rutkai, E, Lukavsky, P.J, Valasek, L. | | Deposit date: | 2009-12-16 | | Release date: | 2010-01-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The indispensable N-terminal half of eIF3j/HCR1 co-operates with
its structurally conserved binding partner eIF3b/PRT1-RRM and eIF1A in
stringent AUG selection
To be Published
|
|
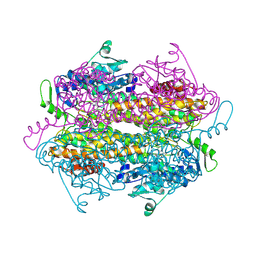
2OHY
 
 | |
7K32
 
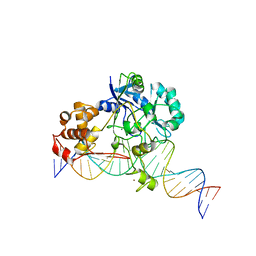 | | Crystal structure of Endonuclease Q complex with 27-mer duplex substrate with an abasic lesion at the active site | | Descriptor: | DNA (27-MER), Endonuclease Q, MAGNESIUM ION, ... | | Authors: | Shi, K, Moeller, N.M, Banerjee, S, Yin, L, Orellana, K, Aihara, H. | | Deposit date: | 2020-09-10 | | Release date: | 2021-03-17 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Structural basis for recognition of distinct deaminated DNA lesions by endonuclease Q.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7K33
 
 | | Crystal structure of Endonuclease Q complex with 27-mer duplex substrate with an abasic lesion at the active site | | Descriptor: | DNA (27-MER), Endonuclease Q, MAGNESIUM ION, ... | | Authors: | Shi, K, Moeller, N.M, Banerjee, S, Yin, L, Orellana, K, Aihara, H. | | Deposit date: | 2020-09-10 | | Release date: | 2021-03-17 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Structural basis for recognition of distinct deaminated DNA lesions by endonuclease Q.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7K31
 
 | | Crystal structure of Endonuclease Q complex with 27-mer duplex substrate with dI at the active site | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, DNA (27-MER), ... | | Authors: | Shi, K, Moeller, N.M, Banerjee, S, Yin, L, Orellana, K, Aihara, H. | | Deposit date: | 2020-09-10 | | Release date: | 2021-03-17 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | Structural basis for recognition of distinct deaminated DNA lesions by endonuclease Q.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7K30
 
 | | Crystal structure of Endonuclease Q complex with 27-mer duplex substrate with dU at the active site | | Descriptor: | 1,2-ETHANEDIOL, DNA (27-MER), Endonuclease Q, ... | | Authors: | Shi, K, Moeller, N.M, Banerjee, S, Yin, L, Orellana, K, Aihara, H. | | Deposit date: | 2020-09-10 | | Release date: | 2021-03-17 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Structural basis for recognition of distinct deaminated DNA lesions by endonuclease Q.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
