8CAV
 
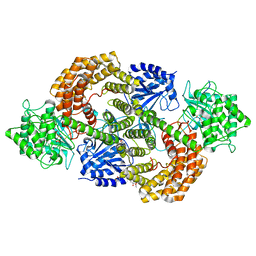 | | Discovery of the lanthipeptide Curvocidin and structural insights into its trifunctional synthetase CuvL | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CuvA, MAGNESIUM ION, ... | | Authors: | Sigurdsson, A, Martins, B.M, Duettmann, S.A, Jasyk, M, Dimos-Roehl, B, Schoepf, F, Gemannter, M, Knittel, C.H, Schnegotyzki, R, Schmid, B, Kosol, S, Gonzalez-Viegas, M, Seidel, M, Huegelland, M, Leimkuehler, S, Dobbek, H, Mainz, A, Suessmuth, R. | | Deposit date: | 2023-01-24 | | Release date: | 2023-06-14 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Discovery of the Lanthipeptide Curvocidin and Structural Insights into its Trifunctional Synthetase CuvL.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
4WM8
 
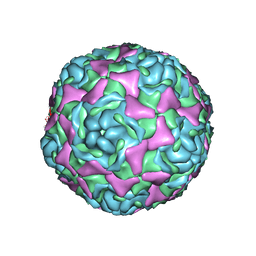 | | Crystal Structure of Human Enterovirus D68 | | Descriptor: | DECANOIC ACID, VP1, VP2, ... | | Authors: | Liu, Y, Sheng, J, Fokine, A, Meng, G, Long, F, Kuhn, R.J, Rossmann, M.G. | | Deposit date: | 2014-10-08 | | Release date: | 2015-01-14 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Virus structure. Structure and inhibition of EV-D68, a virus that causes respiratory illness in children.
Science, 347, 2015
|
|
4NAG
 | | Xanthomonins I III are a New Class of Lasso Peptides Featuringa Seven-Membered Macrolactam Ring | | Descriptor: | HEXANE-1,6-DIOL, Xanthomonin I | | Authors: | Hegemann, J.D, Zimmermann, M, Zhu, S, Steuber, H, Harms, K, Xie, X, Marahiel, M.A. | | Deposit date: | 2013-10-22 | | Release date: | 2014-04-30 | | Last modified: | 2014-05-21 | | Method: | X-RAY DIFFRACTION (0.81 Å) | | Cite: | Xanthomonins I-III: A New Class of Lasso Peptides with a Seven-Residue Macrolactam Ring.
Angew.Chem.Int.Ed.Engl., 53, 2014
|
|
7CAT
 
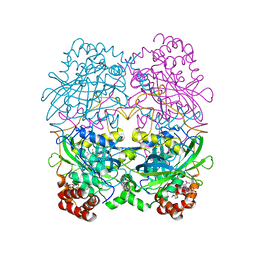 | | The NADPH binding site on beef liver catalase | | Descriptor: | CATALASE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Murthy, M.R.N, Reid III, T.J, Sicignano, A, Tanaka, N, Fita, I, Rossmann, M.G. | | Deposit date: | 1984-11-15 | | Release date: | 1985-04-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The NADPH binding site on beef liver catalase.
Proc.Natl.Acad.Sci.USA, 82, 1985
|
|
3J42
 
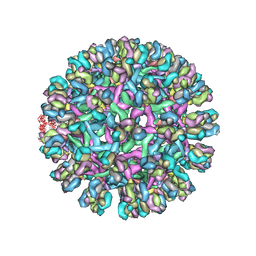 | | Obstruction of Dengue Virus Maturation by Fab Fragments of the 2H2 Antibody | | Descriptor: | Envelope protein E, Ig heavy chain V region MOPC 21, Igh protein chimera, ... | | Authors: | Wang, Z, Pennington, J.G, Jiang, W, Rossmann, M.G. | | Deposit date: | 2013-06-13 | | Release date: | 2013-07-17 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY (21 Å) | | Cite: | Obstruction of Dengue Virus Maturation by Fab Fragments of the 2H2 Antibody.
J.Virol., 87, 2013
|
|
4Z26
 
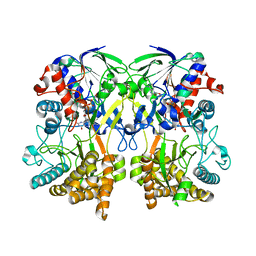 | | Mimivirus R135 (residues 51-702) | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Putative GMC-type oxidoreductase R135 | | Authors: | Klose, T, Rossmann, M.G. | | Deposit date: | 2015-03-28 | | Release date: | 2015-05-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.915 Å) | | Cite: | A Mimivirus Enzyme that Participates in Viral Entry.
Structure, 23, 2015
|
|
7B9S
 
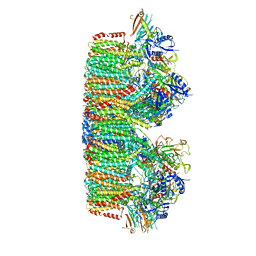 | | Structure of the mycobacterial ESX-5 Type VII Secretion System hexameric pore complex | | Descriptor: | EccB5, EccC5, EccD5, ... | | Authors: | Chojnowski, G, Ritter, C, Beckham, K.S.H, Mullapudi, E, Rettel, M, Savitski, M.M, Mortensen, S.A, Ziemianowicz, D, Kosinski, J, Wilmanns, M. | | Deposit date: | 2020-12-14 | | Release date: | 2021-07-07 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structure of the mycobacterial ESX-5 type VII secretion system pore complex.
Sci Adv, 7, 2021
|
|
2A28
 | | Atomic-resolution crystal structure of the second SH3 domain of yeast Bzz1 determined from a pseudomerohedrally twinned crystal | | Descriptor: | BZZ1 protein | | Authors: | Kursula, P, Kursula, I, Lehmann, F, Zou, P, Song, Y.H, Wilmanns, M. | | Deposit date: | 2005-06-22 | | Release date: | 2006-09-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.07 Å) | | Cite: | Structural genomics of yeast SH3 domains
To be Published
|
|
1WAA
 
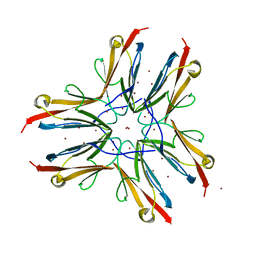 | | IG27 protein domain | | Descriptor: | TITIN, ZINC ION | | Authors: | Vega, M.C, Valencia, L, Zou, P, Wilmanns, M. | | Deposit date: | 2004-10-25 | | Release date: | 2006-07-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Mechanical Network in Titin Immunoglobulin from Force Distribution Analysis.
Plos Comput.Biol., 5, 2009
|
|
1Z9X
 
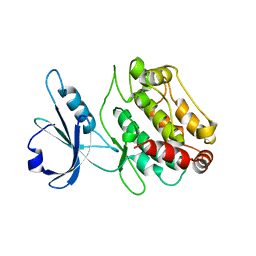 | | Human DRP-1 kinase, W305S S308A D40 mutant, crystal form with 3 monomers in the asymmetric unit | | Descriptor: | Death-associated protein kinase 2 | | Authors: | Kursula, P, Lehmann, F, Shani, G, Kimchi, A, Wilmanns, M. | | Deposit date: | 2005-04-05 | | Release date: | 2006-10-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.93 Å) | | Cite: | A structural insight into the double-locking mechanism of the human death-associated DRP-1 kinase
To be Published
|
|
2NSU
 
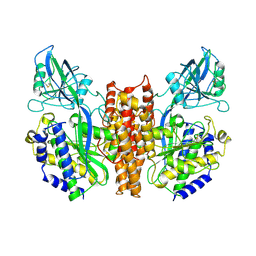 | |
3C6R
 
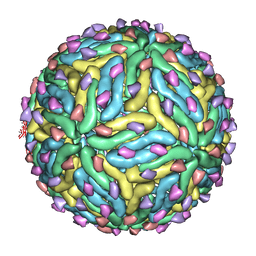 | | Low pH Immature Dengue Virus | | Descriptor: | Envelope protein, Peptide pr | | Authors: | Yu, I, Zhang, W, Holdway, H.A, Li, L, Kostyuchenko, V.A, Chipman, P.R, Kuhn, R.J, Rossmann, M.G, Chen, J. | | Deposit date: | 2008-02-05 | | Release date: | 2008-04-22 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (25 Å) | | Cite: | Structure of the immature dengue virus at low pH primes proteolytic maturation
Science, 319, 2008
|
|
2A4Z
 
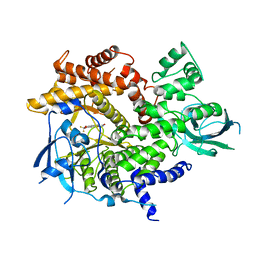 | | Crystal Structure of human PI3Kgamma complexed with AS604850 | | Descriptor: | (5E)-5-[(2,2-DIFLUORO-1,3-BENZODIOXOL-5-YL)METHYLENE]-1,3-THIAZOLIDINE-2,4-DIONE, Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit, gamma isoform | | Authors: | Camps, M, Ruckle, T, Ji, H, Ardissone, V, Rintelen, F, Shaw, J, Ferrandi, C, Chabert, C, Gillieron, C, Francon, B, Martin, T, Gretener, D, Perrin, D, Leroy, D, Vitte, P.-A, Hirsch, E, Wymann, M.P, Cirillo, R, Schwarz, M.K, Rommel, C. | | Deposit date: | 2005-06-30 | | Release date: | 2005-09-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Blockade of PI3Kgamma suppresses joint inflammation and damage in mouse models of rheumatoid arthritis
NAT.MED. (N.Y.), 11, 2005
|
|
1M0F
 
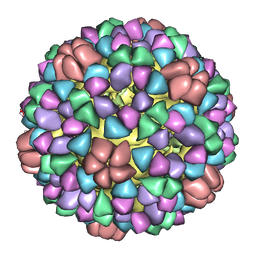 | | Structural Studies of Bacteriophage alpha3 Assembly, Cryo-electron microscopy | | Descriptor: | Capsid Protein F, Major Spike Protein G, Scaffolding Protein B, ... | | Authors: | Bernal, R.A, Hafenstein, S, Olson, N.H, Bowman, V.D, Chipman, P.R, Baker, T.S, Fane, B.A, Rossmann, M.G. | | Deposit date: | 2002-06-12 | | Release date: | 2002-12-25 | | Last modified: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (16 Å) | | Cite: | Structural Studies of Bacteriophage alpha3 Assembly
J.Mol.Biol., 325, 2003
|
|
2AB7
 
 | | Solution structures and characterization of HIV RRE IIB RNA targeting zinc finger proteins | | Descriptor: | ZINC ION, ZNF29G29R | | Authors: | Mishra, S.H, Shelley, C.M, Darby, M.K, Germann, M.W. | | Deposit date: | 2005-07-14 | | Release date: | 2005-08-02 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structures and characterization of human immunodeficiency virus Rev responsive element IIB RNA targeting zinc finger proteins.
Biopolymers, 83, 2006
|
|
1ORV
 
 | | Crystal Structure of Porcine Dipeptidyl Peptidase IV (CD26) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, SULFATE ION, ... | | Authors: | Engel, M, Hoffmann, T, Wagner, L, Wermann, M, Heiser, U, Kiefersauer, R, Huber, R, Bode, W, Demuth, H.U, Brandstetter, H. | | Deposit date: | 2003-03-16 | | Release date: | 2003-05-06 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Crystal Structure of Dipeptidyl Peptidase IV (CD26) Reveals its Functional Regulation and Enzymatic Mechanism
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
2A5U
 
 | | Crystal Structure of human PI3Kgamma complexed with AS605240 | | Descriptor: | (5E)-5-(QUINOXALIN-6-YLMETHYLENE)-1,3-THIAZOLIDINE-2,4-DIONE, Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit, gamma isoform | | Authors: | Camps, M, Ruckle, T, Ji, H, Ardissone, V, Rintelen, F, Shaw, J, Ferrandi, C, Chabert, C, Gillieron, C, Francon, B, Martin, T, Gretener, D, Perrin, D, Leroy, D, Vitte, P.-A, Hirsch, E, Wymann, M.P, Cirillo, R, Schwarz, M.K, Rommel, C. | | Deposit date: | 2005-07-01 | | Release date: | 2005-09-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Blockade of PI3Kgamma suppresses joint inflammation and damage in mouse models of rheumatoid arthritis
NAT.MED. (N.Y.), 11, 2005
|
|
2AB3
 | | Solution structures and characterization of HIV RRE IIB RNA targeting zinc finger proteins | | Descriptor: | ZINC ION, ZNF29 | | Authors: | Mishra, S.H, Shelley, C.M, Darby, M.K, Germann, M.W. | | Deposit date: | 2005-07-14 | | Release date: | 2005-08-02 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structures and characterization of human immunodeficiency virus Rev responsive element IIB RNA targeting zinc finger proteins.
Biopolymers, 83, 2006
|
|
4K8A
 
 | | Fragment-based discovery of Focal Adhesion Kinase Inhibitors | | Descriptor: | 3-bromo-5-(2H-tetrazol-5-yl)pyridine, Focal adhesion kinase 1 | | Authors: | Graedler, U, Bomke, J, Musil, D, Dresing, V, Lehmann, M, Hoelzemann, G, Esdar, C, Krier, M, Heinrich, T. | | Deposit date: | 2013-04-18 | | Release date: | 2013-09-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Fragment-based discovery of focal adhesion kinase inhibitors.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
3D18
 
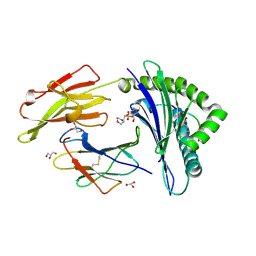 | | Crystal structure of HLA-B*2709 complexed with a variant of the latent membrane protein 2 peptide (LMP2(L)) of epstein-barr virus | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Beta-2-microglobulin, ... | | Authors: | Beltrami, A, Gabdulkhakov, A, Rossmann, M, Ziegler, A, Uchanska-Ziegler, B, Saenger, W. | | Deposit date: | 2008-05-05 | | Release date: | 2009-05-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Crystal structure of HLA-B*2709 complexed with a variant of the latent membrane protein 2 peptide (LMP2(L)) of epstein-barr virus
To be Published
|
|
1J0X
 
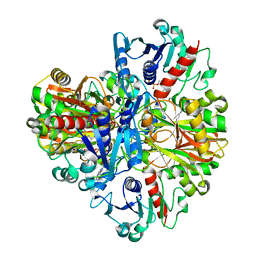 | | Crystal structure of the rabbit muscle glyceraldehyde-3-phosphate dehydrogenase (GAPDH) | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, glyceraldehyde-3-phosphate dehydrogenase | | Authors: | Cowan-Jacob, S.W, Kaufmann, M, Anselmo, A.N, Stark, W, Grutter, M.G. | | Deposit date: | 2002-11-25 | | Release date: | 2003-12-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure of rabbit-muscle glyceraldehyde-3-phosphate dehydrogenase.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
2QZD
 | | Fitted structure of SCR4 of DAF into cryoEM density | | Descriptor: | Complement decay-accelerating factor | | Authors: | Hafenstein, S, Bowman, V.D, Chipman, P.R, Bator Kelly, C.M, Lin, F, Medof, M.E, Rossmann, M.G. | | Deposit date: | 2007-08-16 | | Release date: | 2007-10-30 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY (14 Å) | | Cite: | Interaction of decay-accelerating factor with coxsackievirus b3.
J.Virol., 81, 2007
|
|
2GO5
 
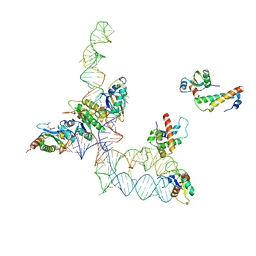 | | Structure of signal recognition particle receptor (SR) in complex with signal recognition particle (SRP) and ribosome nascent chain complex | | Descriptor: | SRP RNA, Signal recognition particle 19 kDa protein (SRP19), Signal recognition particle 54 kDa protein (SRP54), ... | | Authors: | Halic, M, Gartmann, M, Schlenker, O, Mielke, T, Pool, M.R, Sinning, I, Beckmann, R. | | Deposit date: | 2006-04-12 | | Release date: | 2006-06-13 | | Last modified: | 2011-07-13 | | Method: | ELECTRON MICROSCOPY (7.4 Å) | | Cite: | Signal Recognition Particle Receptor Exposes the Ribosomal Translocon Binding Site
Science, 312, 2006
|
|
2QZF
 
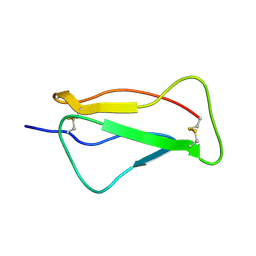 | | SCR1 of DAF from 1ojv fitted into cryoEM density | | Descriptor: | Complement decay-accelerating factor | | Authors: | Hafenstein, S, Bowman, V.D, Chipman, P.R, Bator Kelly, C.M, Lin, F, Medof, M.E, Rossmann, M.G. | | Deposit date: | 2007-08-16 | | Release date: | 2007-10-30 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY (14 Å) | | Cite: | Interaction of decay-accelerating factor with coxsackievirus b3.
J.Virol., 81, 2007
|
|
3CSZ
 
 | |
